Abstract
Pharmacological targeting via small molecule-based chemical probes has recently acquired an emerging importance as a valuable tool to delineate molecular mechanisms. Induction of apoptosis via CD95/Fas and TRAIL-R1/2 is triggered by the formation of the death-inducing signaling complex (DISC). Caspase-8 activation at the DISC is largely controlled by c-FLIP proteins. However molecular mechanisms of this control have just started to be uncovered. In this study we report the first-in-class chemical probe targeting c-FLIPL in the heterodimer caspase-8/c-FLIPL. This rationally designed small molecule was aimed to imitate the closed conformation of the caspase-8 L2′ loop and thereby increase caspase-8 activity after initial processing of the heterodimer. In accordance with in silico predictions, this small molecule enhanced caspase-8 activity at the DISC, CD95L/TRAIL-induced caspase activation, and subsequent apoptosis. The generated computational model provided further evidence for the proposed effects of the small molecule on the heterodimer caspase-8/c-FLIPL. In particular, the model has demonstrated that boosting caspase-8 activity by the small molecule at the early time points after DISC assembly is crucial for promoting apoptosis induction. Taken together, our study allowed to target the heterodimer caspase-8/c-FLIPL and get new insights into molecular mechanisms of its activation.
Similar content being viewed by others
Introduction
Apoptosis is the programme of cell death that is essential for all multicellular organisms. An apoptotic signal can be induced by a variety of factors including death receptor (DR) activation [1]. The apoptotic DR signaling cascade is triggered by the activation of a corresponding DR, in particular, CD95/Fas or TRAIL-R1/2 (DR4/DR5), which results in the formation of a death-inducing signaling complex (DISC). DISC comprises DR, FADD, procaspases-8, -10, and c-FLIP, serving a central platform for procaspase-8 activation, which initiates the subsequent apoptotic response [2]. The molecular architecture of the DISC is designated by a framework of strictly defined interactions between death domains and death effector domains (DEDs). In particular, recently it has been shown that procaspase-8 at the DISC forms so-called DED chains or filaments via DED interactions (that we term filaments thereafter), which serve as a platform for dimerization and subsequent activation of procaspase-8 [3,4,5].
DED filament assembly is crucial for procaspase-8a/b dimerization and subsequent activation [3, 6]. In the course of dimerization, procaspase-8 undergoes a conformational changes that trigger the rearrangement of the L2 loop in the zymogen structure [7]. The L2 loop contains the active cysteine and, accordingly, its rearrangement in the course of dimerization naturally forms the active center of procaspase-8a/b. This is followed by the cleavage of the L2 loop at Asp374 into the L2 (“processed” L2 fragment) and L2′ parts followed by their subsequent structural rearrangement. The cleavage of the L2 loop at Asp374 leads to the generation of p43/p41 (denoted thereafter as p43) and p12 cleavage products, which are further auto-catalytically processed via proteolysis at Asp384 and Asp210/216 resulting in the formation of the active caspase-8 heterotetramer p102/p182 [6, 8, 9].
Initiation of caspase-8 activation at the DISC and DED filaments are largely controlled by c-FLIP proteins [10]. Three c-FLIP isoforms including Long (L), Short (S), and Raji (R), i.e., c-FLIPL, c-FLIPS, and c-FLIPR and two cleavage products have been characterized so far [11,12,13]. All three isoforms possess two DED domains. c-FLIPL also contains catalytically inactive caspase-like domains (p20 and p12). The short c-FLIP isoforms, c-FLIPS and c-FLIPR, block DR-induced apoptosis by inhibiting procaspase-8 activation at the DISC upon their high expression [13,14,15]. Recently, it was reported that short c-FLIP isoforms block caspase-8 activation by interrupting the chains of procaspase-8 at the DISC [16]. In contrast, it was also reported that short c-FLIP isoforms incorporate into DED chains and block caspase-8 activation by forming inactive heterodimers [15]. c-FLIPL at the DISC can act both in a proapoptotic as well as in an antiapoptotic manner [16,17,18]. The proapoptotic action of c-FLIPL is found to take place upon its moderate levels of expression, while upon high expression it acts solely antiapoptotic [10, 14].
The proapoptotic function of c-FLIPL has been suggested to be mediated by the formation of procaspase-8/c-FLIPL heterodimers in which the L2 loop of procaspase-8 is stabilized in the active conformation through interactions with c-FLIPL, as this enhances the catalytic activity of the caspase-8 enzyme [7, 18,19,20]. In particular, a unique feature observed in the structure of procaspase-8/c-FLIPL heterodimer is a so-called “closed” conformation of the unprocessed L2′ fragment (denoted thereafter L2′ loop), which apparently stabilizes the active center of caspase-8 and thereby promotes a catalytic activity of the heterodimer [7]. The cleavage of the L2 loop of procaspase-8 at Asp374 was suggested to lead to the translocation of the L2′ loop from the heterodimer interface, which accordingly results in a change from a “closed” to an “open” conformation. This might lead to destabilization of the active center and subsequently a diminished activity of caspase-8.
To validate this hypothesis, we took advantage of the state-of–the-art methods of in silico biology and constructed a small molecule-based chemical tool that was designed to mimic the stabilizing effect of the L2′ loop in a “closed” conformation. In particular, we rationally designed a small molecule that binds to c-FLIPL at the interface of the heterodimer caspase-8/c-FLIPL, aiming at stabilization of the active center of caspase-8 in the caspase-8/c-FLIPL heterodimer after processing at Asp374. The optimized chemical compound enhanced caspase-8 activity at the DISC and promoted cell death. Moreover, generated computational model of the DISC further supported the suggested mechanism. Taken together, our study uncovers the potential of targeting the caspase-8/c-FLIPL heterodimer via structure-based design leading to the development of compounds that can serve as drugs promoting apoptosis in cancer cells.
Material and methods
Virtual Screening (VS)
VS was carried out using the Glide molecular docking software [21, 22] from the Schrödinger Small-Molecule Drug Discovery Suite 2015-1 (Schrödinger, Inc). Molecular docking was performed in the standard-precision (SP) and extra-precision (XP) modes. Prior to VS, protein structures were processed using the “protein preparation wizard module” in the Schrödinger Suite 2015-1 [23]. Protein minimization was carried out using “MacroModel” from the Schrödinger Suite 2015-1 and OPLS_2005 force field. Restrained energy minimization was done with the convergence of heavy atoms to RMSD value of 0.5 Å. The details of structural modeling in silico are given in Supplementary Material. VS was performed using the ZINC12 library of commercially available compounds prepared for molecular docking, which contains more than 16 million purchasable compounds [24]. At the first step VS was executed using Glide in the SP mode selecting about 100,000 compounds with the best Glide SP scoring function values (Supplementary Fig. 1a). The next round of VS was performed in the XP mode, resulting in a selection of compounds with the best Glide XP scoring function values for further visual inspection and selection of final hits for experimental validation (Supplementary Fig. 1a). In addition, we applied the same VS pipeline for 3770 active substances from the PubChem BioAssay database (AID 62435), which contains the results of high-throughput screening directed on identification of small molecule compounds sensitizing to TRAIL-induced apoptosis through caspase-8 pathway. This allowed us to select two more compounds, FLIPinQ and FLIPinR.
Cell lines
Human cervical cancer HeLa-CD95 cells (CD95-overexpressing cells) [25] and HeLa-CD95-FL cells (CD95/c-FLIPL-overexpressing cells) were maintained in DMEM/Ham’s F-12 media (Merck Millipore, Germany), supplemented with 10% heat-inactivated fetal calf serum, 1% penicillin-streptomycin, and 0.0001% puromycin in 5% CO2. Human acute myeloid leukemia MV4-11 cells and T leukemia Jurkat cells were maintained in RPMI 1640 (Thermo Fisher Scientific Inc., USA), supplemented with 10% heat-inactivated fetal calf serum and 1% penicillin-streptomycin in 5% CO2.
Antibodies and reagents
The following antibodies were used for western blot analysis: polyclonal anti-caspase-3 antibody (#9662), polyclonal anti-PARP antibody (#9542), monoclonal anti-RIPK1 XP antibody (#3493) from Cell Signaling Technology, USA; polyclonal anti-actin antibody (A2103) from Sigma-Aldrich, Germany; polyclonal anti-CD95 antibody (sc-715) from Santa Cruz, USA; polyclonal anti-mCherry antibody (ab183628) from Abcam, UK; monoclonal anti-caspase-10 antibody (M059-3) from MBL International Corporation, USA; monoclonal anti-FADD antibody (clone 1C4), monoclonal anti-caspase-8 antibody (clone C15), and monoclonal c-FLIP antibody (clone NF6). Horseradish peroxidase-conjugated goat anti-mouse IgG1, -2b, goat anti-rabbit and rabbit anti-goat were from Santa Cruz, USA. Recombinant TRAIL (KillerTRAIL™) was from Enzo Life Sciences, Germany. The monoclonal anti-APO-1 antibody (mouse-IgG3) was used for immunoprecipitations (IPs). All chemicals were of analytical grade and purchased from Merck (Germany) or Sigma (Germany). The anti-APO-1, C15, NF6, and 1C4 antibodies were kindly provided by Prof. P. H. Krammer (DKFZ, Heidelberg). Recombinant LZ-CD95L was produced as described [14].
Analysis of total cell lysates by western blot analysis
The western blot analysis of total cellular lysates was performed in accordance with our previous reports [26]. The quantification of processing of caspases and their substrates via semiquantitative western blot was performed using Image LabBeta 5.1 Software (BioRad). The normalization was performed to actin signal.
CD95-DISC-IP
The CD95-DISC-IP from 5 × 106 HeLa-CD95 cells were done as described before using anti-APO-1 antibodies [27]. In addition, DISC-IPs were washed four times with PBS, which was followed by western blot analysis or caspase-8 activity assays.
Caspase-8 activity assay
1,2 × 104 cells per well were seeded in a 96-well plate 1 day before experiment. Fifty microliters of the Caspase-Glo® 8 reagent together with MG-132 inhibitor was added to each well. The luminescence intensity was analyzed by a microplate reader Infinite M200pro (Tecan, Switzerland). The measurements were performed according to the manufacturer’s instructions (Caspase-Glo® 8 Assay, Promega, Germany). Every condition was performed in duplicate.
Measuring caspase-8 activity at the DISC
Each of the protein A bead samples with CD95-DISC-IPs was resuspended in 95 µL of CHAPS-Buffer (50 mM HEPES pH7.2; 50 mM NaCl; 10 mM EDTA; 5% glycerol; 10 mM DTT; 0.1% CHAPS) and transferred into a 96-well plate. Caspase-8 activity was measured according to the manufacturer’s instructions (Caspase-Glo® 8 Assay, Promega, Germany). Every condition was performed in duplicate. The luminescence intensity was analyzed by the microplate reader Infinite M200pro (Tecan, Switzerland).
Cell viability quantification by ATP assay
1,2 × 104 cells per well (for adherent cells) were seeded in a 96-well plate 1 day prior stimulation. 2 × 104 cells per well (for suspension cells) were seeded in a 96-well plate before experiment. Fifty microliters of the CellTiter-Glo® solution was added to each well. Measurements were performed according to the manufacturer’s instructions (CellTiter-Glo® Luminescent Cell Viability Assay, Promega, Germany). The luminescence intensity was analyzed by the microplate reader Infinite M200pro (Tecan, Switzerland). The ATP amount of nontreated cells was taken as one relative unit (RU). Every condition was performed in duplicate.
Caspase-3/7 activity assay
1,2 × 104 cells per well (for adherent cells) were seeded in a 96-well plate 1 day prior stimulation. 2 × 104 cells per well (for suspension cells) were seeded in a 96-well plate before experiment. Fifty microliters of the Caspase-Glo® 3/7 solution was added to each well. The luminescence intensity was analyzed by a microplate reader Infinite M200pro (Tecan, Switzerland) according to the manufacturer’s instructions (Caspase-Glo® 3/7 Assay, Promega, Germany). The caspase activity of nontreated cells was taken as one RU. Every condition was performed in duplicate.
IncuCyteTM live cell imaging
HeLa-CD95 cells in a 96-well plate (5 × 103 cells/well) seeded 1 day before stimulation were preincubated with FLIPinBγ for 2 h. Subsequently, cells were stimulated with CD95L (50 ng/mL). To monitor the loss of the cell membrane integrity, cells were treated with IncuCyte™ Cytotox Red Reagent (1:4000) [28]. To monitor caspase activity, the cells were treated with IncuCyte™ Caspase-3/7 Green Apoptosis Assay Reagent (1:5000). The cells were imaged every hour by IncuCyte® ZOOM (Essen BioScience, USA) with a 10× objective for a period of 6 h. Every condition was performed in triplicate. Data were analyzed by the IncuCyte® ZOOM Software (2016A).
Cell death analysis using imaging flow cytometry
5 × 105 cells per well were seeded 1 day before stimulation in a 6-well plate. Afterward they were treated with or without 30 µM FLIPinBγ for 2 h, which was followed by stimulation with 1667 ng/mL CD95L for 4 h. Quantification of early and late apoptotic cells was performed using Annexin-V-FITC/SytoxOrange staining and imaging flow cytometry via FlowSight® (AMNIS®, Luminex) with subsequent analysis as described previously [29].
Statistical analysis
Graphpad Prism 8 software was used to generate and perform one-way ANOVA tests (paired, normal distribution, multiple comparison test: Dunnett’s test). T-tests were performed for statistical comparison of two different conditions within one analysis (paired, parametric, two-tailed) using Graphpad Prism 8 software or using Scipy python library. Box plots were plotted using the Python matplotlib and seaborn libraries.
Modeling
Ordinary differential equations (ODEs) were used to model caspase-8 activation at the DISC with or without FLIPinBγ. A detailed description of modeling procedures including parameter fitting is given in Supplementary Material. In brief, the model was fitted to the data on caspase-3 activity in the cells and in vitro caspase-8 activity at the DISC. In vitro caspase-8 activity was estimated from the rate of caspase-8 substrate cleavage, assuming that the caspase-8 substrate is present in excess. Identifiability analysis of parameters was carried out by implementing the profile likelihood method [30]. Profile likelihoods for each parameter were evaluated by fixing corresponding parameters in a wide range of values, while reoptimizing all other parameters. The 95% finite sample confidence intervals for the parameters were estimated.
Results
VS identified the compounds targeting c-FLIP (FLIPins)
A “closed” conformation of the unprocessed L2′ loop of procaspase-8 in the procaspase-8/c-FLIPL heterodimer was suggested to be crucial for its proapoptotic function, due to its stabilizing effect on the active center of caspase-8 (Fig. 1a). In the “closed” conformation, the side chains of amino acid residues of the unprocessed L2′ loop occupy a groove on the c-FLIPL protein located between the β6 and α3 regions (β6/α3 groove) [7]. The switch of the L2′ loop from a “closed” to an “open” conformation due to the L2 loop cleavage is expected to abolish the interactions within the β6/α3 groove of c-FLIPL and subsequently diminish the activity of the heterodimer (Fig. 1b). In this regard, the L2 loop cleavage leads to the generation of caspase-8-p43 and p10 cleavage products, resulting in a complex comprising caspase-8-p43/p10 and c-FLIPL (denoted thereafter as p43/p10/c-FLIPL) (Fig. 1b). Accordingly, we assumed that if the β6/α3 groove will be occupied with a rationally designed small molecule mimicking the stabilizing effect of the L2′ loop in a “closed” conformation, it would rescue the activity of the p43/p10/c-FLIPL complex (Fig. 1b). In order to identify such compounds, a VS was performed (Supplementary Fig. 1a).
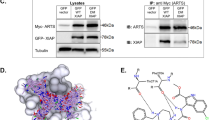
a Model of heterodimer structure of procaspase-8 (shown in dark blue) and c-FLIPL (shown in light blue) [PDB ID 3H11]. L2′ loop in the “closed” conformation is indicated. Amino acid residues of L2′ loop are shown in gold color. L2 loop and cleavage sites are indicated with dashed lines. b Model of heterodimer structure of c-FLIPL [PDB ID 3H11] and processed caspase-8-p43/p10 [PDB ID 4PRZ] with a FLIPinBγ (shown in green) bound to the putative small molecule binding site. The molecular surface of c-FLIPL and caspase-8-p43/p10 binding site regions is shown in brown and blue colors, respectively. Amino acid residues of L2′ loop in “open” conformation are shown in gold color. c Predicted interactions of FLIPinBγ with binding site residues of the heterodimer complex; amino acid residues of c-FLIPL and caspase-8 are denoted in brown and blue colors, respectively. The water molecule from the structure [PDB ID 3H11] at the FLIPinBγ binding interface is indicated. Residues potentially involved in allosteric stabilization of caspase-8 L2 loop are highlighted with red color. Active site C360 is shown in CPK representation.
To carry out the VS, the structure of the p43/p10/c-FLIPL complex with the L2′ loop in an “open” conformation had to be computed (Supplementary Material). The structure of the p43/p10/c-FLIPL complex was derived from two previously obtained structures: the crystal structure of procaspase-8/c-FLIPL heterodimer of [PDB ID 3H11] [7] and the structure of the p18/p10 subunits of caspase-8 [PDB ID 4PRZ] [31]. The generated 3D structure of the p43/p10/c-FLIPL complex was used for VS (Fig. 1b).
Structural rearrangements in the course of the transformation from the “closed” to the “open” conformation of the L2′ loop might lead to formation of a cavity on the binding interface of p43/p10/c-FLIPL complex, according to in silico model (Fig. 1b). Therefore, we expected that VS would allow to identify compounds binding within this cavity to c-FLIPL, thereby imitating the stabilizing effect of the L2′ loop in a “closed” conformation and leading to an increase in caspase-8 activity (Supplementary Fig. 1a). As a result of our VS, 5000 compounds with the best scoring function values were selected for further visual inspection (Supplementary Fig. 1a). Visual verification was aimed at filtering of possible docking artifacts, as well as at selection of compounds able to mimic key interactions observed in the L2′ loop in the procaspase-8/c-FLIPL heterodimer structure (Supplementary Fig. 1b, c). Finally, we selected the 18 best compounds termed FLIPins (FLIP inhibitors) for in vitro experimental tests (Supplementary Fig. 1d).
FLIPinB/FLIPinBγ enhances CD95L-/TRAIL-mediated loss of cell viability
Death ligand (DL) stimulation of the sensitive cells leads to the formation of the DISC and DED filaments followed by caspase-8 activation and cell death. To perform initial experimental screening, the effect of FLIPins was examined by measuring the viability of Jurkat cells upon stimulation with FLIPin/CD95L in comparison with CD95L-only treatment (Supplementary Fig. 2). FLIPins, which showed significant loss of viability upon their administration to Jurkat cells, were excluded from further consideration (Supplementary Fig. 3). Subsequently, only six compounds were left for further analysis, from which the compound termed FLIPinB showed the best properties in decreasing cell viability.
FLIPinB effects on CD95L- or TRAIL-induced cell viability loss were further exploited in Jurkat, HeLa-CD95 (HeLa cells overexpressing CD95), and HeLa-CD95-FL (HeLa-CD95 cells overexpressing c-FLIPL) cells (Supplementary Fig. 4). Administration of FLIPinB in Jurkat, HeLa-CD95 and HeLa-CD95-FL cells enhanced CD95L- and TRAIL-induced loss of cell viability (Supplementary Fig. 4a, c–e) and increased caspase-3/7 activity (Supplementary Fig. 4b, f). These data provide the first evidence that the compound FLIPinB promotes CD95L- and TRAIL-induced loss of cell viability and increases caspase activation.
Subsequently, the structural properties of FLIPinB were further optimized using computer design methods. In order to construct a water soluble compound and improve its binding affinity to the caspase-8/c-FLIPL heterodimer, several optimization steps were introduced (Supplementary Material). The modification of the FLIPinB structure led to the small molecule FLIPinBγ (Supplementary Fig. 5a). The key interactions observed in the predicted FLIPinBγ/p43/p10/c-FLIPL complex are shown in Fig. 1c. FLIPinBγ occupies a cavity formed between the β6-sheet strand and the α3-helix of c-FLIPL. FLIPinBγ forms hydrogen bonds with the carbonyl oxygen of Ser318 and a water molecule from the crystallographic structure [PDB ID 3H11], which was kept for the VS. Importantly, the same interaction features are observed for the L2′ loop in the procaspase-8/c-FLIPL complex (Supplementary Fig. 5b, c). In addition, the sulfonamide group is able to form hydrogen bonds with the hydroxyl group of Thr407 of c-FLIPL and caspase-8 carboxyl group of Glu396 (Fig. 1c).
The designed small compound FLIPinBγ efficiently promoted a loss of cell viability in a dose-dependent manner induced by CD95L in several cell lines. In particular, FLIPinBγ enhanced CD95L-induced cell viability loss in HeLa-CD95 and Jurkat cells (Supplementary Fig. 6a, b) as well as in acute myeloid leukemia MV4-11 cells (Supplementary Fig. 6c). Furthermore, co-treatment with FLIPinBγ increased the rate of CD95L-induced apoptotic cell death in HeLa-CD95-FL cells as was monitored by the elevated level of annexin-V-positive cells (Fig. 2a). These observations were supported by live cell imaging with IncuCyte technology and measuring the penetration into the cells of Cytotox Red, a dye which penetrates into dying cells with a compromised plasma membrane (Fig. 2b). Importantly, FLIPinBγ enhanced TRAIL-induced penetration of Cytox Red into the cells, which was most prominent in the first hours after TRAIL treatment (Fig. 2b). Taken together, these experiments have demonstrated that the optimized compound FLIPinBγ promotes CD95L- and TRAIL-mediated loss of cell viability and cell death.
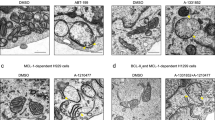
a HeLa-CD95-FL cells were pretreated with 30 µM FLIPinBγ for 2 h and stimulated with CD95L (1667 ng/mL) for 4 h. Cell death was measured using Annexin-V-FITC (A) and SytoxOrange (S)-staining and imaging flow cytometry. Mean and standard deviation of early and late apoptotic cells of three independent experiments are shown, *P < 0.05. Three representative pictures for each cell condition generated by AMNIS® are included. Abbreviation: brightfield (bf). b HeLa-CD95 cells were pretreated with 40 µM FLIPinBγ for 2 h and stimulated with TRAIL (100 ng/mL). Cells were stained with Cytotox Red Reagent. The fluorescence of cells was detected by the IncuCyte™ live cell analysis system (Essen Bioscience). At the right part, representative images after 4 h of TRAIL stimulation are shown (bf brightfield, T = 100 ng/mL TRAIL, F + T = 40 µM FLIPinBγ + 100 ng/mL TRAIL).
FLIPinBγ enhances CD95L-/TRAIL-mediated caspase activity
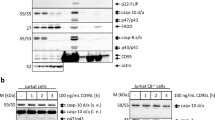
To ensure that FLIPinBγ promotes DL-mediated cell viability loss and cell death via the apoptotic branch of the pathway, we analyzed its effects on caspase activity. Both FLIPinBγ/TRAIL and FLIPinBγ/CD95L co-treatment led to an increase of caspase-3/7 activity in Jurkat cells, compared with DL-treatment only (Supplementary Fig. 6d, e). This was consistent with the analysis of caspase-3/7 activity via live cell imaging, which showed an increase in caspase-3/7 activity in single cells upon FLIPinBγ/CD95L or FLIPinBγ/TRAIL coadministration in HeLa-CD95 cells (Fig. 3a, b). These data were further supported by western blot analysis, demonstrating that FLIPinBγ increases CD95L-induced cleavage of procaspase-8 to p30 and p18 [9], as well as RIPK1 and PARP processing (Fig. 3c). In this regard, a slight increase of RIPK1 cleavage indicated that FLIPinBγ might act via increasing caspase-8 activity, as RIPK1 is one of the caspase-8 substrates [6]. Collectively, these data demonstrate that FLIPinBγ enhances caspase activation induced by DLs and that it acts on the apoptotic branch of the pathway.
a, b HeLa-CD95 cells were pretreated with 40 µM FLIPinBγ for 2 h and stimulated with CD95L (50 ng/mL) or TRAIL (100 ng/mL). Cells were stained with caspase-3/7 Green Apoptosis Assay Reagent. Caspase activity was visualized with live cell imaging. The fluorescence of caspase-3/7-positive cells was detected by the IncuCyte™ live cell analysis system (Essen Bioscience). In the upper part representative images after 2 h of CD95L stimulation are shown in a (bf brightfield, C = 50 ng/mL CD95L, F + C = 40 µM FLIPinBγ + 50 ng/mL CD95L). c HeLa-CD95 cells were pretreated with 20 µM FLIPinBγ for 2 h, followed by stimulation with CD95L (60 ng/mL) for 3 h. Total cellular lysates were analyzed by western blot using the indicated antibodies. One representative experiment out of three independent experiments is shown. Protein quantification was carried out using the normalization to actin.
c-FLIP proteins are characterized by a short half-life, which plays a central role in the regulation of their inhibitory action via fine-tuning of their intracellular concentration [10]. The binding of FLIPinBγ to c-FLIPL might cause a conformational change leading to recruitment of a putative DUB and subsequent c-FLIPL proteosomal degradation. To check if FLIPinBγ triggers a decrease of the intracellular c-FLIPL levels and thereby induces apoptosis, we analyzed whether the addition of FLIPinBγ to HeLa-CD95 cells changes the level of c-FLIPL (Supplementary Fig. 6f). Time-dependent analysis of c-FLIPL levels by western blot demonstrated no influence of FLIPinBγ on c-FLIPL expression in the first hours after its administration. Furthermore, no impact of FLIPinBγ treatment on the expression of other core components of the DISC was detected (Supplementary Fig. 6f). Thus, it was concluded that FLIPinBγ does not act via modulating the protein level of c-FLIPL.
FLIPinBγ acts via enhancement of caspase-8 activity at the DISC
DISC is a central platform for procaspase-8 activation. The predicted in silico mechanism of the FLIPinBγ effect involves the increased activity of the procaspase-8/c-FLIPL heterodimer directly at the DISC. The suggested mechanism of FLIPinBγ action assumes that it binds to the procaspase-8/c-FLIPL complex after initial processing of procaspase-8 to p43 and p10 (Fig. 4a). We assume that this leads to the stabilization of the heterodimer and an increase of caspase-8 activity in the FLIPinBγ/p43/p10/c-FLIPL complex (Fig. 4a).
a Scheme presenting the proposed mechanism of FLIPinBγ action: FLIPinBγ binds to the caspase-8/c-FLIPL complex after initial processing of procaspase-8 to p43 and p10, which leads to the enhancement of caspase-8 activity in the FLIPinBγ/p43/p10/c-FLIPL complex. The procaspase-8 is shown in red, procaspase-8 cleavage products p43 and p10 are shown in dark blue, FLIPinBγ is shown in green, and c-FLIPL is shown in light blue. b HeLa-CD95 cells were pretreated with or without 30 µM FLIPinBγ for 2 h and afterward stimulated with 60 ng/mL CD95L for 3 h. Caspase-8 activity was measured using Caspase-Glo®-8 Assay after 90 min of incubation. The experimental data points are shown as small circles. Variation of caspase-8 activity within three independent experiments is shown with box plot depicting median, 25/75% boundaries and whiskers indicating maximum and minimum values. Stars indicate statistical significance (*P; 0.05) according to two-tailed Student’s t-test. c HeLa-CD95 cells were preincubated with or without 20 µM FLIPinBγ for 2 h, followed by stimulation with 60 ng/mL CD95L for 3 h. CD95-DISC-IPs were performed and subsequently analyzed by Caspase-Glo®-8 Assay. Variation of average caspase-8 activity within three independent experiments at 0, 30, 90, and 150 min of incubation is shown with box plot depicting median, 25/75% boundaries and whiskers indicating maximum and minimum values. Stars indicate statistical significance (*P; 0.05) according to two-tailed Student’s t-test. d To control DISC formation CD95-DISC-IPs were analyzed by western blot and probed for the indicated proteins. Actin was used as loading control. One representative experiment out of three is shown. (BC = control IP without antibody). e Kinetics of caspase-8 activation for CD95L-stimulated (blue) and CD95L/FLIPinBγ co-stimulated (red) CD95-DISC-IPs for each individual experiment. Confidence intervals were calculated for technical replicates (n = 2) of each experiment and shown as error bands.
To test this hypothesis, caspase-8 activity was tested in CD95L-stimulated HeLa-CD95 cells with or without pretreatment with FLIPinBγ. Co-treatment with FLIPinBγ increased caspase-8 activity, supporting the proposed mechanism of FLIPinBγ action (Fig. 4b, Supplementary Fig. 7a, b). Further, the CD95 DISC was immunoprecipitated from HeLa-CD95 cells with or without pretreatment with FLIPinBγ using anti-CD95 (anti-APO-1) antibodies. Subsequently, the caspase-8 activity in the DISC-IPs was analyzed using Caspase-8-Glo assay (Fig. 4c, Supplementary Fig. 7c). DISC-IPs contained all core components of the DISC: CD95, FADD, procaspases-8 and -10 and their cleavage products (Fig. 4d). Co-treatment with FLIPinBγ increased caspase-8 activity at the DISC, indicating that FLIPinBγ acts directly at the DISC by specifically increasing the caspase-8 activity (Fig. 4c, e, Supplementary Fig. 7c). This experimental analysis further supports the computational predictions on the mechanism of FLIPinBγ.
Validation of the cell response to FLIPinBγ treatment with a computational model
To get detailed quantitative insights into the mechanism of FLIPinBγ function at the DED filaments and thereby to understand procaspase-8/c-FLIPL heterodimer action, we used the cutting edge technology of mathematical modeling [32, 33]. To this end, a biochemical model of procaspase-8 activation at the DED filament has been created based on ODEs. The topology of the model included the formation of the homo- (procaspase-8/procaspase-8) and heterodimers (procaspase-8/c-FLIPL) at the DED filaments (Fig. 5a). After homo- and heterodimerization, procaspase-8 undergoes autocatalytic activation in the DED filament, which is followed by intra- and intermolecular processing of the dimers into p43 and p10 [34]. This is accompanied by the caspase-8-mediated cleavage of procaspase-3 to caspase-3 and its subsequent activation, leading to cell death. The initial binding constants and related parametrization were based on our previous models of the DISC generated in HeLa-CD95 cells [25]. The ratios between procaspase-8 and c-FLIP at the DISC were based on quantitative proteomics analysis performed in our previous work and indicating substoichiometric amounts of c-FLIP in the DED filament with a ratio of 1 to 10 (c-FLIP to procaspase-8) [4]. This subsequently results in the decreased ratios of procaspase-8/c-FLIPL heterodimers to procaspase-8/procaspase-8 homodimers and rather low contribution of procaspase-8/c-FLIPL heterodimers to enhancement of caspase-8 activity in HeLa-CD95 cells as reported by us previously [14]. The generated model was trained and validated with experimental data for caspase-8 activity at the DISC (Fig. 5b), total cellular caspase-3/7 activity (Fig. 5c), and penetration of Cytox Red into the cells that was used as an indicator of cell death (Fig. 5d). The simulations generated with selected model parameters were fitting well to the experimental data (Fig. 5b–d).
a The topology of ODE model describing the activation of caspase-8 in the DED filament. kd—dimerization rate for procaspase-8/procaspase-8 and procaspase-8/caspase-8 dimers; γ·kd—dimerization rate for c-FLIPL/procaspase-8 and caspase-8/caspase-8 dimers; kp—processing rate; kcd and kcd_c3 rates for cell death substrate cleavage by caspase-8 and caspase-3, respectively. kp3, kp3_c3 rates of procaspase-3 cleavage by caspase-8 and caspase-3 respectively. b–d Simulations of the model and experimental data used for the model training. The diagrammes show caspase-8 activity at the DISC (b), caspase-3/7 activity (c), and the penetration of Cytox Red into the cells that was used as an indicator of cell death (d). Lanes present model simulations, while points experimentally measured values. Treatment with CD95L is indicated in blue, while the co-treatment CD95L/FLIPinBγ is shown in red.
The modeling predicted that FLIPinBγ would stabilize p43/p10/c-FLIPL complex up to 4 h after CD95L stimulation resulting in its sustained activity over this period of time (Fig. 6a). In fact, the in silico simulations have shown that the concentration of the p43/p10/c-FLIPL complex increases from 1 to 3 nM within the first hours after addition of CD95L (Fig. 6a). This in turn results in the subsequent increase of the concentration of active caspase-8 upon addition of FLIPinBγ for 4 h after stimulation (Fig. 6a). This prediction was fitting well to the experimental data on caspase activation and finally provided an explanation why the effects of FLIPinBγ on caspase activity are only detectable in the first hours after DL stimulation.
Model simulations for the dynamics of active caspase-8 at the DISC (a) and the penetration of Cytox Red into the cells that was used as an indicator of cell death (b). The concentration of the CD95L used for simulation was 60 ng/mL. Treatment with CD95L is marked in blue, while the co-treatment CD95L/FLIPinBγ is shown in red. c HeLa-CD95 cells were pretreated with 40 µM FLIPinBγ for 2 h and stimulated with CD95L (50 ng/mL). Cells were stained with Cytotox Red Reagent. The penetration of Cytox Red into the cells that was used as an indicator of cell death was detected by the IncuCyte™ live cell analysis system (Essen Bioscience). In the right part representative images after 4 h of CD95L stimulation are shown (bf brightfield, C = 50 ng/mL CD95L, C + F = 40 µM FLIPinBγ + 50 ng/mL CD95L). Heat maps corresponding to predictions of the degree of cell death (d) and active caspase-8 (e) are shown. The dependence upon the time of CD95L administration and the concentration of c-FLIPL for CD95L-only treatment (left) and CD95L/FLIPinBγ co-treatment (center) are presented. d (right) The model predictions for FLIPinBγ enhancement of cell death after 1 h of CD95L stimulation depending upon c-FLIPL/procaspase-8 ratio. e (right) The model predictions for the time-dependent generation of active caspase-8 at the DISC for 1/2 (c-FLIPL/procaspase-8) ratio.
To test how the model would compute the timing of cell death upon the addition of FLIPinBγ the simulations for apoptosis induction with 60 ng/mL of CD95L with or without FLIPinBγ were performed. The model has predicted the increase of the cell death within the first four hours after stimulation, which disappeared at later time points (Fig. 6b). Modeling predictions were validated by measuring CD95L-induced penetration of Cytox Red into the cells that was used as an indicator of cell death in HeLa-CD95 cells via live cell imaging using the IncuCyte technology as described previously (Fig. 6c). In accordance with in silico predictions, FLIPinBγ enhanced CD95L-induced penetration of Cytox Red into the cells in vitro in the first hours after CD95L/FLIPinBγ co-treatment (Fig. 6c).
HeLa-CD95 cells that were used for the modeling are characterized by relatively low c-FLIP expression with ~1 to 10 (c-FLIP to procaspase-8) ratio at the DED filaments (Fig. 6d). The model simulations show the enhancement of cell death (Fig. 6d) and caspase-8 activity (Fig. 6e) up to several fold upon a moderate increase of c-FLIPL level. Hence, predictions in silico have demonstrated that a moderate increase of c-FLIPL level improves the effects of FLIPinBγ. This emerges from the increased number of caspase-8/c-FLIPL heterodimers in the DED filament subsequently resulting in the enhanced amount of active caspase-8 that in turn promotes apoptosis (Fig. 6e). In particular, the highest sensitization effects of FLIPinBγ on cell death were observed at approximately one to two ratio of c-FLIPL to procaspase-8 (Fig. 6d). Hence, according to the model predictions upon the increase of c-FLIPL levels, FLIPinBγ has much higher potential to accelerate caspase-8 activation and apoptosis.
Taken together, our modeling approach fully supported the predicted mechanism of FLIPinBγ action manifesting in the FLIPinBγ-mediated increase of the stability of heterodimer and thereby prolonging its “catalytic life” at the DED filament.
Discussion
In this study, we used advanced methods of in silico computational biology to design a small molecule targeting the heterodimer caspase-8/c-FLIPL, which plays a key role in apoptosis regulation. The design strategy of the small molecule was to mimick the stabilizing effect on the L2′ loop in a “closed” conformation, which should in turn lead to the increase of caspase-8 activity at the DISC and more efficient apoptosis induction. The latter is based on the hypothesis that the switch of the L2′ loop from a “closed” to an “open” conformation, as induced by L2 loop processing, might lead to the destabilization of the p43/p10/c-FLIPL complex. The experimental data that were obtained with the small molecule FLIPinBγ support the predicted in silico mechanism. Our data strongly indicate that the stabilization of the active center of caspase-8 in the heterodimer procaspase-8/c-FLIPL is required for its activation. Moreover, our results support the hypothesis of L2′ loop translocation as the key event in the deactivation of this heterodimer. Furthermore, co-treatment with FLIPinBγ and DL caused enhanced apoptotic cell death and caspase-8 activity at the DISC providing evidence for the mechanism suggested for FLIPinBγ action in silico.
The suggested molecular mechanism of FLIPinBγ might involve allosteric regulation, which has been also reported for other caspases, as well as a stabilizing role for heterodimer complex formation. Caspases are known to have an allosteric site in their intersubunit homodimerization interface [35, 36]. In particular, such allosteric sites of caspase-1, caspase-7, and caspase-8 have been recently targeted by small molecules. Allosteric inhibitors of caspase-7 and caspase-1 were shown to lock the inactive zymogen conformation [35, 36]. Accordingly, we suggest an existence of previously undescribed allosteric site of caspase-8 activation within the interface of the heterodimer caspase-8/c-FLIPL. A small molecule targeting the interface of caspase-8 homodimer was shown to enhance TRAIL-induced apoptosis but at high compound concentration [37]. Therefore, we assumed that targeting the heterodimerization interface of the caspase-8/c-FLIPL complex could be a promising strategy for further design of CD95L and TRAIL potentiating compounds.
Cellular signal transmission that regulates apoptosis represents an ideal system to go into quantitative studies using methods of the emerging field of systems biology [32, 33]. The computational modeling of the apoptotic pathway dynamics is very advanced, and detailed experimentally validated models focusing on multiple aspects are available [32]. In this study we have generated a link between in silico methods to modeling the apoptotic signaling pathways and cancer therapy by extending the model of the DISC and apoptosis signaling with the effect of the specific small molecule FLIPinBγ that increases the activity of the caspase-8/c-FLIPL heterodimer and thereby promoting apoptosis. Importantly, the generated model can describe the biochemical reaction kinetics of FLIPinBγ action at the DISC, predict an increase in heterodimer concentration leading to the increase of caspase-8 activity, and, most importantly, precisely predict the response of cancer cell population to the addition of FLIPinBγ. Moreover, the model allows to predict the action of FLIPinBγ depending on the c-FLIPL expression. This is a crucial step for therapies that might use the possibilities of modeling of interactions of drugs specific for cell death pathways. Interestingly, for growth factor related pathways, the model based drug design has already made the step to clinical studies [38].
The constructed model has shown that boosting of procaspase-8 processing at initial time points after CD95L/TRAIL stimulation promotes apoptosis induction. The latter might be explained by a high degradation rate of the DISC complex as well as by the rate limiting process of caspase-8 oligomerization to form catalytically active subunits. Indeed, a half-life of the DISC was estimated to be only 1 h [34], while according to our model it requires significantly more time to activate all procaspase-8 molecules at the DISC. Moreover, we can conclude that enhancing procaspase-8 processing rate at the first hours after DL stimulation is the general strategy to achieve higher cell death rates. This is in line with the previous reports on the contribution of the rate of procaspase-8 activation to the cell death [34, 39]. In this regard, our approach showed that extending the time of heterodimer activity via stabilizing effects of small molecules might serve as a promising strategy in the future research.
The moderate proapoptotic activity of the designed compound FLIPinBγ might be explained by several factors. A short half-life of the caspase-8/c-FLIPL complex might require a high concentration of FLIPinBγ compound to achieve a stabilizing effect. The moderate activity of FLIPinBγ might be due to the low abundance of c-FLIPL in the DED filaments. Indeed, a quantitative proteomics analysis of DED filaments demonstrated that c-FLIP is present in a ten times lower concentration in the filaments as compared with procaspase-8 [4]. Therefore, there is only a limited number of procaspase-8/c-FLIPL heterodimers present in each chain limiting the effects of FLIPinBγ. Furthermore, according to prediction of the kinetic model FLIPinBγ activity can be enhanced up to several times upon increase of c-FLIPL/procaspase-8 ratio (Fig. 6d). In this regard, consideration that a number of cancer cells have a high level of c-FLIPL allows to suggest that the development of compounds based on FLIPinBγ in general has an extremely high therapeutic potential.
In contrast to the major regulators of the intrinsic apoptosis pathway, to which specific small molecule-based inhibitors have already been successfully developed and are currently in clinical trials, the extrinsic pathway is only starting to be addressed [40]. Taken together, this work provides a basis for the development of new therapies against cancer via specific enhancement of the activity of the key enzyme of the extrinsic apoptosis-caspase-8 and paving new avenues for specifically targeting DR-induced apoptosis.
References
Krammer PH, Arnold R, Lavrik IN. Life and death in peripheral T cells. Nat Rev Immunol. 2007;7:532–42.
Lavrik IN, Krammer PH. Regulation of CD95/Fas signaling at the DISC. Cell Death Differ. 2012;19:36–41.
Dickens LS, Boyd RS, Jukes-Jones R, Hughes MA, Robinson GL, Fairall L, et al. A death effector domain chain DISC model reveals a crucial role for caspase-8 chain assembly in mediating apoptotic cell death. Mol Cell. 2012;47:291–305.
Schleich K, Warnken U, Fricker N, Ozturk S, Richter P, Kammerer K, et al. Stoichiometry of the CD95 death-inducing signaling complex: experimental and modeling evidence for a death effector domain chain model. Mol Cell. 2012;47:306–19.
Fu TM, Li Y, Lu A, Li Z, Vajjhala PR, Cruz AC, et al. Cryo-EM structure of caspase-8 tandem DED filament reveals assembly and regulation mechanisms of the death-inducing signaling complex. Mol Cell. 2016;64:236–50.
Hughes MA, Harper N, Butterworth M, Cain K, Cohen GM, MacFarlane M. Reconstitution of the death-inducing signaling complex reveals a substrate switch that determines CD95-mediated death or survival. Mol Cell. 2009;35:265–79.
Yu JW, Jeffrey PD, Shi Y. Mechanism of procaspase-8 activation by c-FLIPL. Proc Natl Acad Sci USA. 2009;106:8169–74.
Golks A, Brenner D, Schmitz I, Watzl C, Krueger A, Krammer PH, et al. The role of CAP3 in CD95 signaling: new insights into the mechanism of procaspase-8 activation. Cell Death Differ. 2006;13:489–98.
Hoffmann JC, Pappa A, Krammer PH, Lavrik IN. A new C-terminal cleavage product of procaspase-8, p30, defines an alternative pathway of procaspase-8 activation. Mol Cell Biol. 2009;29:4431–40.
Ozturk S, Schleich K, Lavrik IN. Cellular FLICE-like inhibitory proteins (c-FLIPs): fine-tuners of life and death decisions. Exp Cell Res. 2012;318:1324–31.
Scaffidi C, Schmitz I, Krammer PH, Peter ME. The role of c-FLIP in modulation of CD95-induced apoptosis. J Biol Chem. 1999;274:1541–8.
Golks A, Brenner D, Krammer PH, Lavrik IN. The c-FLIP-NH2 terminus (p22-FLIP) induces NF-kappaB activation. J Exp Med. 2006;203:1295–305.
Golks A, Brenner D, Fritsch C, Krammer PH, Lavrik IN. c-FLIPR, a new regulator of death receptor-induced apoptosis. J Biol Chem. 2005;280:14507–13.
Fricker N, Beaudouin J, Richter P, Eils R, Krammer PH, Lavrik IN. Model-based dissection of CD95 signaling dynamics reveals both a pro- and antiapoptotic role of c-FLIPL. J Cell Biol. 2010;190:377–89.
Schleich K, Buchbinder JH, Pietkiewicz S, Kahne T, Warnken U, Ozturk S, et al. Molecular architecture of the DED chains at the DISC: regulation of procaspase-8 activation by short DED proteins c-FLIP and procaspase-8 prodomain. Cell Death Differ. 2016;23:681–94.
Hughes MA, Powley IR, Jukes-Jones R, Horn S, Feoktistova M, Fairall L, et al. Co-operative and hierarchical binding of c-FLIP and caspase-8: a unified model defines how c-FLIP isoforms differentially control cell fate. Mol Cell. 2016;61:834–49.
Chang DW, Xing Z, Pan Y, Algeciras-Schimnich A, Barnhart BC, Yaish-Ohad S, et al. c-FLIP(L) is a dual function regulator for caspase-8 activation and CD95-mediated apoptosis. EMBO J. 2002;21:3704–14.
Micheau O, Thome M, Schneider P, Holler N, Tschopp J, Nicholson DW, et al. The long form of FLIP is an activator of caspase-8 at the Fas death-inducing signaling complex. J Biol Chem. 2002;277:45162–71.
Boatright KM, Deis C, Denault JB, Sutherlin DP, Salvesen GS. Activation of caspases-8 and -10 by FLIP(L). Biochemical J. 2004;382:651–7.
Pop C, Oberst A, Drag M, Van Raam BJ, Riedl SJ, Green DR, et al. FLIP(L) induces caspase 8 activity in the absence of interdomain caspase 8 cleavage and alters substrate specificity. Biochem J. 2011;433:447–57.
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem. 2004;47:1739–49.
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem. 2006;49:6177–96.
Sastry GM, Adzhigirey M, Day T, Annabhimoju R, Sherman W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput-Aided Mol Des. 2013;27:221–34.
Irwin JJ, Sterling T, Mysinger MM, Bolstad ES, Coleman RG. ZINC: a free tool to discover chemistry for biology. J Chem Inf Model. 2012;52:1757–68.
Neumann L, Pforr C, Beaudouin J, Pappa A, Fricker N, Krammer PH, et al. Dynamics within the CD95 death-inducing signaling complex decide life and death of cells. Mol Syst Biol. 2010;6:352.
Schmidt JH, Pietkiewicz S, Naumann M, Lavrik IN. Quantification of CD95-induced apoptosis and NF-kappaB activation at the single cell level. J Immunol Methods. 2015;423:12–7.
Pietkiewicz S, Eils R, Krammer PH, Giese N, Lavrik IN. Combinatorial treatment of CD95L and gemcitabine in pancreatic cancer cells induces apoptotic and RIP1-mediated necroptotic cell death network. Exp Cell Res. 2015;339:1–9.
Hillert LK, Bettermann-Bethge K, Nimmagadda SC, Fischer T, Naumann M, Lavrik IN. Targeting RIPK1 in AML cells carrying FLT3-ITD. Int J Cancer. 2019;145:1558–69.
Pietkiewicz S, Schmidt JH, Lavrik IN. Quantification of apoptosis and necroptosis at the single cell level by a combination of imaging flow cytometry with classical annexin V/propidium iodide staining. J Immunol Methods. 2015;423:99–103.
Raue A, Kreutz C, Maiwald T, Bachmann J, Schilling M, Klingmuller U, et al. Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics. 2009;25:1923–9.
Vickers CJ, Gonzalez-Paez GE, Litwin KM, Umotoy JC, Coutsias EA, Wolan DW. Selective inhibition of initiator versus executioner caspases using small peptides containing unnatural amino acids. ACS Chem Biol. 2014;9:2194–8.
Spencer SL, Sorger PK. Measuring and modeling apoptosis in single cells. Cell. 2011;144:926–39.
Lavrik IN. Systems biology of death receptor networks: live and let die. Cell Death Dis. 2014;5:e1259.
Kallenberger SM, Beaudouin J, Claus J, Fischer C, Sorger PK, Legewie S, et al. Intra- and interdimeric caspase-8 self-cleavage controls strength and timing of CD95-induced apoptosis. Sci Signal. 2014;7:ra23.
Hardy JA, Lam J, Nguyen JT, O’Brien T, Wells JA. Discovery of an allosteric site in the caspases. Proc Natl Acad Sci USA. 2004;101:12461–6.
Scheer JM, Romanowski MJ, Wells JA. A common allosteric site and mechanism in caspases. Proc Natl Acad Sci USA. 2006;103:7595–600.
Bucur O, Gaidos G, Yatawara A, Pennarun B, Rupasinghe C, Roux J, et al. A novel caspase 8 selective small molecule potentiates TRAIL-induced cell death. Sci Rep. 2015;5:9893.
Schoeberl B, Kudla A, Masson K, Kalra A, Curley M, Finn G, et al. Systems biology driving drug development: from design to the clinical testing of the anti-ErbB3 antibody seribantumab (MM-121). NPJ Syst Biol Appl. 2017;3:16034.
Roux J, Hafner M, Bandara S, Sims JJ, Hudson H, Chai D, et al. Fractional killing arises from cell-to-cell variability in overcoming a caspase activity threshold. Mol Syst Biol. 2015;11:803.
Roberts AW, Davids MS, Pagel JM, Kahl BS, Puvvada SD, Gerecitano JF, et al. Targeting BCL2 with Venetoclax in Relapsed Chronic Lymphocytic Leukemia. N Engl J Med. 2016;374:311–22.
Acknowledgements
We acknowledge the center of dynamic systems, funded by the EU-programme European Regional Development Fund, Volkswagen Foundation (VW 90315), Wilhelm Sander-Stiftung (2017.008.01), DFG (LA 2386), and Russian Science Foundation 14-44-00011 for supporting our work. We thank Dr Sabine Pietkiewicz, Claudia Arndt, and Dr Nastya Shunaeva-Zotova for the experimental help. We acknowledge Prof. Peter H. Krammer (DKFZ, Heidelberg) for providing us with anti-APO-1, C15, NF6, and 1C4 antibodies.
Author information
Authors and Affiliations
Contributions
LH performed experiments and contributed to the manuscript text. NI performed computational modeling and worked a lot on the manuscript text. DB, CK, and JE contributed experimentally. SP and NK contributed to the theoretical design. VI and IL designed a study and wrote a manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by S. Fulda
Rights and permissions
About this article
Cite this article
Hillert, L.K., Ivanisenko, N.V., Busse, D. et al. Dissecting DISC regulation via pharmacological targeting of caspase-8/c-FLIPL heterodimer. Cell Death Differ 27, 2117–2130 (2020). https://doi.org/10.1038/s41418-020-0489-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-020-0489-0
This article is cited by
-
The role of cell death in SARS-CoV-2 infection
Signal Transduction and Targeted Therapy (2023)
-
cFLIP downregulation is an early event required for endoplasmic reticulum stress-induced apoptosis in tumor cells
Cell Death & Disease (2022)
-
The non-apoptotic function of Caspase-8 in negatively regulating the CDK9-mediated Ser2 phosphorylation of RNA polymerase II in cervical cancer
Cellular and Molecular Life Sciences (2022)
-
The role of death domain proteins in host response upon SARS-CoV-2 infection: modulation of programmed cell death and translational applications
Cell Death Discovery (2020)
-
Pharmacological targeting of c-FLIPL and Bcl-2 family members promotes apoptosis in CD95L-resistant cells
Scientific Reports (2020)