Abstract
Esophageal squamous cell carcinoma (ESCC) is a deadly malignancy with notable metabolic reprogramming, yet the pivotal metabolic feature driving ESCC progression remains elusive. Here, we show that methionine cycle exhibits robust activation in ESCC and is reversely associated with patient survival. ESCC cells readily harness exogenous methionine to generate S-adenosyl-methionine (SAM), thus promoting cell proliferation. Mechanistically, methionine augments METTL3-mediated RNA m6A methylation through SAM and revises gene expression. Integrative omics analysis highlights the potent influence of methionine/SAM on NR4A2 expression in a tumor-specific manner, mediated by the IGF2BP2-dependent stabilization of methylated NR4A2 mRNA. We demonstrate that NR4A2 facilitates ESCC growth and negatively impacts patient survival. We further identify celecoxib as an effective inhibitor of NR4A2, offering promise as a new anti-ESCC agent. In summary, our findings underscore the active methionine cycle as a critical metabolic characteristic in ESCC, and pinpoint NR4A2 as a novel methionine-responsive oncogene, thereby presenting a compelling target potentially superior to methionine restriction.
Similar content being viewed by others
Introduction
Esophageal squamous cell carcinoma (ESCC), a major histological type of esophageal cancer, exhibits an inferior prognosis as characterized by a median overall survival (OS) ranging from 7 to 14 months and a 5-year OS rate below 20% [1]. To better treat this disorder, it is imperative to comprehensively unveil its underlying pathobiological characteristics. As we and others demonstrate that metabolic reprogramming has emerged as a critical factor in tumorigenesis and tumor progression, producing new and valuable insights into cancer pathobiology [2,3,4,5,6]. Previous investigations have revealed the significant upregulation of various metabolic pathways in ESCC, such as proline/fatty acid/polyamine biosynthesis and glutamine/histidine/uridine metabolism [7,8,9]. Nevertheless, the key metabolic signature of ESCC in clinical tissue samples remains elusive. In our recent studies utilizing tissue and serum samples from patients with ESCC, we conducted comprehensive metabolomic analyses to dissect the metabolic features of this disease [10, 11]. However, in these studies, we focus on the perturbations but not the activity of enriched metabolic pathways in ESCC. In the current study, we re-analyzed the tissue metabolic data derived from our previous study [10] from the perspective of pathway activity and discovered a significant activation of the methionine cycle in clinical ESCC tissues. Nevertheless, a nutritional epidemiology study shows that higher intake of methionine is not associated with ESCC risk [12]. This finding confounds the role of the methionine in ESCC.
Clinical observations using positron emission tomography imaging have provided compelling evidence of an overt increase in intratumoral methionine uptake in patients with brain tumor, glioma, or multiple myeloma [13,14,15], indicating a pronounced reliance on this essential amino acid by human tumors. Methionine metabolism primarily involves in three key metabolic pathways: the methionine cycle, transsulfuration pathway, and methionine salvage pathway [16,17,18]. By regulating multiple cellular processes, methionine metabolism exerts control over cancer traits, including cancer growth, stemness, and therapeutic sensitivity [16,17,18,19]. For instance, methionine metabolism is tightly coupled to one-carbon metabolic network through the remethylation of homocysteine, thereby amplifying nucleotide synthesis [20]. Furthermore, methionine plays a crucial role in maintaining cellular redox homeostasis by providing homocysteine to fuel the transsulfuration pathway and generate the essential antioxidant glutathione [16, 18]. Additionally, methionine cycle is involved in chromatin and nucleic acid methylation through the supply of S-adenosyl-methionine (SAM), a universal methyl donor [21, 22]. The addiction of cancer cells to methionine has prompted investigations of methionine-restricted diets in combination with chemotherapy in both preclinical models and clinical trials, demonstrating promising anti-cancer efficacy [23,24,25]. However, concerns regarding the appropriate level and duration of methionine restriction (MR) have arisen, given the amino acid’s importance in maintaining muscle mass, neuronal functions, and redox balance [16, 18, 24, 26]. To maximize the anti-cancer potential of MR while minimizing latent adverse effect, it is imperative to identify key downstream players in methionine metabolism that contribute to tumor progression.
It is worth noting that the methionine cycle is intimately linked to intracellular RNA methylation by providing SAM [27, 28]. Among the various modifications of messenger RNA (mRNA), N6-methyladenosine (m6A) is the most prevalent and it significantly impacts multiple mRNA processes, including alternative splicing, nuclear exportation, decay, stabilization, and translation [29,30,31]. The regulation of intracellular mRNA m6A methylation is orchestrated by a complex machinery consisting of writer, reader, and eraser proteins [29, 30, 32]. The core subunits of the writer complex comprise methyltransferase-like 3 (METTL3), methyltransferase-like 14 (METTL14), methyltransferase-like 16 (METTL16), while well-known eraser proteins are fat mass and obesity-associated protein (FTO) and alkB homolog 5 (ALKBH5). Reader proteins involved in this process include YT521-B homology domain-containing family proteins (YTHDFs) and insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs) [30,31,32,33]. Increasing lines of evidence highlight the importance of mRNA m6A methylation in cancer initiation and progression [28, 34, 35]. For instance, METTL3-mediated mRNA m6A modification, followed by YTHDF2-dependent degradation of the m6A-methylated mRNA of suppressor of cytokine signaling 2, a tumor suppressor, promotes liver cancer growth and migration [33].
Based on our preliminary findings demonstrating the ectopic activation of the methionine cycle in ESCC, we have formulated a hypothesis suggesting that this metabolic pathway can generate an abundant supply of SAM. Consequently, we propose that this elevated SAM level can perturb the RNA m6A pattern and induce changes in the expression of specific genes important for ESCC development. This study aims to thoroughly investigate and validate this hypothesis, therefore elucidating the importance of methionine addiction for ESCC and identifying methionine-responsive genes that could serve as new therapeutic targets for ESCC treatment.
Results
Methionine cycle is hyperactive in clinical ESCC tissues and predicts unfavorable patient survival
In our previous study for the metabolic features in clinical ESCC tissues [10], we concentrate on the perturbations of enriched metabolic pathways using the algorithm of metabolite set enrichment analysis [36, 37]. To assess the changes in the activity of metabolic pathways in clinical ESCC tissues, we used the reported ESCC cohort as a discovery cohort (n = 24, Supplementary Table S1) [10] and conducted a reanalysis of tissue metabolomic dataset from this cohort. Our analysis revealed significant alterations in 56.50% of measured metabolites, with 83.19% of these differential metabolites upregulated in ESCC tissues compared to paired normal adjacent tissues (NATs) (Fig. 1A). Using Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway-based differential abundance analysis, we identified 27 enriched metabolic pathways, with nine of these pathways exhibiting upregulation (≥0.5 differential abundance score) in the tumor tissues (Fig. 1B). Notably, methionine metabolism was ranked first among these nine pathways (Fig. 1B), with elevated levels for nine out of 10 measured metabolites in this pathway (Supplementary Fig. S1A), indicating the aberrant activation of methionine metabolism in ESCC tissues. As previously described [18], the methionine cycle, a pathway in methionine metabolism, generates SAM to provide methyl groups for the methylation of diverse biological macromolecules (Fig. 1C). Our findings suggested the potential stimulation of this metabolic pathway in clinical ESCC tissues, as evidenced by increased levels of two out of four metabolites involved in the pathway, namely methionine and homocysteine (Hcy) (Supplementary Fig. S1A).
A Heatmap showing all measured metabolites in clinical ESCC tissues and paired NATs from the discovery cohort. The upper and lower panels represent differential (56.50%, 113/200) and non-differential (43.50%, 87/200) metabolites in tumor tissues, respectively. B Metabolic pathway enrichment analysis of clinical ESCC tissues from the discovery cohort. The differential abundance score represents the average change of measured metabolites in a pathway in ESCC tissues relative to paired NATs. Upregulation and downregulation of all measured metabolites in a pathway are indicated by red and cyan bars, respectively. Pathway size is defined by the quantity of measured metabolites participating in a particular metabolic pathway. C Schematic of the methionine cycle, highlighting the amino acid transporter and metabolic enzymes involved in this pathway (shown in red). D Differential metabolites of the methionine cycle between clinical ESCC tissues and paired NATs in validation cohorts 1 and 2. OS and DFS curves of patients with high and low PRS in validation cohort 1 (E) and validation cohort 2 (F). The cut-off value for PRS is determined in validation cohort 1 using the 70th percentile. The 95% confidence limits of each survival curve are represented by shadows. G Forest plot displaying OS hazard ratios of PRS and several well-known clinical parameters derived from a multivariate Cox regression model fitted using the combined validation cohort 1 and validation cohort 2.
Next, we enrolled two independent validation cohorts (n = 94 and n = 112, respectively; Supplementary Tables S2 and S3) to corroborate the activation of methionine cycle in clinical ESCC tissues. Unsupervised principal component analysis (PCA) of the tissue metabolomic data from these cohorts demonstrated low dispersion of quality control samples, indicating minimal systemic measurement variability (Supplementary Fig. S1B, C). Consistent with the findings in the discovery cohort, the levels of four metabolites involved in the methionine cycle (methionine, SAM, S-adenosyl-homocysteine (SAH), and Hcy) were markedly upregulated in clinical ESCC tissues compared to paired NATs in both validation cohorts (Fig. 1D), providing further confirmation of the overactivation of this pathway in ESCC. Subsequently, we investigated whether the activity of the methionine cycle in ESCC tissues correlated with patient survival. Using four intratumoral metabolites involved in the methionine cycle (illustrated in Fig. 1D), we developed a predictive PCA model for validation cohort 1 as described in the Materials and Methods section. This model was utilized to generate prognosis-risk score (PRS), determined by the first component value, for each patient in validation cohort 1, and to predict the PRS for each case in validation cohort 2. High PRS values predicted inferior overall survival (OS) and disease-free survival (DFS) in patients with ESCC in both validation cohorts (Fig. 1E, F). Multivariate Cox regression analysis demonstrated that high PRS values predicted poor OS and DFS in the combined validation cohort 1 and validation cohort 2, independent of well-established prognostic factors such as age, gender, treatment protocol, and TNM stage (Fig. 1G and Supplementary Fig. S1D). Notably, high PRS values were correlated with increased levels of methionine cycle metabolites (Supplementary Fig. S1E,F). These results highlighted that the significance of the upregulated methionine cycle as a new and independent predictive biomarker for unfavorable survival in patients with ESCC.
To further verify the aberrant activation of the methionine cycle in ESCC, we examined the expression patterns of the transporter and metabolic enzymes involved in this pathway, as depicted in Fig. 1C. First, we reanalyzed the tissue proteomic dataset from the previously reported discovery cohort [10] and observed a significant increase in the intratumoral levels of SLC7A5, a transporter responsible for methionine uptake, as well as three key metabolic enzymes involved in the methionine cycle: methionine adenosyltransferase 2A (MAT2A), methionine adenosyltransferase 2B (MAT2B), and adenosylhomocysteinase (AHCY) (Supplementary Fig. S2A). Second, we obtained a public gene expression dataset, GSE23400 [38], which consisted of 53 pairs of clinical ESCC tissues and matched NATs, to serve as an external validation dataset. Analysis of this dataset revealed upregulation of SLC7A5 and the three metabolic enzymes mentioned above at the transcriptional level in ESCC tissues (Supplementary Fig. S2B). To further verify our findings, we performed validation assay in the aforementioned two validation cohorts. As some tissue samples in validation cohort 1 and validation cohort 2 were depleted, we enrolled a new validation cohort 3 (n = 41, Supplementary Table S4). Immunohistochemistry (IHC) staining of tissue sections demonstrated an overt elevation in the protein levels of SLC7A5, MAT2A, MAT2B, and AHCY in clinical ESCC tissues as relative to NATs in all three validation cohorts (Supplementary Fig. S2C–H). Collectively, our results provided comprehensive evidence of the ectopic activation of the methionine cycle at both the metabolite and metabolic enzyme levels. Furthermore, we established a reverse association between this metabolic pathway and the prognosis of patients with ESCC.
High methionine promotes ESCC growth via SAM but low methionine causes adverse effect
As methionine is a well-known essential amino acid for human body, an important question arises regarding whether the active methionine cycle observed in clinical ESCC tissues signifies an avid uptake and utilization of exogenous methionine by ESCC cells. To address this question, we initially selected two representative ESCC cell lines, KYSE150 and Eca109, for in vitro assay. When compared to the cells cultured in a methionine-free medium, those cultured in a methionine-containing medium revealed increased levels of intracellular methionine and its downstream metabolite, SAM (Fig. 2A, B). Subsequently, we conducted ex vivo assays using freshly harvested ESCC tissues from five patients. Similar to the in vitro findings, the ESCC tissues cultured in a complete medium exhibited significantly higher levels of intratumoral methionine and SAM compared to the matched ESCC tissues cultured in a methionine-free medium (Fig. 2C). These results demonstrated that both the ESCC cell lines and primary fresh ESCC tissues from patients effectively consumed exogenous methionine to generate SAM. Moreover, ESCC cells cultivated in media enriched with SAH or Hcy displayed markedly elevated levels of intracellular SAH or Hcy compared to their counterparts cultured in methionine, SAH, and Hcy-free medium (Supplementary Fig. S3A, B). This observation suggested that ESCC cells were capable of assimilating exogenous SAH and Hcy.
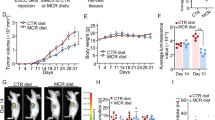
Comparison of intracellular methionine (A) and its downstream metabolite SAM (B) levels between ESCC cells cultured in medium with 10 μM methionine and methionine-free medium for 24 h. C Comparison of intratumoral methionine and SAM levels between primary fresh ESCC tissues cultured in medium with 10 μM methionine and methionine-free medium for 24 h. Five ESCC tissues from five patients were collected and cultured accordingly. Each ESCC tissue was halved and the acquired segments were cultured in methionine-containing and methionine-free media, respectively. Influence of dietary methionine on the growth of heterotopic ESCC tumors (D and E) and intratumoral expression of cyclin B1 and PCNA (F and G) in the subcutaneous xenograft mouse model. H Establishment of 4-NQO-induced orthotopic ESCC mouse model: Following a 16-week induction period, ESCC mice were randomly divided into two groups and administered pure water or water containing 10 mg/mL methionine. Propylene glycol (PG) solution, dissolved in water at a ratio of 1:60 (vol:vol), was used as vehicle. I Representative hematoxylin and eosin (H&E) and IHC staining of the esophagi from three groups of mice derived from the model described in H. Keratin 14 is used as a marker of esophageal epithelial squamous cells. J Quantification of the percentage of neoplastic cells with strong Ki-67 staining in esophageal tumors derived from 4-NQO-induced ESCC mice with or without methionine administration. Error bars represent mean ± standard error of the mean (SEM). *p < 0.05, **p < 0.01, ***p < 0.001 (Student’s t test).
We then sought to decipher the significance of methionine and SAM in the malignance of ESCC. First, in vitro assays were conducted using four ESCC cell lines, including KYSE150, Eca109, KYSE450, and KYSE30. The results demonstrated that methionine and SAM, but not SAH and Hcy, fostered the proliferation of ESCC cells in a dose-dependent manner (Supplementary Fig. S3C–F). The impact of exogenous methionine and SAM on the growth of ESCC cells may be confounded by the presence of diverse amino acids in standard fetal bovine serum (FBS) incorporated into the culture medium. To address this potential interference, a methionine-free medium supplemented with 10% dialyzed FBS was employed for validation purposes. The results validated that both exogenous methionine and SAM indeed boosted the growth of ESCC cells in a dose-dependent manner in the dialyzed medium (Supplementary Fig. S3G,H). Furthermore, methionine and SAM promoted G2/M phase progression and upregulated the expression of a G2/M progression marker cyclin B1 and a cell proliferation marker proliferating cell nuclear antigen (PCNA) in ESCC cells (Supplementary Fig. S3I–K). Second, in vivo investigations were carried out using a subcutaneous xenograft mouse model. It was observed that dietary methionine dramatically expedited heterotopic ESCC tumor growth in a dose-dependent manner, accompanied by enhanced intratumoral expression of cyclin B1 and PCNA (Fig. 2D–G). Mouse diets with distinct levels of methionine (Supplementary Table S5) were prepared as previously depicted [39]. Third, in vivo studies were performed using a 4-nitroquinoline 1-oxide (4-NQO)-induced ESCC mouse model. Oral administration of methionine via drinking water remarkably accelerated orthotopic ESCC tumor growth and upregulated the intratumoral expression of another cell proliferation marker, Ki-67 (Fig. 2H–J). Together, these findings demonstrated that exogenous methionine and its downstream metabolite SAM were essential for ESCC growth.
In line with previous findings showing the anti-cancer potency of MR against gastric cancer, colorectal cancer, and soft-tissue sarcoma [23,24,25], low methionine markedly hindered ESCC cell growth in vitro and in vivo (Fig. 2D–J and Supplementary Fig. S3C). Of note, as compared to high methionine diet, low methionine diet caused a substantial 11.6–14.7% body weight loss, a typical symptom of cancer-associated cachexia [40], in mice harboring ESCC xenograft tumors after 31 days of tumor transplantation (Supplementary Fig. S3L), indicating a considerable adverse effect of MR. This raised the necessity of identifying methionine-responsive oncogenes instead of MR as new therapeutic targets against ESCC.
RNA m6A methylation of ESCC is augmented through a methionine-SAM-MELLT3 cascade
Given the known role of methionine and its downstream metabolite SAM in promoting RNA m6A methylation [27, 28], and the importance of RNA m6A methylation in modulating gene expression [29, 30], we speculated that methionine would revise the expression of certain oncogenes via altering RNA m6A modification to foster ESCC growth. First, we sought to ascertain whether the m6A levels were elevated in intratumoral RNAs of clinical ESCC tissues. To address this, we extracted RNA samples from 48 ESCC tissues and their matched NATs derived from patient cohort 2 and examined the m6A intensity of these specimens using a dot blot assay. The result revealed a substantial increase in RNA m6A methylation in clinical ESCC tissues as relative to paired NATs (Fig. 3A, B). Of importance, upon removal of methionine from the culture medium, both ESCC cell lines and primary fresh ESCC tissues from patients exhibited an evident abatement of RNA m6A methylation (Fig. 3C, D). Notably, SAM supplementation dramatically restored this RNA modification (Fig. 3E). These findings conclusively demonstrated the essential role of methionine and its metabolite SAM in the maintenance of RNA m6A methylation in ESCC.
A Dot blot assay comparing RNA m6A content between clinical ESCC tissues and paired NATs from validation cohort 2 (n = 48). B Quantification of RNA m6A levels of paired tissues described in A, with p value calculated using paired Wilcoxon rank-sum test. Impact of methionine removal from culture medium on RNA m6A abundance in ESCC cell lines (C) and primary fresh ESCC tissues from patients (D) after 24 h of in vitro culture. E Recovery of intracellular RNA m6A content in the absence of methionine through supplementation with 100 μM SAM for 24 h. F Proteomic analysis of the expression of three writer enzymes and two cofactors in the RNA m6A machinery between clinical ESCC tissues and paired NATs from discovery cohort. G, H Influence of METTL3 abrogation on the intracellular RNA m6A intensity in ESCC cells in the presence of methionine. I Influence of STM2457 treatment on the intracellular RNA m6A abundance in ESCC cells in the presence of methionine. Comparison of METTL3 expression between ESCC tissues and NATs from validation cohort 1 (J and K) and validation cohort 2 (L and M). Representative IHC images (J and L) and statistical violin-box-scatter plots (K and M) are presented, with p values calculated using Wilcoxon rank-sum test. NC non-target control.
Subsequently, we aimed to uncover the underlying mechanism responsible for the installation of m6A in RNA in ESCC cells. Reanalysis of the proteomic data from the discovery cohort of our previously published study [10] manifested a significant upregulation of three m6A writer enzymes, namely METTL3, METTL14, and METTL16, along with two cofactors, WTAP and RBM15B, in clinical ESCC tissues (Fig. 3F). Among these writer enzymes, METTL3 has been reported to enhance the m6A modification in RNA transcripts in ESCC cells [41]. Therefore, we hypothesized that METTL3 mediated the methionine-induced RNA m6A methylation in ESCC cells. Indeed, the deletion of METTL3 severely impaired the methionine-elicited RNA m6A modification in ESCC cells (Fig. 3G, H). Consistently, treatment with a well-established METTL3 inhibitor, STM2457 [42], also remarkably restrained the methionine-induced RNA m6A signal in ESCC cells (Fig. 3I). Moreover, analysis of clinical samples revealed that the expression of METTL3 was prominently increased in clinical ESCC tissues from three validation cohorts as compared to NATs (Fig. 3J–M and Supplementary Fig. S4).
Collectively, our findings demonstrated that RNA m6A modification was notably upregulated in clinical ESCC tissues through a methionine-SAM-METTLE3 cascade.
Integrative omics analysis reveals a positive linkage between the expression of NR4A2 and the m6A intensity in its mRNA under methionine stimulation
Methionine metabolism has been shown to impact RNA m6A methylation by providing the universal methyl donor SAM, consequently modifying gene expression [30, 43]. Based on our findings, we hypothesized that methionine would alter intracellular RNA m6A patterns and gene expression profiles to promote ESCC progression. To investigate this, we conducted RNA-sequencing (RNA-seq) analyses using KYSE150 cells cultured in vitro and esophageal tissues from 4-NQO-induced ESCC mice to identify methionine-regulated genes. There were 8,527 genes to be identified in both in vitro and in vivo models. Gene-wise (inter-sample) correlation analysis revealed a striking discrepancy between the in vitro and in vivo conditions regarding the impact of methionine administration on gene expression, as shown by a median gene-wise correlation value of −0.024 and a low percentage (4.2%) of genes with significant positive Spearman correlations (Supplementary Fig. S5A). Subsequently, we observed significant perturbations in the expression of 533 genes in the in vitro model and 578 genes in the in vivo model upon methionine treatment (Benjamini–Hochberg adjusted p < 0.05, fold change (FC) > 2 or <−2) (Fig. 4A, B). Importantly, Venn diagram revealed seven genes consistently modulated by methionine in both in vitro and in vivo settings, including nuclear receptor subfamily 4 group A member 2 (NR4A2), aldehyde dehydrogenase 3 family member B1 (ALDH3B1), ankyrin repeat domain 37 (ANKRD37), growth arrest specific 7 (GAS7), SRY-box transcription factor 9 (SOX9), dual specificity phosphatase 5 (DUSP5), and 4-hydroxy-2-oxoglutarate aldolase 1 (HOGA1) (Fig. 4C).
A Hierarchically clustered heatmap showing differentially expressed genes between KYSE150 cells cultured in medium with 10 μM methionine and methionine-free medium. B Hierarchically clustered heatmap displaying differentially expressed genes in neoplastic esophagi of 4-NQO-induced ESCC mice fed with methionine-containing water and pure water. C Venn diagram showing seven genes consistently modulated by methionine in both KYSE150 cells in vitro and 4-NQO-induced ESCC mice in vivo. D Predominant consensus motif, GGACU, identified from the top 1000 m6A peaks in KYSE150 cells. E Pie charts illustrating the distribution of m6A peaks in different regions across the entire set of mRNA transcripts of KYSE150 cells treated with or without methionine. Methionine-elicited m6A peaks are predominantly enriched in 3′-boundaries and 3′-UTRs of mRNAs. F Integrative analysis of methionine-elicited expression changes and m6A peaks for seven methionine-modulated genes in C in esophageal tissues of 4-NQO-induced mice in vivo and KYSE150 cells in vitro. G Analysis of m6A peaks in NR4A2 mRNAs between KYSE150 cells cultured in medium with 10 μM methionine and methionine-free medium, identifying four strongly methionine-provoked m6A peaks. H The fusion of NR4A2 promoter sequence with firefly luciferase reporter and the influence of methionine supplement on the relative F-luc mRNA abundance of NR4A2 promoter reporter in HEK 293T cells. UTR untranslated regions, CDS coding sequence.
Subsequently, we employed methylated RNA immunoprecipitation sequencing (MeRIP-seq) to investigate the influence of methionine on RNA m6A signature in KYSE150 cells. Consistent with previous study [44], the consensus m6A motif of GGACU was highly enriched in the immunopurified mRNAs of KYSE150 cells (Fig. 4D). Of note, the methionine-induced m6A peaks were predominantly located near 3′-boundaries and in 3′-untranslated regions (UTRs) of mRNAs (Fig. 4E, Supplementary Fig. S5B, C). Further analysis revealed that methionine treatment significantly augmented m6A modification at 3200 sites while attenuating it at 3,029 sites within RNA transcripts (p < 0.005, FC > 2 or < −2) (Supplementary Table S6). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis revealed that m6A-modified transcripts were enriched in cancer-related pathways (such as pathways in cancer, Notch signaling pathway, and pancreatic cancer pathway) and biological processes (such as cell cycle and gap junction pathways, and AGE-RAGE signaling pathway in diabetic complications) (Supplementary Fig. S5D).
We then performed an integrative analysis of the seven genes consistently regulated by methionine in vitro and in vivo, using RNA-seq and MeRIP-seq data. Among these genes, NR4A2, ALDH3B1, ANKRD37, GAS7, SOX9, and DUSP5 exhibited upregulation, while HOGA1 showed downregulation upon methionine treatment. Notably, m6A peaks were observed only in NR4A2, ALDH3B1, ANKRD37, SOX9, and DUSP5 (Fig. 4F). This finding suggested that methionine may elicit gene expression changes through m6A modification. We noticed that, in the presence of methionine, there were four major m6A peaks in both NR4A2 and DUSP5 mRNAs, and three m6A peaks in the NR4A2 mRNA had FC values > 10 (Fig. 4F and Supplementary Fig. S5E–J, and Supplementary Table S6). Therefore, we focused on NR4A2 for further investigation. Of note, four major m6A peaks in the NR4A2 mRNA elicited by methionine were located in the exon 3, exon 6, exon 8, and 3′-UTR (Fig. 4G). Therefore, methionine-induced m6A methylation in the NR4A2 mRNA was closely linked to the upregulation of NR4A2 expression in the presence of methionine. Furthermore, we constructed a reporter minigene harboring the promoter sequence of NR4A2. The supplement of methionine did not perturb the expression of the NR4A2 promoter (Fig. 4H), suggesting that methionine treatment increased the expression of NR4A2 not via upregulating its transcription. Taken together, our results demonstrated that methionine administration remarkably perturbed the patterns of gene expression and RNA m6A methylation in ESCC cells, particularly inducing the expression and m6A modification of NR4A2.
Methionine stimulates NR4A2 expression by enhancing its mRNA stability through a SAM-METTL3-m6A-IGF2BP2 cascade
Our analyses using RNA-seq, quantitative reverse transcription polymerase chain reaction (qRT-PCR), western blot and IHC assays revealed that methionine induced NR4A2 expression in ESCC at both the mRNA and protein levels in both in vitro and in vivo models (Figs. 4F and 5A, B). Importantly, methionine specifically elicited NR4A2 expression in tumor cells, as there was no change in NR4A2 expression in the presence of methionine in a non-transformed esophageal epithelial cell line, Het-1A (Fig. 5A). Furthermore, SAM also stimulated NR4A2 expression in ESCC cells at both the mRNA and protein levels (Supplementary Fig. S6A), indicating that methionine-elicited NR4A2 expression occurred through its downstream metabolite SAM.
A, B Influence of methionine on NR4A2 expression at mRNA and protein levels in ESCC cells and the non-transformed esophageal epithelial cell line Het-1A. Cells were cultured for 24 h. B Effect of methionine administration via drinking water on the protein abundance of NR4A2 in the neoplastic cells of esophagi in the 4-NQO-induced ESCC mouse model. Treatment with the methylation inhibitor DAA at 10 μM for 24 h affecting methionine-elicited RNA m6A methylation (C) and NR4A2 expression at mRNA and protein levels (D) in ESCC cells. E Perturbation of methionine-elicited NR4A2 expression at both mRNA and protein levels in ESCC cells by METTL3 ablation. F NR4A2 mRNA degradation curves over time in ESCC cells treated with or without methionine in the presence of the transcription inhibitor actinomycin D. G mRNA degradation curves of NR4A2 over time in ESCC cells with or without METTL3 deletion under the treatment of actinomycin D. Cells were cultured in complete medium. H RIP-qPCR revealing the association of NR4A2 mRNA with IGF2BPs in KYSE150 cells. I mRNA degradation curves of NR4A2 over time in ESCC cells with or without IGF2BP2 deletion under the treatment of actinomycin D. Cells were cultured in complete medium. J Disturbance of NR4A2 protein levels in ESCC cells by IGF2BP2 deletion. Cells were cultured in complete medium. K Schematic diagram showing three predicted m6A motifs within NR4A2 mRNA with high confidence, and construction of three NR4A2 mutants containing these motifs. Mut1 and Mut2 represent full-length coding sequence mutants of NR4A2 with A-T substitutions (shown in red) within m6A consensus. Mut3 represents a 715-bp fragment of the 3′-UTR of NR4A2 with an A-T substitution (shown in red) within m6A consensus. L mRNA degradation curves of wild-type NR4A2 (NR4A2-WT), NR4A2 Mutant 1 (NR4A2-Mut1), and NR4A2 Mutant 2 (NR4A2-Mut2) in the presence of methionine, along with NR4A2-WT in the absence of methionine, under the treatment of actinomycin D. M Relative F-luc mRNA levels of NR4A2-WT or NR4A2 Mutant 3 (NR4A2-Mut3) reporters in HEK 293T cells with or without methionine treatment. NC non-target control, WT wild-type, Mut mutant, F-luc firefly luciferase, R-luc renilla luciferase. Error bars represent mean ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001 (Student’s t test).
To understand how methionine and SAM triggered NR4A2 expression in ESCC cells, we hypothesized that these metabolites activated NR4A2 expression through METTL3-mediated mRNA m6A methylation. Indeed, treatment of ESCC cells with a global methylation inhibitor, 3-deazaadenosine (DAA), substantially declined methionine and SAM-induced NR4A2 expression at both the mRNA and protein levels (Fig. 5C, D, Supplementary Fig. S6B, C). Moreover, deletion of METTL3 in ESCC cells dramatically restrained methionine and SAM-elicited global RNA m6A methylation (Fig. 3G, H and Supplementary Fig. S6D, E) and attenuated NR4A2 expression at both the mRNA and protein levels (Fig. 5E). Additionally, treatment with a METTL3 inhibitor STM2457 also downregulated NR4A2 expression (Supplementary Fig. S6F). These findings demonstrated that methionine and SAM upregulated NR4A2 expression at the RNA level through METTL3-mediated m6A methylation.
Previous study has shown that m6A modification can promote mRNA stability and storage, thereby increasing gene expression output [31]. In line with this, treatment with the transcription inhibitor actinomycin D revealed that methionine administration overtly retarded NR4A2 mRNA decay (Fig. 5F). Conversely, abrogation of METTL3 remarkably expedited the degradation of NR4A2 mRNA induced by methionine in the presence of actinomycin D (Fig. 5G). Collectively, these findings indicated that methionine-induced NR4A2 expression in ESCC cells was mediated by METTL3-m6A-triggered mRNA stabilization.
Next, we explored which reader protein was responsible for interpreting the m6A modification in NR4A2 mRNA. Insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs), including IGF2BP1/2/3, are well-established m6A readers that recognize the consensus GG(m6A)C sequence, primarily located near stop codons and in 3′-UTRs, thereby promoting the stability and storage of target mRNA transcripts [31]. Given one of the four m6A peaks in NR4A2 mRNA induced by methionine was located in the 3′-UTR, we speculated that IGF2BPs would recognize this m6A peak and promote mRNA stability. To investigate this, we performed RNA immunoprecipitation-quantitative polymerase chain reaction (RIP-qPCR) analysis and found that IGF2BP2 exhibited the highest affinity for NR4A2 mRNA in KYSE150 cells (Fig. 5H). We hypothesized that IGF2BP2 recognized the m6A sites in NR4A2 mRNA and stabilized it. Indeed, deletion of IGF2BP2 remarkably accelerated NR4A2 mRNA decay and reduced the protein abundance of NR4A2 in ESCC cells (Fig. 5I, J). Thus, methionine could intensify the stability of NR4A2 mRNA and increase its expression output through a SAM-MELLT3-m6A-IGF2BP2 cascade.
Finally, we aimed to determine which m6A in NR4A2 mRNA was required for methionine-induced mRNA stabilization. Based on the MeRIP-seq data, we predicted three m6A consensus sequences within NR4A2 mRNA with high confidence, located in exon 6, exon 8, and 3′-UTR, respectively (Fig. 5K). Consequently, we constructed two mutants containing the full-length coding sequence of NR4A2 with A-T substitutions in m6A consensus sequences in exon 6 or exon 8 (Fig. 5K). In the presence of methionine, these two mutants did not alter NR4A2 mRNA decay rate compared to the wild-type NR4A2, as shown by qRT-PCR results (Fig. 5L). Furthermore, we generated reporter minigenes containing wild-type or mutant 3′-UTR fragments of NR4A2 (Fig. 5K and Supplementary Fig. S6G). Mutation of the m6A consensus sequence in the 3′-UTR led to a 25.9% decrease in methionine-induced NR4A2 expression compared to the wild-type 3′-UTR fragment of NR4A2 (Fig. 5M). Together, these results demonstrated that m6A modification in the 3′-UTR of NR4A2 was required for the methionine-elicited stabilization of NR4A2 mRNA.
NR4A2 facilitates ESCC growth both in vitro and in vivo
After identifying NR4A2 as a methionine-responsive gene, we studied its biological roles. We observed that NR4A2 deletion hindered ESCC cell proliferation, while enforced NR4A2 expression augmented ESCC cell multiplication (Fig. 6A–D). Furthermore, NR4A2 ablation led to cell cycle arrest at G2/M phase (Fig. 6E, F), indicating that NR4A2 was vital for ESCC cell propagation in vitro.
A Western blot assay confirming NR4A2 deletion in ESCC cells using the CRISPR-Cas9 approach and the influence of NR4A2 ablation on PCNA expression. B Western blot assay confirming NR4A2 overexpression of in ESCC cells and the effect of enforced NR4A2 expression on PCNA levels. Cell proliferation curves over time showing the impact of NR4A2 deletion (C) and NR4A2 overexpression (D) on ESCC cell propagation in complete medium. P values at each time point were calculated by comparing with the NC cells or cells transfected with blank vector. E, F Flow cytometric analysis revealing G2/M phase arrest of ESCC cells with NR4A2 deletion. Cells were cultured in complete medium. Three biological replicates were performed. G–I Influence of NR4A2 abrogation on ESCC xenograft tumor growth under conditions of high and low dietary methionine. Tumor growth curves, tumor image, and statistical analysis of tumor weight are presented in G, H, and I, respectively. J Western blot analysis revealing the effect of NR4A2 knockout on intratumoral expression of PCNA and cyclin B1 under conditions of high and low dietary methionine. NC non-target control. Error bars represent mean ± SEM. *p < 0.05, ***p < 0.001 (Student’s t test).
We utilized a xenograft tumor mouse model to verify the oncogenic role of NR4A2 in ESCC. High dietary methionine dramatically expedited the tumor growth of implanted control ESCC cells but not ESCC cells lacking NR4A2 (Fig. 6G–I). Consistently, in the presence of high dietary methionine, intratumoral expression of PCNA and Cyclin B1 was remarkably repressed by NR4A2 deletion (Fig. 6J). Additionally, under low dietary methionine conditions, NR4A2 abrogation did not further impede ESCC tumor growth and decline intratumoral expression of PCNA and Cyclin B1 (Fig. 6I–J). These results demonstrated that NR4A2 expression was crucial for methionine-induced ESCC tumor amplification in vivo. Notably, NR4A2 abrogation generated an anti-ECCC effect in the presence of high dietary methionine, comparable to the effect of a low methionine diet (Fig. 6I), indicating that NR4A2 inhibition could serve as a promising alternative to MR.
NR4A2 is highly expressed in ESCC and reversely linked to patient prognosis
Finally, we examined the expression of NR4A2 in clinical ESCC tissues and assessed its clinical significance. First, qRT-PCR analysis of tissue samples from validation cohort 2 revealed significantly elevated NR4A2 mRNA levels in ESCC tissues compared to paired NATs (Fig. 7A). Second, IHC analysis of tissue samples from validation cohorts 1, 2, and 3 showed a substantial increase in NR4A2 protein abundance in ESCC tissues as relative to NATs (Fig. 7B, C, Supplementary Fig. S7A, B). Third, western blot analysis of tissue samples from validation cohort 2 demonstrated a marked upregulation of NR4A2 protein content in ESCC tissues compared to paired NATs (Supplementary Fig. S7C, D). Collectively, these results indicated that NR4A2 expression was elevated in clinical ESCC tissues at both the mRNA and protein levels.
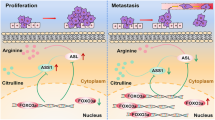
A qRT-PCR examination comparing NR4A2 mRNA levels between clinical ESCC tissues and paired NATs derived from validation cohort 2 (n = 48). P values were computed using the paired Wilcoxon rank-sum test. IHC staining analysis comparing NR4A2 protein levels between clinical ESCC tissues and NATs derived from validation cohort 1 (B) and validation cohort 2 (C). Representative IHC images and statistical violin-box-scatter plots are shown. P values were calculated using the Wilcoxon rank-sum test. OS (D) and DFS (E) curves of patients with high and low NR4A2 in validation cohort 1. The 70th percentile of NR4A2 IHC scores is used as the cut-off value. The 95% confidence limits of each survival curve are displayed by shadows. F–M Correlation analysis between NR4A2 and SLC7A5, MAT2A, MAT2B, or METTL3 in clinical ESCC tissues derived from validation cohort 1 (F–I) or combined validation cohort 2 and validation cohort 3 (J–M). Correlation coefficient (r) values and statistical p values were acquired by Spearman correlation analysis. Confidential intervals of regression lines fitted by a linear model are shown by shadows. N A new tumor-promoting mechanism of methionine in ESCC unveiled by this study. When compared to low methionine, high methionine stimulated methionine cycle and accelerated intracellular SAM production to intensify m6A methylation in NR4A2 mRNA in a METTL3-dependent manner. Next, a reader protein IGF2BP2 recognized a specific m6A site in 3′-UTR of NR4A2 mRNA, promoting its stability and storage. Consequently, this resulted in an increase in NR4A2 expression, therefore boosting ESCC growth.
Prognostic analysis using IHC data from validation cohort 1 revealed that high NR4A2 expression in ESCC tissues predicted unfavorable OS and DFS of patients (Fig. 7D, E). As depicted above, increased activity of methionine cycle predicted inferior prognosis in patients with ESCC (Fig. 1E, F). These findings suggested the potential involvement of the methionine cycle-NR4A2 axis in the disease progression of ESCC patients.
Furthermore, the IHC staining data of clinical ESCC tissues from validation cohorts 1, 2, and 3 showed an intimate and positive correlation between the IHC scores of NR4A2 and those of SLC7A5, MAT2A, MAT2B, and METTL3, respectively (Fig. 7F–M). This underscored the existence of a methionine cycle-METTL3-mRNA m6A-NR4A2 cascade in ESCC.
In summary, the current study presented a new tumor-promoting mechanism of methionine in ESCC (Fig. 7N). When compared to low methionine, high methionine provoked methionine cycle and expedited intracellular SAM generation in ESCC cells. SAM supplied active methyl group and enhanced m6A methylation in NR4A2 mRNA, primarily in a METTL3-dependent manner. Subsequently, a reader protein IGF2BP2 bound to a specific m6A site in 3′-UTR of NR4A2 mRNA, promoting its stability and storage. Ultimately, this led to a significant increase in gene expression output of NR4A2, thereby facilitating ESCC growth.
Celecoxib is a NR4A2 inhibitor with evident anti-ESCC efficacy
Previous studies find that the transcription of NR4A2 can be inhibited by cyclooxygenase-2 (COX-2) inhibitors, including parecoxib and celecoxib [45, 46]. Thus, we performed a screen using parecoxib, rofecoxib, and celecoxib, and found that only celecoxib obviously downregulated the protein abundance of NR4A2 in ESCC cells in a dose-dependent manner (Fig. 8A). However, celecoxib treatment did not perturb NR4A2 transcription in ESCC cells (Fig. 8B). Of note, in the presence of cycloheximide (CHX), an agent blocking the elongation phase of eukaryotic protein translation, supplement of celecoxib dramatically accelerated the degradation of NR4A2 (Fig. 8C). These findings suggested that celecoxib could enhance the protein stability of NR4A2. Subsequently, we ascertained whether this compound could bind to NR4A2 to exert its function. First, molecular docking analysis revealed that celecoxib could bind to NR4A2 with a binding energy value of −5.61 kcal/mol (Fig. 8D). Second, cellular thermal shift assay (CETSA) demonstrated that celecoxib supplement caused a remarkable thermal shift for NR4A2 protein (Fig. 8E), further indicating a direct interaction between this compound and NR4A2 protein.
A Influence of well-known COX-2 inhibitors at distinct concentrations on intracellular NR4A2 protein abundance of ESCC cells. B Effect of celecoxib at different concentrations on intracellular NR4A2 mRNA levels of ESCC cells. C Influence of celecoxib at 40 μM on intracellular NR4A2 protein abundance of ESCC cells over time under the condition of CHX treatment. The curves on the right side of the western blot images indicate the quantification of protein levels. D Celecoxib (green) interacting with NR4A2 motifs in the docking model. The hydrogen bond between this compound and NR4A2 was highlighted by red dashed line. E Western blot assays showing intracellular NR4A2 protein abundance of ESCC cells treated with or without celecoxib in the CETSA experiments. The curves on the right side of the western blot images indicate the quantification of protein levels. F Impact of NR4A2 deletion or NR4A2 overexpression on the sensitivity of KYSE150 cells to celecoxib treatment. G–I Influence of celecoxib and CDDP on KYSE150 xenograft tumor growth under high dietary methionine condition. Tumor growth curves, tumor image, and statistical analysis of tumor weight are presented in G, H, and I, respectively. J Western blot analysis displaying the influence of celecoxib administration on intratumoral NR4A2 expression under high dietary methionine condition. Error bars represent mean ± SEM. *p < 0.05, ***p < 0.001 (Student’s t test).
Subsequently, we investigated whether celecoxib exerted anti-ESCC efficacy in a NR4A2-dependent manner. First, in vitro surveys showed that NR4A2 deletion declined the sensitivity of KYSE150 cells to celecoxib, whereas ectopic NR4A2 expression substantially augmented the sensitivity of KYSE150 cells to this compound (Fig. 8F). Second, in vivo study demonstrated that oral administration of celecoxib dramatically suppressed the growth of subcutaneous KYSE150 xenograft tumors and downregulated intratumoral NR4A2 levels under high dietary methionine condition (Fig. 8G–J). Of note, celecoxib administration in vivo yielded an overt anti-ESCC effect in the presence of high dietary methionine, comparable to the effect of a low methionine diet or a well-known chemotherapeutic agent cisplatin (CDDP) under high dietary methionine condition (Fig. 8I).
Finally, we analyzed the safety of celecoxib administration in mice harboring KYSE150 xenograft tumors. When compared to high methionine diet, low methionine diet caused a significant reduction in the weight of the bodies, kidneys, and livers, but not the spleens (Supplementary Fig. S8). Similarly, under high dietary methionine condition, cisplatin (CDDP) administration evidently reduced the weight of the bodies, kidneys, livers, and spleens (Supplementary Fig. S8). However, in the presence of high dietary methionine, celecoxib administration only downregulated the weight of the livers and spleens, but not the bodies and kidneys (Supplementary Fig. S8). Notably, in the presence of high dietary methionine, there was a remarkable reduction in the liver weight in CDDP group as relative to celecoxib group (Supplementary Fig. S8C). Collectively, these results demonstrated that the safety of celecoxib administration in vivo was superior to that of a MR diet or CDDP treatment.
Discussion
Altered metabolism is a core hallmark of cancer and plays a direct role in cancer initiation and progression [6, 47]. Recent studies have reported the impact of diet on cancer growth and progression by modifying cancer cell metabolism [48,49,50]. Among these, methionine, an essential amino acid for humans, has been found to be pivotal for the expansion of tumor-initiating cells in lung cancer [19]. Additionally, dietary methionine can impinge upon the sensitivity of certain cancer cells, including colorectal cancer and soft-tissue sarcoma cells, to chemotherapy and radiotherapy [25]. In this study, we conducted a tissue metabolic survey in four patient cohorts, providing strong evidence of the aberrantly provoked methionine cycle in clinical ESCC tissues. Furthermore, we observed an inverse correlation between the activity of this metabolic pathway and patient survival. Importantly, our study demonstrated that dietary methionine was readily utilized by ESCC to generate SAM, thereby facilitating ESCC growth. These findings contribute to the growing body of evidence supporting methionine as a tumor-promoting metabolite in ESCC.
As a well-studied epigenetic event, m6A modification in mRNA plays a key role in posttranscriptional gene expression regulation. While it is widely accepted that methionine is readily absorbed by various cancer cells to generate the universal methyl donor SAM [17], whether and how methionine alters the RNA m6A methylation pattern of ESCC cells remains unclear. In our study, we demonstrated a remarkable upregulation of m6A abundance in RNA samples from clinical ESCC tissues via a methionine-SAM-METTL3 cascade. RNA-seq and MeRIP-seq assays revealed substantial changes in gene expression and mRNA m6A methylation signatures upon methionine treatment in ESCC cells. Importantly, we identified NR4A2 as a prominent downstream gene that was positively regulated by methionine through a SAM-METTL3-RNA m6A-IGF2BP2 cascade. Notably, we demonstrate that methionine strongly elicits NR4A2 expression in ESCC cells, but shows negligible influence on NR4A2 expression in non-transformed esophageal epithelial cells, potentially suggesting that this amino acid does not involve in the tumorigenesis of ESCC, consistent with the previous finding in a nutritional epidemiology study [12]. Furthermore, we found that m6A modification in the 3′-UTR of NR4A2 is important for IGF2BP2-induced stability of NR4A2 mRNA, accounting for approximately 25.9% of its stability. Therefore, further investigation is needed to ascertain other m6A sites contributing to IGF2BP2-mediated stability of NR4A2 mRNA. In conclusion, methionine addiction exerts a profound influence on the molecular traits of ESCC cells through intricate epitranscriptional regulation.
NR4A2 is a member of nuclear receptor subfamily 4A and its role in solid tumors is still a subject of debate [51]. For instance, NR4A2 exhibits oncogenic functions in cervical, prostate, and colon cancers by promoting malignant transformation and refraining intrinsic apoptosis [52, 53], while acting as a tumor suppressor in bladder cancer by eliciting cell apoptosis [54]. In our study, we provide evidence supporting NR4A2 as an oncogene in ESCC, as it boosts cell growth in the presence of methionine and negatively affects patient survival. Considering the adverse effect of a MR diet, such as severe body weight loss observed in this study, NR4A2, a newly identified methionine-responsive oncogene with specifically high expression in clinical ESCC tissues, represents a compelling target for ESCC treatment in the presence of a normal diet. Indeed, celecoxib, an inhibitor targeting NR4A2, shows overt anti-ESCC growth efficacy and safety in the presence of a methionine-rich diet.
In the end, several important issues raised by our study. Firstly, the impact of methionine/SAM on the methylation patterns of various intracellular biological macromolecules demands thorough exploration. SAM, as the sole source of the universal methyl donor, serves as an indispensable substrate in all methylation reactions, encompassing the methylation of RNA, DNA, histones, and non-histone proteins [18]. Secondly, the influence of methionine/SAM on the m6A modification of other RNAs necessitates comprehensive assessment. Previous studies report that m6A modification is prevalent across various RNAs, encompassing mRNAs, transfer RNAs, ribosomal RNAs, circular RNAs, micro RNAs, and long non-coding RNAs [29, 55]. Thirdly, the mechanisms by which methionine determines the unique mRNA m6A methylation pattern in ESCC cells need to be further explored. While METTL3 has been shown to intensify m6A modification in adenomatous polyposis coli (APC) mRNA in ESCC cells [41], methionine-induced mRNA transcripts with prominent m6A peaks do not include APC (Supplementary Table S6). Therefore, the underlying mechanism by which methionine triggers a distinct mRNA m6A signature in ESCC requires further investigation. Fourthly, the mechanisms through which methionine regulates gene expression in ESCC should be thoroughly addressed. Although Fig. 4 shows upregulation of GAS7 and downregulation of HOGA1 in the presence of methionine, no m6A peaks were identified in the mRNAs of these genes. Hence, there must be other unknown mechanisms by which methionine modulates the expression of these genes. Fifthly, it is important to carefully determine whether NR4A2 inhibition is a viable alternative to MR for ESCC therapy. We demonstrate that, in a subcutaneous xenograft mouse model, NR4A2 deletion leads to a significant anti-ESCC effect in the presence of a methionine-rich diet, similar to the effect of a MR diet. In this mouse model, oral celecoxib administration also yields an evident anti-ESCC effect under high dietary methionine condition, comparable to the effect of a MR diet or CDDP under high dietary methionine condition. Further animal models are necessary to confirm whether NR4A2 inhibition, as a new approach that can bypass the adverse effect of MR, can provide considerable therapeutic benefit for ESCC. Sixthly, the tumor-promoting mechanism of NR4A2 in ESCC warrants urgent investigation, given its paradoxical roles in cancer as reported in previous studies. Ultimately, it is imperative to unravel the impact of other methionine-induced genes, such as ALDH3B1, ANKRD37, GAS7, SOX9, and DUSP5, on the phenotypic characteristics of ESCC cells in future investigations. Notably, ALDH3B1 is found to be important for glioma cell proliferation [56]. The heightened expression of SOX9 in clinical cancer tissues promotes neoplastic growth of cancer cells [57]. Furthermore, DUSP5, known for its overexpression in human thyroid carcinoma (TC) tissues, plays a pivotal role in fostering TC cell metastasis and anchorage-independent growth [58]. Given these findings, it is imperative to access the involvement of these methionine-responsive genes in the progression of ESCC, thereby advancing our understanding of ESCC pathogenesis.
Materials and methods
All materials and methods are available in the Supplementary Information file. All the full and uncropped WB images in this paper are uploaded as Supplementary Information.
Data availability
All datasets are available from the corresponding author on reasonable request.
References
Wang ZX, Cui C, Yao J, Zhang Y, Li M, Feng J, et al. Toripalimab plus chemotherapy in treatment-naive, advanced esophageal squamous cell carcinoma (JUPITER-06): A multi-center phase 3 trial. Cancer Cell. 2022;40:277–88 e273.
Chen WL, Jin X, Wang M, Liu D, Luo Q, Tian H, et al. GLUT5-mediated fructose utilization drives lung cancer growth by stimulating fatty acid synthesis and AMPK/mTORC1 signaling. JCI Insight. 2020;5:e131596.
Chen WL, Wang JH, Zhao AH, Xu X, Wang YH, Chen TL, et al. A distinct glucose metabolism signature of acute myeloid leukemia with prognostic value. Blood. 2014;124:1645–54.
Chen WL, Wang YY, Zhao A, Xia L, Xie G, Su M, et al. Enhanced Fructose Utilization Mediated by SLC2A5 Is a Unique Metabolic Feature of Acute Myeloid Leukemia with Therapeutic Potential. Cancer Cell. 2016;30:779–91.
DeBerardinis RJ, Keshari KR. Metabolic analysis as a driver for discovery, diagnosis, and therapy. Cell. 2022;185:2678–89.
Vander Heiden MG, DeBerardinis RJ. Understanding the Intersections between Metabolism and Cancer Biology. Cell. 2017;168:657–69.
Sun C, Li T, Song X, Huang L, Zang Q, Xu J, et al. Spatially resolved metabolomics to discover tumor-associated metabolic alterations. Proc Natl Acad Sci USA. 2019;116:52–57.
Yang T, Hui R, Nouws J, Sauler M, Zeng T, Wu Q. Untargeted metabolomics analysis of esophageal squamous cell cancer progression. J Transl Med. 2022;20:127.
Zhang H, Wang L, Hou Z, Ma H, Mamtimin B, Hasim A, et al. Metabolomic profiling reveals potential biomarkers in esophageal cancer progression using liquid chromatography-mass spectrometry platform. Biochem Biophys Res Commun. 2017;491:119–25.
Jin X, Liu L, Wu J, Jin X, Yu G, Jia L, et al. A multi-omics study delineates new molecular features and therapeutic targets for esophageal squamous cell carcinoma. Clin Transl Med. 2021;11:e538.
Liu L, Wu J, Shi M, Wang F, Lu H, Liu J, et al. New Metabolic Alterations and A Predictive Marker Pipecolic Acid in Sera for Esophageal Squamous Cell Carcinoma. Genomics Proteomics Bioinformatics. 2022;20:670–87.
Xiao Q, Freedman ND, Ren J, Hollenbeck AR, Abnet CC, Park Y. Intakes of folate, methionine, vitamin B6, and vitamin B12 with risk of esophageal and gastric cancer in a large cohort study. Br J Cancer. 2014;110:1328–33.
Glaudemans AW, Enting RH, Heesters MA, Dierckx RA, van Rheenen RW, Walenkamp AM, et al. Value of 11C-methionine PET in imaging brain tumours and metastases. Eur J Nucl Med Mol Imaging. 2013;40:615–35.
Kim S, Chung JK, Im SH, Jeong JM, Lee DS, Kim DG, et al. 11C-methionine PET as a prognostic marker in patients with glioma: comparison with 18F-FDG PET. Eur J Nucl Med Mol Imaging. 2005;32:52–59.
Luckerath K, Lapa C, Albert C, Herrmann K, Jorg G, Samnick S, et al. 11C-Methionine-PET: a novel and sensitive tool for monitoring of early response to treatment in multiple myeloma. Oncotarget. 2015;6:8418–29.
Cavuoto P, Fenech MF. A review of methionine dependency and the role of methionine restriction in cancer growth control and life-span extension. Cancer Treat Rev. 2012;38:726–36.
Kaiser P. Methionine Dependence of Cancer. Biomolecules. 2020;10:568.
Sanderson SM, Gao X, Dai Z, Locasale JW. Methionine metabolism in health and cancer: a nexus of diet and precision medicine. Nat Rev Cancer. 2019;19:625–37.
Wang Z, Yip LY, Lee JHJ, Wu Z, Chew HY, Chong PKW, et al. Methionine is a metabolic dependency of tumor-initiating cells. Nat Med. 2019;25:825–37.
Lamers Y. Indicators and methods for folate, vitamin B-12, and vitamin B-6 status assessment in humans. Curr Opin Clin Nutr Metab Care. 2011;14:445–54.
Fukumoto K, Ito K, Saer B, Taylor G, Ye S, Yamano M, et al. Excess S-adenosylmethionine inhibits methylation via catabolism to adenine. Commun Biol. 2022;5:313.
Liu L, Amorin R, Moriel P, DiLorenzo N, Lancaster PA, Penagaricano F. Maternal methionine supplementation during gestation alters alternative splicing and DNA methylation in bovine skeletal muscle. BMC Genomics. 2021;22:780.
Goseki N, Yamazaki S, Shimojyu K, Kando F, Maruyama M, Endo M, et al. Synergistic effect of methionine-depleting total parenteral nutrition with 5-fluorouracil on human gastric cancer: a randomized, prospective clinical trial. Jpn J Cancer Res. 1995;86:484–9.
Hoffman RM. Clinical Studies of Methionine-Restricted Diets for Cancer Patients. Methods Mol Biol. 2019;1866:95–105.
Gao X, Sanderson SM, Dai Z, Reid MA, Cooper DE, Lu M, et al. Dietary methionine influences therapy in mouse cancer models and alters human metabolism. Nature. 2019;572:397–401.
Shea TB, Rogers E. Lifetime requirement of the methionine cycle for neuronal development and maintenance. Curr Opin Psychiatry. 2014;27:138–42.
Bokar JA, Shambaugh ME, Polayes D, Matera AG, Rottman FM. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA. 1997;3:1233–47.
Lan Q, Liu PY, Haase J, Bell JL, Huttelmaier S, Liu T. The Critical Role of RNA m(6)A Methylation in Cancer. Cancer Res. 2019;79:1285–92.
Fu Y, Dominissini D, Rechavi G, He C. Gene expression regulation mediated through reversible m(6)A RNA methylation. Nat Rev Genet. 2014;15:293–306.
Huang L, Wang C, Xu H, Peng G. Targeting citrate as a novel therapeutic strategy in cancer treatment. Biochim Biophys Acta Rev Cancer. 2020;1873:188332.
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–95.
Zhang C, Fu J, Zhou Y. A Review in Research Progress Concerning m6A Methylation and Immunoregulation. Front Immunol. 2019;10:922.
Chen M, Wei L, Law CT, Tsang FH, Shen J, Cheng CL, et al. RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology. 2018;67:2254–70.
Chen XY, Zhang J, Zhu JS. The role of m(6)A RNA methylation in human cancer. Mol Cancer. 2019;18:103.
Sun T, Wu R, Ming L. The role of m6A RNA methylation in cancer. Biomed Pharmacother. 2019;112:108613.
Xia J, Wishart DS. MSEA: a web-based tool to identify biologically meaningful patterns in quantitative metabolomic data. Nucleic Acids Res. 2010;38:W71–77.
Chong J, Soufan O, Li C, Caraus I, Li S, Bourque G, et al. MetaboAnalyst 4.0: towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018;46:W486–W494.
Su H, Hu N, Yang HH, Wang C, Takikita M, Wang QH, et al. Global gene expression profiling and validation in esophageal squamous cell carcinoma and its association with clinical phenotypes. Clin Cancer Res. 2011;17:2955–66.
Mentch SJ, Mehrmohamadi M, Huang L, Liu X, Gupta D, Mattocks D, et al. Histone Methylation Dynamics and Gene Regulation Occur through the Sensing of One-Carbon Metabolism. Cell Metab. 2015;22:861–73.
Rohm M, Schafer M, Laurent V, Ustunel BE, Niopek K, Algire C, et al. An AMP-activated protein kinase-stabilizing peptide ameliorates adipose tissue wasting in cancer cachexia in mice. Nat Med. 2016;22:1120–30.
Wang W, Shao F, Yang X, Wang J, Zhu R, Yang Y, et al. METTL3 promotes tumour development by decreasing APC expression mediated by APC mRNA N(6)-methyladenosine-dependent YTHDF binding. Nat Commun. 2021;12:3803.
Yankova E, Blackaby W, Albertella M, Rak J, De Braekeleer E, Tsagkogeorga G, et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature. 2021;593:597–601.
Kim J, Lee G. Metabolic Control of m(6)A RNA Modification. Metabolites. 2021;11:80.
Wang Q, Chen C, Ding Q, Zhao Y, Wang Z, Chen J, et al. METTL3-mediated m(6)A modification of HDGF mRNA promotes gastric cancer progression and has prognostic significance. Gut. 2020;69:1193–205.
Zagani R, Hamzaoui N, Cacheux W, de Reynies A, Terris B, Chaussade S, et al. Cyclooxygenase-2 inhibitors down-regulate osteopontin and Nr4A2-new therapeutic targets for colorectal cancers. Gastroenterology. 2009;137:1358–66. e1351-1353
Hibino S, Chikuma S, Kondo T, Ito M, Nakatsukasa H, Omata-Mise S, et al. Inhibition of Nr4a Receptors Enhances Antitumor Immunity by Breaking Treg-Mediated Immune Tolerance. Cancer Res. 2018;78:3027–40.
Hanahan D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022;12:31–46.
Bose S, Allen AE, Locasale JW. The Molecular Link from Diet to Cancer Cell Metabolism. Mol Cell. 2020;78:1034–44.
Lien EC, Vander Heiden MG. A framework for examining how diet impacts tumour metabolism. Nat Rev Cancer. 2019;19:651–61.
Locasale JW. Diet and Exercise in Cancer Metabolism. Cancer Discov. 2022;12:2249–57.
Safe S, Karki K. The Paradoxical Roles of Orphan Nuclear Receptor 4A (NR4A) in Cancer. Mol Cancer Res. 2021;19:180–91.
Holla VR, Mann JR, Shi Q, DuBois RN. Prostaglandin E2 regulates the nuclear receptor NR4A2 in colorectal cancer. J Biol Chem. 2006;281:2676–82.
Ke N, Claassen G, Yu DH, Albers A, Fan W, Tan P, et al. Nuclear hormone receptor NR4A2 is involved in cell transformation and apoptosis. Cancer Res. 2004;64:8208–12.
Inamoto T, Papineni S, Chintharlapalli S, Cho SD, Safe S, Kamat AM. 1,1-Bis(3′-indolyl)-1-(p-chlorophenyl)methane activates the orphan nuclear receptor Nurr1 and inhibits bladder cancer growth. Mol Cancer Ther. 2008;7:3825–33.
Jiang X, Liu B, Nie Z, Duan L, Xiong Q, Jin Z, et al. The role of m6A modification in the biological functions and diseases. Signal Transduct Target Ther. 2021;6:74.
Wang Z, Mo Y, Tan Y, Wen Z, Dai Z, Zhang H, et al. The ALDH Family Contributes to Immunocyte Infiltration, Proliferation and Epithelial-Mesenchymal Transformation in Glioma. Front Immunol. 2021;12:756606.
Aldaz P, Otaegi-Ugartemendia M, Saenz-Antonanzas A, Garcia-Puga M, Moreno-Valladares M, Flores JM, et al. SOX9 promotes tumor progression through the axis BMI1-p21(CIP). Sci Rep. 2020;10:357.
Buffet C, Hecale-Perlemoine K, Bricaire L, Dumont F, Baudry C, Tissier F, et al. DUSP5 and DUSP6, two ERK specific phosphatases, are markers of a higher MAPK signaling activation in BRAF mutated thyroid cancers. PLoS One. 2017;12:e0184861.
Funding
This work was supported by the National Key R&D Program of China (2023YFC3503200, 2023YFC3503201, 2022YFC3500200, 2022YFC3500202), National Natural Science Foundation of China (31970708, 82002953, 32170778, 82004177, 82072567), Shanghai Frontier Research Base of Disease and Syndrome Biology of Inflammatory cancer transformation (2021KJ03-12), National Scientific and Technological Major Special Project of China (2019ZX09201004-002-013), Tracking Program for Eastern Scholar at Shanghai Institutions of Higher Learning, Shanghai High-level Talent Leadership Program of Traditional Chinese Medicine [ZY(2021-2023)-0403], Scientific Research Project of Industry Development Center of Shanghai University of Traditional Chinese Medicine (602076D), the grant from Nantong Health Commission (MA2021024), Shanghai “Science and Technology Innovation Action Plan” Medical Innovation Research Project-Shanghai Clinical Research Center of Traditional Chinese Medicine Oncology (21MC1930500), Shanghai 13th Five-Year Plan Key Specialty of Traditional Chinese Medicine Oncology (shslczdzk03701), Three-year Action Plan for Shanghai TCM Development and Inheritance Program [ZY(2021-2023)-0401], and Health Commission of Pudong New Area Health and Family Planning Scientific Research Project (PW2019E-1).
Author information
Authors and Affiliations
Contributions
Conceptualization: W-LC; Studies in vitro and in vivo: XJ (Xin Jing) and DL. Patient enrollment, clinical samples, and clinical data: LL, JW, CW, SW, FY, XY, M-SW, Z-WW, and J-BL; Pathology, H&E, and IHC staining assays: XJ (Xing Jin), GY, XJ (Xiaoxia Jin), Y-WX, and DJ; Omics study and data analysis: XJ (Xing Jin), DL, YN, and W-LC; Bioinformatics and statistical analysis: XJ (Xing Jin), DL, and W-LC; Manuscript writing: XJ (Xin Jing) and W-LC; Writing - review and editing: W-LC, YO, WJ, and GM.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jin, X., Liu, L., Liu, D. et al. Unveiling the methionine cycle: a key metabolic signature and NR4A2 as a methionine-responsive oncogene in esophageal squamous cell carcinoma. Cell Death Differ (2024). https://doi.org/10.1038/s41418-024-01285-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41418-024-01285-7