Abstract
Acquired platinum resistance poses a significant therapeutic impediment to ovarian cancer patient care, accounting for more than 200,000 deaths annually worldwide. We previously identified that overexpression of the antioxidant superoxide dismutase 1 (SOD1) in ovarian cancer is associated with a platinum-resistant phenotype via conferring oxidative stress resistance against platinum compounds. We further demonstrated that enzymatic inhibition using small-molecule inhibitors or silencing of SOD1 via RNA interference (RNAi) increased cisplatin sensitivity and potency in vitro. We launched this study to explore the potential therapeutic applications of SOD1 silencing in vivo in order to reverse cisplatin resistance using a graphene-based siRNA delivery platform. PEGylated graphene oxide (GO) polyethyleneimine (GOPEI-mPEG) nanoparticle was complexed with SOD1 siRNA. GOPEI-mPEG-siSOD1 exhibited high biocompatibility, siRNA loading capacity, and serum stability, and showed potent downregulation of SOD1 mRNA and protein levels. We further observed that cisplatin and PEI elicited mitochondrial dysfunction and transcriptionally activated the mitochondrial unfolded protein response (UPRmt) used as a reporter for their respective cytotoxicities. SOD1 silencing was found to augment cisplatin-induced cytotoxicity resulting in considerable tumour growth inhibition in cisplatin-sensitive A2780 and cisplatin-resistant A2780DDP subcutaneous mouse xenografts. Our study highlights the potential therapeutic applicability of RNAi-mediated targeting of SOD1 as a chemosensitizer for platinum-resistant ovarian cancers.
Similar content being viewed by others
Introduction
Ovarian cancer, with an annual worldwide incidence of 313,000 cases and mortality of 207,000, is considered the second most lethal gynaecological malignancy [1]. Despite the favourable overall survival when detected in early stage 1—attributed principally to its asymptomatic disease progression—most cases are diagnosed in advanced stages 2 and 3 with a 5-year survival of 31%. The overall global survival has only increased modestly in previous decades due to the constrained availability of treatment options and the clinically acquired chemotherapy resistance [2]. Despite most patients responding to debulking surgery and combinational treatment with platinum and taxols, almost half the patients eventually develop recurrence and become resistant or refractory to additional platinum-based therapeutic interventions [2].
Despite their adverse side effect profile, platinum compounds are still considered potent as the first-line drug of choice as mono- or synergistic therapy for many solid tumours [3,4,5,6]. Besides nuclear DNA adduct formation and subsequent induction of apoptotic signalling, platinum drugs also elicit mitochondrial dysfunction through oxidative stress and mitochondrial DNA (mtDNA) damage [7, 8]. Contingent on the intrinsic platinum sensitivity of the respective malignancy, platinum drugs initially exhibit robust therapeutic efficiency. However, the development of acquired platinum resistance of recurrent tumours renders their subsequent clinical applicability ineffective. Lung, prostate, and colorectal cancers are intrinsically resistant to platinum; acquired resistance is also more common in epithelial ovarian cancer [9]. Implicitly, preventing or reversing this well-documented clinical phenomenon could have profound and widespread clinical therapeutic benefits.
Intrinsic and acquired platinum resistance have been linked to reduced drug uptake, increased efflux, enhanced detoxification, elevated scavenger levels, increased oxidative stress tolerance, upregulated DNA repair mechanisms, and the reprogramming of cellular metabolism to evade cisplatin-induced death [10, 11]. Cancer cells may exhibit one, or more of the aforementioned peculiar mechanisms that ultimately determine their net sensitivity to platinum [12, 13]. However, to date, the targeting of drug-transporting pathways of MDR-1, ATP-7A/7B, CTR1, MRP, and DNA repair pathways, including BRCA1/2, ERCC1, and MMR, provided only modest improvement in survival clinically [14]. In addition, the utilization of MDR1 inhibitors, including but not limited to Zosuquidar, MK-571, and PSC-833, showed moderate efficacy in terms of platinum chemosensitization in ovarian cancer [15]. Thus far, the exact mechanisms and determinant factors driving platinum resistance have not been fully elucidated. Due to genetic and patient sample heterogeneity, patient-specific expression levels of potentially robust platinum resistance biomarkers make the discovery process burdensome.
In the quest for a novel therapeutic target for cisplatin resistance, we previously identified the ROS-neutralizing SOD1 to be overexpressed in cisplatin-resistant ovarian cancer cell lines using quantitative label-free comparative proteomics analysis [16]. The 32 kDa Cu2+/Zn2+ SOD1 is an abundantly expressed intracellular homodimeric metalloenzyme that converts superoxide anions to hydrogen peroxide that is subsequently transformed by catalase to oxygen and water [17]. Superoxide neutralization by SOD1 is a crucial mechanism in counteracting oxidative damage, as SOD1 knockout Drosophila exhibits reduced lifespan, infertility, and hypersensitivity to oxidative stress [18]. Other studies concluded that RNAi-mediated SOD1 silencing provokes senescence in human fibroblasts and induces apoptosis in HeLa cells via ROS-mediated induction of TP53 [19]. Further, SOD1 plays an essential role in cytoplasmic NRF2-mediated adaptation to oxidative stress and in the mitochondria via UPRmt [20]. Thus, our previous results suggest the role of SOD1 overexpression as a defense mechanism of ovarian cancer cells to counteract platinum-induced oxidative stress and modulation of ROS-mediated redox signalling [16]. We further concluded that enzymatic inhibition of SOD1 using copper/zinc chelating agents of TETA and ATN-224 reversed platinum resistance [21]. Subsequently, to overcome the off-target side effects caused by non-specific small molecule metal chelators, we further corroborated the chemosensitizing effect of SOD1 downregulation via RNAi in vitro [22].
RNAi is a validated gene therapeutic method to control post-transcriptional gene regulation in various disease states [23]. However, the exogenous introduction of therapeutic siRNA for in vivo applications faces numerous obstacles, including easy degradation, short half-life, filtration by the kidneys, poor cellular uptake due to inherent negative charge, structural instability, and degradation by RNases [24]. Therefore, a meticulous design of any siRNA delivery system is fundamental to fully capture the potential of siRNA therapeutics. Non-viral gene delivery vectors such as cationic lipids, cationic polymers, and polysaccharides, in particular, have gained momentum due to their efficacy and biosafety compared to viral vectors for delivering DNA or RNA cargo into the cells [25].
The two-dimensional graphene oxide (GO), due to its convenient applicability for non-covalent or covalent functionalization via abundant epoxy, carboxyl, and hydroxyl surface groups, has been widely used in combination with various cationic polymers for siRNA and gene delivery [26, 27]. GO is also highly biocompatible in most in vitro and in vivo test systems due to its high colloidal stability, water dispersibility, and surface-to-volume area ratio [28]. We previously synthesized graphene-based siRNA, small molecule, and combined drug delivery systems for various applications [29]. Consequently, in this study, a novel graphene-based nanoparticle platform (GOPEI-mPEG) was prepared by sequential coupling of nano-graphene oxide with cationic polymer polyethyleneimine (PEI) to achieve SOD1 siRNA delivery and further with polyethylene glycol (PEG) to increase biocompatibility and control surface charge. In addition, the incorporation of GO, due to the fixed lateral dimensions of the 2D sheets, can allow for improved control of a more uniform and reproducible final hydrodynamic nanoparticle size, compared to using PEI-PEG polyplexes alone [30, 31]. Our study aimed to evaluate the in vivo chemosensitizing efficacy of SOD1 siRNA by GOPEI-mPEG delivery system in cisplatin-resistant mouse xenograft models.
Materials and methods
Nanoparticle preparation
Graphene oxide (GO) synthesis
GO was prepared with a modified Hummer’s method [32]. The mixture of NaCl (35 g) and native graphite flakes (1 g, Alfa Aesar, Haverhill, MA, USA) was ground with a mortar until the colour of the mixture turned grey, then dissolved in deionized water (DIW). NaCl was extracted by multiple washing steps and ultracentrifugation at 8000 rpm for 5 min. The product was dried at 90 °C for 12 h and stirred in a three-necked bottle with H2SO4 (23 mL) for 8 h. In an ice bath, KMnO4 (3 g) was added to the mixture keeping the temperature below 20 °C, until the colour turned dark green. The solution was further stirred in a dimethyl-silicone oil bath at 38 °C for 30 min and at 70 °C for an additional 45 min until the colour of the solution turned dark brown. The mixture was diluted with DIW (5 mL) and stirred for 10 min. Next, the solution was diluted with DIW (40 mL) and stirred for 15 min at 100 °C. Subsequently, the mixture of DIW (10 mL) and H2O2 (10 mL) was added and stirred for 10 min, and the final product was washed twice with 5% HCl (50 mL) and numerous times with DIW via ultracentrifugation. The prepared GO batch was dialyzed using 100 kDa dialysis bags. Next, GO (8 mL) was washed twice with DIW, and the –COOH groups were activated by adding NaOH (1.8 g). Finally, the suspension was diluted with DIW up to 15 mL and stirred at 55 °C for 4 h. Next, the solution was neutralized by adding HCl (5 mL), and the mixture was washed multiple times until the pH became neutral. Following a 30-min bath sonication and centrifugation at 13,000 rpm for 5 min, the concentration of the supernatant was measured with an Evolution 201 UV–Vis spectrophotometer (Thermo-Fisher Scientific, Carlsbad, CA, USA).
GOPEI synthesis
The conjugation of COOH groups of GO and primary amines of PEI was achieved with zero-length carboxyl to amine carbodiimide (EDC) cross-linking. Carboxyl groups were activated with EDC forming an active but unstable O-acylisourea intermediate displaced by nucleophilic attack from the primary amines of PEI. These primary amines formed an amide bond with the activated carboxylic groups. In brief, GO (5 mg) was diluted up to 15 mL in phosphate-buffered saline (PBS) and was bath sonicated for 30 min in ice before adding 30, 60, or 90 mg of PEI25 kDa (Sigma-Aldrich, St. Louis, MO, USA) dissolved in 200 µL of PBS (HyClone Laboratories Inc, Logan, UT, USA). Following 5 min of sonication, NaOH (20 µL) and 5 mg of Pierce™ Premium Grade EDC [1-Ethyl-3-(3-dimethyl aminopropyl) carbodiimide] (Thermo-Fisher Scientific) were added, respectively, then sonicated for 5 min and stirred for 30 min. GOPEI was prepared using three different PEI concentrations; 30, 60, and 90 mg/mL corresponding with GOPEI1X, GOPEI2X, GOPEI3X, respectively. Next, 10 mg of EDC in 400 µL PBS was added under continuous stirring, sonicated for 30 min, and then stirred overnight. The final EDC concentration was 1 mg/mL in a 10-fold molar or weight excess to GO. The prepared GOPEI nanoparticles were purified with ultrafiltration using a 100 kDa Amicon Ultra-15 centrifugal filter (Merck Millipore Ltd., Darmstadt, Germany).
GOPEI-mPEG synthesis
Methyl-PEG (2000 kDa, 5 mg) (Xi'an Ruixi Biological Technology Co., Ltd, Shaanxi, China) in PBS was diluted to 15 mL and sonicated for 5 min. NaOH (20 µL) and EDC (5 mg) were added and sonicated for 30 min. Next, GOPEI (30 mg) was added, and the solution was sonicated for 5 min before adding EDC (10 mg), then stirred overnight. The nanoparticle was purified with ultrafiltration using a 100 kDa filter (Millipore Sigma, Burlington, MA, USA).
Nanoparticle characterisation
Atomic force microscopy (AFM)
The size and surface morphology of GO and GOPEI were characterized with AFM. The nanoparticle samples (200 μL) were transferred onto a 1.5 cm × 1.5 cm muscovite mica sheet and imaged with Veeco Dimension 3100 atomic force microscope (Veeco Instruments, Bruker, MA, USA). The images were analyzed using V700 (Veeco) and Nanoscope v.7.00b19 (Veeco) software.
Transmission electron microscopy (TEM)
The lateral size and qualitative structural properties of GO and GOPEI were assessed with TEM. The nanoparticle samples were transferred onto a 20 cm × 20 cm muscovite mica sheet, then imaged by a Tecnai G2 F20 S-Twin transmission electron microscope (Thermo-Fisher Scientific), and analyzed by Digital Micrograph 3.0 (Thermo-Fisher Scientific) software.
Dynamic light scattering (DLS)
The average size distribution (ASD), the polydispersity index (PDI), and surface charge (ζ-potential) of GO, GOPEI, GOPEI-mPEG nanoparticles were determined with DLS. The samples (100 μL) were first filtered, then diluted up to 2 mL with DIW and transferred into a ZEN0112-low volume disposable sizing cuvette for ASD and PDI and into a DTS1060C-Clear disposable zeta cell for ζ-potential analysis. The measurements were performed with a Zetasizer Nanoseries Nano-25 (Malvern, United Kingdom) in batch mode at 25 °C in triplicates with 120 sec equilibration time. The results were analyzed with the Zetasizer (Malvern, United Kingdom) software.
Fourier transform infrared (FT-IR) spectroscopy
The confirmation of successful covalent amide bond formation between GO and PEI was determined with FT-IR spectroscopic analysis for GO, GOPEI, GOPEI-mPEG samples with a Cary 600 series (Agilent Technologies, Santa Clara, CA, USA) FTIR Spectrometer, and all obtained results were analyzed with the Resolution Pro (Agilent Technologies, Santa Clara, CA, USA) and displayed with the GraphPad Prism 7 software (GraphPad Software Inc, Boston, MA, USA).
Ultraviolet-visible (UV–Vis) spectroscopy
The nanoparticle absorbance spectra and the siRNA oligonucleotide loading capacity were measured and confirmed with UV–Vis spectroscopy. The GO (0.01 mg/mL), PEI (0.005 and 0.01 mg/mL), GOPEI, GOPEI-siSCR (scrambled negative control), GOPEI-mPEG, and GOPEI-mPEG-siSCR samples complexed at w/w ratio of 3:1 (nanoparticle : siRNA) at RT for 30 min and scrambled negative control siRNA (siSCR) only samples (50 μL) were dissolved in DIW (2 mL) in a 10 mm quartz cuvette (3.5 mL). The absorbance spectrum was recorded compared to DIW as a calibration control. The UV–Vis spectra were measured with an Evolution 201 UV–Vis spectrophotometer (Thermo-Fisher Scientific) between 190 and 800 nm in absorbance mode and were analyzed with the Thermo Cue (Thermo-Fisher Scientific) software.
Thermogravimetric analysis (TGA)
The composition and PEI surface coating of lyophilized GO and GOPEI nanoparticles were quantitatively assessed with an STA 449 F3 Jupiter® thermal analyzer (Netzsch, Germany). Under inert nitrogen purge (50 mL/min), the samples were heated (25–800 °C, 50 °C/min), and the change of the initial nanoparticle mass (5 mg) was measured as a function of time and temperature. Ceramic crucibles with loosely fitted lids containing the samples were held at 25 °C for 5 min prior to measurements to allow for equilibration between the crucible and the furnace. To correct the bias caused by the buoyancy effect, a blank measurement with an empty ceramic crucible under the same temperature program as the samples were performed. Results were analyzed with the Netzsch Proteus Thermal Analysis software and displayed as a thermogravimetric temperature vs mass curve with the following formula:
where Wt=0 is the initial sample weight, and W Sample ∆t is the measured sample weight at different time points.
Colloidal stability studies
To assess the dispersibility, stability, and aggregation potential of GO, GOPEI, and GOPEI-mPEG, the nanoparticles were dissolved in various serum-containing and serum-free salt solutions, including DIW, PBS, normal saline (0,9%), RPMI (HyClone Laboratories Inc, Logan, UT, USA), RPMI with 5% v/v FBS (Biological Industries, Cromwell, CT, USA), RPMI with 10% v/v FBS, 50% v/v FBS and 50% v/v PBS, RPMI with 10% FBS in 1 M HCl (pH = 6.4). As mentioned earlier, the solutions without nanoparticles served as negative controls. All nanoparticles were filtered under sterile conditions before mixing with the solutions in sterilized glass vials with a 0.45 µM pore diameter syringe filter (Merck Millipore Ltd., Darmstadt, Germany) and were left at RT for six months. The aggregation potential of nanoparticles was assessed by inspection at 1, 4, 12, and 24 weeks and by DLS measurements at 4 weeks.
Gel retardation assay (GRA)
The nanoparticles' siSCR loading and retention capacity, including GO, GOPEI, and Lipofectamine 2000TM (Lipo2000), were investigated with GRA. GO was mixed with siSCR at w/w ratios of 2:1, 4:1, 6:1, 8:1, 10:1, 12:1, 14:1, 16:1, 18:1, and 20:1 respectively, while GOPEI and Lipo2000 at a w/w ratios of 0.25:1, 0.5:1, 0.75:1, 1:1, 1.25:1, 1.5:1, 1.75:1, 2:1, 2.25:1, 2.5:1, respectively, while siSCR with sense 5′-UUCUCCGAACGUGUCACGUTT-3′ and antisense 5′-ACGUGACACGUUCGGAGAATT-3′ sequences (Suzhou GenePharma, Suzhou, China) served as the negative control in DNase, RNase free UltrapureTM Distilled Water (Invitrogen, Grand Island, NY, USA). The nanoparticles were left at RT for 30 min to complex with siSCR, then mixed with 5X RNA loading buffer (4 μL) in a total volume of 24 μL. The samples were loaded into an agarose gel (1%) (Sigma-Aldrich) prepared with 15 μL of 10,000 X Gel Red (Biotium Inc, Fremont, CA, USA) in 20X TAE Buffer (Thermo-Fisher Scientific, Carlsbad, CA, USA) and ran for 15 min at 120 V. The gel images were recorded with a ChemiDocTM MP Imaging System (Bio-Rad) or Gel Doc TM XR+ Molecular Imager (Bio-Rad) with the Image LabTM Touch v.2.3.0.07 and Quantity One v.4.6.8 software, respectively.
Serum stability
The siRNA cargo retention properties and the susceptibility of GOPEI complexed siSCR to RNase degradation were measured with a serum stability assay. Naked siSCR (0.66 µg), 50% FBS, 10%, 30%, 50%, 100% FBS dissolved in PBS, 100% FBS, 4 µg GOPEI (with 1X, 2X, and 3X of conjugated PEI concentrations) in pure FBS, DEPC water and pure FBS, 4 µg PEI (5 mg/mL) in DEPC water were investigated. The nanoparticle samples were complexed with siSCR at a 3:1 w/w ratio and kept at RT for 30 min to allow full complexation. Next, all samples were incubated at 37 °C and at predetermined time points of 0, 0.5, 1, 2, 2.5, 3, 6, 8, 12, 16, 24, and 48 h, respectively, a portion of the respective samples mixed with 5X RNA loading buffer (4 µL) (SolarBio, Beijing, China) was loaded into an agarose gel (1%) containing Gel Red (15 µL), and the samples were run for 10 min at 120 V. The gels were imaged with Gel Doc TM XR+ (Bio-Rad) Molecular Imager and analyzed with Quantity One v.4.6.8 (Bio-Rad) software. The stability of the nanoparticles complexed with FAM-labelled siSCR (siSCRFAM) with sense 5′-UUCUCCGAACGUGUCACGUTT3′ and antisense 5′-ACGUGACACGUUCGGAGAATT-3′ sequences (Suzhou GenePharma) in mouse serum was further measured in a fluorescent kinetic study. In a 96-well plate (Corning Inc., Corning, NY, USA) in triplicates, 50 µL of PBS (blank), 1:1 PBS with mouse serum, 1:1 PBS with mouse serum plus siSCR as negative control, GOPEI conjugated siSCR, and Lipo2000 complexed siSCR as positive control were measured. The fluorescence was recorded for 8 h at an excitation wavelength of 488 nm and emission wavelength of 520 nm using a Varioskan LUX multimode microplate reader and analyzed using Skanit Re v.4.1 (Thermo-Fisher Scientific) software.
In vitro assays
Cell culture
A2780 cisplatin (CP) sensitive and A2780DDP CP resistant human ovarian epithelial adenocarcinoma cell lines were a kind gift from Sun Yat-sen University Cancer Center (Department of Gynecology, Guangdong, China). A2780 and A2780DDP cell lines were grown in full media consisting of Dulbecco’s Modified Eagle Media (DMEM, HyClone Laboratories Inc, Logan, UT, USA) media supplemented with 10% FBS (Gibco, Thermo-Fisher Scientific, USA) and 1% penicillin/streptomycin. The cell lines were grown in a humidified incubator at 37 °C with 5% CO2. The cisplatin-resistant phenotype of A2780DDP was maintained by supplementing 20 µg/ml cisplatin (Sigma-Aldrich) every 4 weeks for 24 h. For cell passaging, the media was removed, and cells were washed twice with PBS, then 5 ml of 0.25% (w/v) trypsin-EDTA (Sigma-Aldrich) was added and incubated in 5% CO2 at 37 °C for 6–7 min. Next, 9 mL of full media was added. The cell suspension was centrifuged in a polypropylene conical centrifuge tube (15 mL) with a Sorvall™ ST 16 Centrifuge (Thermo-Fisher Scientific) at RT for 3 min at 800 rpm. Following removing the supernatant, cells were re-suspended and counted manually under an Eclipse TS 100 inverted microscope (Nikon, Japan) with a modified Neubauer hemocytometer using Trypan blue (Invitrogen) to quantify cell viability.
Live cell FAM uptake
Based on graphene-based nanomaterials' ability to quench specific fluorophores' fluorescence, siSCRFAM was complexed with GOPEI and GOPEI-mPEG, and the cellular uptake and siSCRFAM release in A2780DDP cells were measured at excitation and emission wavelengths of 488 nm and 520 nm, respectively. The nanoparticle samples were pre-complexed with siSCRFAM at RT for 30 min in triplicates in a final volume of 50 µL and were aliquoted into a 96-well plate, while DIW, RPMI media, nanoparticles only, and naked siSCRFAM served as negative controls. Under maintained CO2 levels (5%) and temperature (37 °C), the fluorescent signal was recorded for 12 h with a Varioskan LUX (Thermo-Fisher Scientific) plate reader, and the results were analyzed with the SkanItTM (Thermo-Fisher Scientific) software.
Laser confocal microscopy
A2780 and A2780DDP cells (5 × 104) were seeded in collagen-coated (d = 35 mm) gamma-irradiated glass bottom micro-well confocal dishes (MaTek Corporation, Ashland, MA, USA) and incubated overnight in a 5% CO2 incubator at 37 °C. siSCRFAM (2 µg) dissolved in Opti-MEM (500 µl) reduced serum media (Gibco, Carlsbad, CA, USA) served as the negative control, GOPEI (6 µg) and GOPEI-mPEG (6 µg) were complexed with siSCRFAM (2 µg) at w/w ratio of 3:1 for 30 min, and 15 µg of Lipo2000 (1 mg/mL) (Thermo Fisher) as a positive control was complexed with siSCRFAM (2 µg) for 30 min, Lipo2000 and GOPEI without siSCRFAM were used as negative controls, and cells without treatment as blank controls. Next, the media was removed following overnight incubation with the test samples, the dishes were washed with ice-cold PBS thrice, and serum-free RPMI media (500 µL) was added. The confocal dishes were completely covered with aluminium foil and incubated for 5 h in a 5% CO2 incubator at 37 °C. Next, the cells were washed with ice-cold PBS twice and fixed with formaldehyde (3.7%) for 10 min. For cell permeabilization, Triton X-100 (0.2%) (Sigma-Aldrich, St. Louis, MO, USA) was added for 5 min, then washed with PBS thrice. The nuclei of the cells were stained with DAPI (1 mg/ml) nuclear stain (Sigma-Aldrich, St. Louis, MO, USA) dissolved 1:5000 in PBS. Following a 20 min incubation at 40 °C in the dark, the cells were washed with PBS thrice. They were imaged in confocal mode with a Zeiss laser confocal scanning microscope at 10X and 40X resolution objectives. The excitation wavelengths for DAPI and siSCRFAM were 405 and 488 nm, while the emission was recorded in the 460–490 nm and 500–530 nm wavelengths, respectively. The obtained images were analyzed with the Zeiss LSM 880 (Zeiss, Oberkochen, Germany) and ZEN Imaging software 2.3 (Zeiss).
Fluorescence-activated cell sorting (FACS)
The siRNA transfection efficiency of the nanoparticles was quantified with FACS. A2780DDP cells (3 × 105/well) were seeded in six-well plates and incubated overnight prior to transfection. The plates were transfected with GOPEI (0.4, 0.8 and 1.2 µg) complexed siSCRFAM (30, 60, 90 nM) at a w/w ratio of 3:1, Lipo2000 (positive control) complexed siSCRFAM (30, 60, 90 nM), GO (22 µg) complexed siSCRFAM (90 nM), and blank (media only), naked siSCRFAM, GOPEI and Lipo2000 served as negative controls. To assess the siRNA transfection efficiency of GOPEI-siSCRFAM complexes at 0.2:1, 0.8:1, 1:1, 2:1, 5:1, and 10:1 w/w ratios, GOPEI was mixed with siSCRFAM (40 nM) at RT for 30 min. For transfection, the media was first replaced with serum-free DMEM (500 µL), and the test samples suspended in Opti-MEM (500 µL) were added to the wells dropwise. Following transfection, the plates were incubated at 5% CO2 at 37 °C for 5 h, then the media was replaced, and the plates were incubated for at least 48 h until they reached 80% confluency. The adherent cells were harvested by trypsinization and centrifugation and were re-suspended in ice-cold PBS (1 mL). The fluorescence of the cells was measured with a FACSCalibur (BD Biosciences, Franklin Lakes, NJ, USA) flow cytometer by analyzing 10,000 events per measurement. Cells were isolated according to their granularity and size predetermined by the Side Scatter (SSC-H), Forward scatter (FSC-H), and green FAM fluorescence band pass filter (FL1-525 nm) parameters with a BD CellQuest Pro (BD Biosciences) software, and the results were analyzed with the FlowJo software (BD Biosciences).
Quantitative real-time polymerase chain reaction (qRT-PCR)
The baseline and time-course SOD1 mRNA expression levels at 4, 8, 16, 24, 36, and 48 h and the knockdown efficiency of GOPEI and GOPEI-mPEG complexed siSOD1 (1–300 nM) at different w/w ratios (0.2:1 to 10:1) along with the effect of GOPEI treatment (1–8 µg/mL) alone on the SOD1 mRNA expression levels in A2780, and A2780DDP cells and the activation of the UPRmt pathways were measured with qRT-PCR. Cells (5 × 105/well) were seeded in 6-well plates (Corning), incubated for 24 h then transfected with nanoparticle-siRNA complexes for 48 h. Next, the media was removed, and the cells were lysed with 1 mL TRI Reagent (Sigma-Aldrich) for 10 min. The cell lysate was mixed with chloroform (200 µL) and then centrifuged at 12,000 × g for 15 min at 4 °C. The supernatant above the phenol-chloroform fraction was removed, and 2-propanol (500 µL) was added and centrifuged at 12,000 × g for 10 min at 4 °C. Next, the supernatant was discarded, and the pellet was washed with 75% ethanol (1 mL) and centrifuged at 7500 rpm for 5 min at 40 °C. The RNA-containing pellet was air-dried, re-suspended in DEPC water (20 µL), and then incubated for 15 min at 56 °C. The RNA concentration was measured with Nanodrop 2000C (Thermo-Fisher Scientific) using the Nanodrop 2000C v.1.4.1 software (Thermo-Fisher Scientific). A GoScriptTM Reverse Transcription Kit (Promega Corporation, Madison, WI, USA) was used for cDNA synthesis. Total RNA (500 ng) was annealed with Oligo(dT15) primers at 70 °C for 5 min, then cooled immediately in ice water for 5 min and centrifuged for 10 s. SOD1 mRNA level was quantified using the (forward) 5′-ATCCTCTATCCAGAAAACACGG and 5′-GCGTTTCCTGTCTTTGTACTTT (reverse), while β-actin housekeeping gene with 5′-CACCATTGGCAATGAGCGGTTCC-3′ (forward) and 5′-GTAGTTTCGTGGATGCCACAGG-3′ (reverse) primer sequences (Sangon Biotech, Shanghai, China) was used as an internal reference. The samples were mixed with 4.0 µL GoScript™ 5X Reaction Buffer (Promega), 3 nM MgCl2 (3 µL), 1 µl GoScript™ Reverse Transcriptase, 1 µL PCR Nucleotide Mix (0.5 mM each dNTP), and nuclease-free water (6 μL). The samples were annealed at 25 °C for 5 min, extended at 42 °C for 1 h then incubated at 70 °C for 15 min. The cDNA samples were diluted 10-fold and combined with the GoTaq qPCR Master Mix kit (Promega). A final reaction volume was 20 µL consisting of 2X GoTaq qPCR master mix (10 µl), 10 nM reverse and forward SOD1 and β-actin primers (1 µL each), cDNA sample (2 µL), and nuclease-free water (6 µL). The PCR cycling parameters were as follows: activation (95 °C, 1 cycle, 2 min), denaturation (95 °C, 15 s), annealing (40 cycles), and extension at 95 °C (1 min). The pentuplet samples were run under SYBR Green setting with an Applied BiosystemsTM Quantstudio-5 (Thermo-Fisher Scientific) qRT-PCR machine. The results were analyzed with the Quantstudio-5 Design and Analysis v1.4.3 (Thermo-Fisher Scientific) software using the ∆∆Ct method [33].
Western blot (WB)
The baseline protein expression levels of SOD1, the knockdown efficiencies of GOPEI and GOPEI-mPEG complexed siSOD1 (1, 5, 10, 15, 20, 25, and 30 nM, respectively) at various w/w ratios (0.2:1 to 10:1), and the effect of GOPEI (1–8 µg/mL) treatment alone in A2780 and A2780DDP cells were quantified with WB. Three SOD1-specific siRNAs were tested for their knockdown efficiencies: siSOD1-homo-D-333 with sense 5′-CCUCACUUUAAUCCUCUAUTT-3′ and antisense 5′-AUAGAGGAUUAAAGUGAGGTT-3′ sequences, siSOD1-homo-D2757 with sense 5′-CGACGAAGGCCGUGUGCGCTT-3′ and antisense 5′-UCGCACACGGCCUUCGUCGTT-3′ sequences, siSOD1-homo-D2861 with sense 5′-CCCUUAACUCAUCUGUUACTT-3′ and antisense 5′-UUAACAGAUGAGUUAAGGGTT-3′ sequences. All three siRNAs were acquired from Suzhou GenePharma (Suzhou, China). The cells (3 × 105/well) were plated in six-well plates (Corning) and incubated for 24 h in 5% CO2 at 37 °C prior to transfection with siSOD1 complexed nanoparticle samples or nanoparticles only using the same method as for FACS. After 48 h, the media was removed, and the plates were washed twice with ice-cold PBS. For protein extraction, 1 X RIPA buffer (200 µL) (150 mM NaCl, 20 mM Tris-HCl, pH 7.5, 1 mM EGTA, 1 mM Na2EDTA, 2.5 mM sodium pyrophosphate, 1% NP-40, 1% sodium deoxycholate, 1 mM Na3VO4, 1 mM β-glycerophosphate, and 1 µg/mL leupeptin) supplemented with 1X protease inhibitor cocktail (Roche, Basel, Switzerland) in a 1:25 v/v ratio. After determining the protein concentration with a BCA Protein Assay kit (BeyoTime, Shanghai, China), the samples were transferred to a 95 °C heat block for 10 min. Next, the respective protein samples (20 µg) were loaded into 10% polyacrylamide 12 or 15-well precast gels (GenScript, NJ, USA) and ran for 90 min at 120 V in Tris-MOPS-SDS running buffer. The gel was transferred with an eBlot™ L1 Fast Wet Transfer System (GenScript) onto a PVDF membrane, then incubated in blocking buffer (Beyotime, Shanghai, China) for 1 h at RT. The membrane was incubated with mouse anti-human SOD1 (15.9 kDa) monoclonal antibody (Cat# 67480-1-Ig, 1:1000, ProteinTech, Rosemont, IL, USA) and with mouse β-actin (42 kDa) monoclonal antibody (1:2000, Cat# 66009-1-Ig, ProteinTech) overnight at 40 °C. Next, the membrane was washed with 1X TBST thrice for 5 min and incubated with donkey anti-mouse IgG (H + L) IRDye® 800CW (LICOR Biosciences, NE, USA) polyclonal antibody at RT for 1 h, then washed with 1X TBST thrice for 5 min each. Protein bands were displayed by an Odyssey Imaging System (LI-COR Biosciences) and analyzed by the Li-Cor Odyssey 3.0.29 (LI-COR Biosciences) and Image-J (NIH) software.
MTT assay
The cisplatin sensitivity (IC50) of A2780 and A2780DDP cells in a range of 2–28 µg/mL cisplatin (Sigma-Aldrich) concentration, the cytotoxicity of graphene-based nanocarriers, and the cisplatin chemo-sensitizing effect of nanoparticle complexed siSOD1 (D-333) at different concentrations (1–300 nM), and w/w ratios (0.05:1–10:1) upon cisplatin treatment were first determined with MTT assay before confirming it with clonogenic assay. A2780 and A2780DDP cells (3 × 103 cells/well) in 100 µL complete media were seeded into 96-well plates and incubated for 24 h. Next, cells were treated with the respective drug compound, nanoparticle, or both in serum-free media (50 µL per well) and incubated in 5% CO2 at 37 °C for different durations (48–72 h). Next, 15 µL of 5 mg/ml MTT [3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide] (Sigma-Aldrich) solution was added to each well. The plates were transferred onto a plate shaker, shaken for 5 min at 600 rpm, and then incubated in 5% CO2 at 37 °C for 5 h. The formazan crystals were dissolved by adding 135 µL of 10% acidified SDS (Thermo-Fisher Scientific) and further shaken at 500 rpm for 15 min. The absorbance was recorded at 570 nm, and the results (9–18 replicates) were compared to a blank (media only), negative control (cells only), and positive death control (cells treated with Triton X-100 (1%) and analyzed by Varioskan LUX, Skanit Re v.4.1 (Thermo-Fisher Scientific) software. The relative cell viability was calculated as follows:
Cell viability (%) = (OD test sample – OD blank - OD positive control) / (OD negative control – OD blank - OD positive control) × 100%.
Population Doubling
To evaluate the effect of SOD1 knockdown on the proliferation rate, A2780, A2780DDP cells (3 × 103/well) in 100 µL, complete media were seeded into 96-well plates and incubated for 24 h. Next, cells were treated with the respective drug compound, siRNA-nanoparticles, or both in serum-free media (50 µl per well) and incubated in 5% CO2 at 37 °C for 4 days. From day 0 to day 4, one 96-well plate was treated with MTT solution, and the absorbance readings were taken at 24 h, 48 h, 72 h, and 96 h post-seeding, respectively. The total normalized absorbance values of the respective plates were used to plot a cell growth curve. Population doubling-time calculations: Population doubling times were calculated using the slope of the angle of the linear regression analysis of the 5 time points and confirmed with the Doubling Time Software v1.0.10 (http://www.doubling-time.com).
Clonogenic Assay
The cisplatin sensitivity and chemo-sensitizing effects of GOPEI-siSOD1(D-333) and GOPEI-mPEG-siSOD1(D-333) were determined with a clonogenic cell assay. A2780 and A2780DDP cells (~1 × 103 cells/well) were seeded in 6-well plates and then incubated overnight with 5% CO2 at 37 °C. Next, the different drug compounds, siRNA-nanoparticles, or both dissolved in RPMI (1 mL) media and added to the cells, then following 2–5 h of incubation, depending on the respective treatment, the media was replaced with full media, and the plates were incubated for 10–12 days. The colonies were fixed with methanol (25% v/v) for 1 min, stained with crystal violet (0.05% w/v) for 5 min and washed under running tap water. Colonies (min >50 cells) were counted manually, and the relative cell viability of the treatment groups was expressed relative to the control group.
In vivo study
Ethical approval and animal handling
Animal study protocols (ASP) were approved by the Institutional Animal Care and Use Committee (IACUC) of Suzhou GenePharma (Suzhou, China), and all in vivo experiments were conducted according to IAUCC guidelines. The animals used in this study were housed in the facility of Suzhou GenePharma, an AAALAC internationally accredited research centre. All in vivo experiments were conducted according to the Guide for the Care and Use of Laboratory Animals. Female Balb/c 6–8 weeks old nude mice were exposed to 12 h light/dark cycles, kept in specific pathogen-free conditions in filter-topped cages, and fed with standard rodent chow plus water ad libitum. In this study, mice were sacrificed when any tumour volume reached more than 2000 mm3, upon any visible distress expressed by the animals, or more than a 20% decrease in body weight.
Preparation of ovarian cancer cells
A2780 cisplatin-sensitive and A2780DDP cisplatin-resistant ovarian cancer cells limited to maximum 10 passages were grown in T-75 flasks and harvested following sterile techniques in a biological safety cabinet in the logarithmic growth phase at 60–70% confluency. The full cell culture media (DMEM supplemented with 10% FBS) was aspirated and washed thrice with PBS (37 °C). Next, trypsin/EDTA (4 mL) was added, and the plates were incubated at 37 °C in 5% CO2 for 7 min. The cells were dislodged by gentle plate tapping, and the detached cells were visualized under an inverted microscope. Complete cell culture medium (10 mL) was added to inactivate the trypsin solution, and a single cell suspension was obtained by vigorous mixing. The live cells were counted by mixing 100 μL of the cell suspension with 100 μL of trypan blue and counted in a hemocytometer. The cell number was confirmed by combining 100 μL of the cell suspension in 9900 μL of CASY TT Buffer and counted with the CASY TT automated cell counter (OMNI Life Science GmbH & Co. KG, Bermen, Germany). Next, the single cell suspension (13.8 mL) was transferred to a centrifuge tube (15 mL) and centrifuged at 1000 rpm for 3 min at RT. The supernatant was discarded, and the cells were reconstituted in serum-free media at concentrations of 1 × 106, 5 × 106, or 1 × 107 cells mixed with 50% matrigel per mL as per the experimental design for subcutaneous tumour inoculation experiments. The cells were inoculated in the mice subcutaneously within 30 min of preparation.
Nanoparticle hemocompatibility assay
To obtain the red blood cell (RBC) fraction, anticoagulated fresh mouse blood (5 mL) was centrifuged for 10 min at 2000 × g, then washed thrice with PBS. Next, 2% (v/v) RBC suspension buffer was prepared in PBS mixed with 5 concentrations of naked siSCR, GO-siSCR, GOPEI-siSCR, GOPEI-mPEG-siSCR, and Lipo2000-siSCR in a total volume of 100 μL added to 900 μL of 2% RBC suspension. The samples were incubated at 37 °C for 1 h, centrifuged for 10 min at 3000 × g, and the absorbance was measured at 540 nm using a SpectraMax i3 plate reader.
In vivo tumour therapeutic efficacy study of GOPEI-mPEG-siSOD1
To evaluate the therapeutic effect of SOD1 knockdown in vivo, Balb/c nude xenograft mice were injected with cisplatin, naked siSOD1(D-2861)+cisplatin, GOPEI-mPEG-siSCR (control)+cisplatin, and GOPEI-mPEG-siSOD1(D-2861)+cisplatin (n = 3), respectively, at 0.5 mg/kg siRNA-nanoparticles dose. During a 14-day long therapeutic window, mice received 6 doses of respective siSCR or siSOD1(D-2861)-based treatments with or without cisplatin treatment. Tumour sizes were measured by caliper every 2–3 days.
In vivo toxicity and blood biochemistry testing
The potential acute in vivo toxicity of the nanoparticles was evaluated with blood biochemistry testing. Balb/c nude mice were injected with siSCR, GO-siSCR, GOPEI-siSCR, GOPEI-mPEG-siSCR (n = 3) at a 0.5 mg/kg siRNA dose. Blood samples were collected 8 h after the drug administration. Whole blood samples without anticoagulants were centrifuged at 3000 × g at 4 °C for 10 min to obtain the serum fraction. Haematological testing was performed using an automatic haematology analyzer (MEK6400, Nihon Kohden, Japan) for serum alanine aminotransferase (ALT), aspartate aminotransferase (AST), alkaline phosphatase (ALP), total protein (TP), albumin (ALB), creatinine (CREA), uric acid (UA), blood urea nitrogen (BUN) lactate dehydrogenase (LDH) and amylase (AMY).
Statistical analysis
Unless stated otherwise, all experiments were carried out in triplicates (n = 3). The in vitro numerical data were analyzed for statistical significance with Student’s two-tailed paired and unpaired t-tests and were expressed as mean ± SD using GraphPad Prism 7.04 (GraphPad Software, San Diego, CA. USA). In the study design, p < 0.05 was considered statistically significant and was denoted as *, **, ***, and **** for less than 0.05, 0.01, 0.001 and 0.0001, respectively.
Results
Graphene nanoparticle preparation
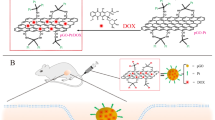
Graphene-oxide (GO) was prepared according to a previous study (Fig. 1A) [32]. Subsequently, polyethyleneimine (PEI) was conjugated onto the GO sheets using EDC chemistry, yielding a GO-PEI (GOPEI) nanocarrier. Further, to increase biocompatibility and decrease surface charge, GOPEI was directly PEGylated on the primary amine groups of PEI (GOPEI-mPEG), allowing for a tunable surface charge configuration. TEM images of GO and GOPEI showed distinct hexagonal lattice structure of GO (Fig. 1B) and a homogenous, well-dispersed nanoparticle distribution exhibiting rough, wrinkled morphological domains of graphene sheets (Fig. 1C, D). A high-resolution image of GOPEI revealed darker sheet intensities relative to GO due to PEI functionalization (Fig. 1D). AFM imaging revealed two-dimensional sheet-like morphology with highly variable particle thickness and a lateral size ranging from 0.05 to 0.40 μm. (Fig. 1E, F) Following PEI conjugation, the sheet thickness of GOPEI increased by ~3–4 nm relative to GO. (Fig. 1G, H) The latter was further confirmed with z-section analysis showing darker sheet intensities on 3D topographic analysis of the GOPEI sample attributed to increased flake thickness due to PEI functionalization (Fig. 1I, J).
A Schematic representation of GOPEI-mPEG preparation. mPEG was directly conjugated to PEI instead of graphene, which created a tunable system to control the ζ-potential of the nanoparticle. Linear methyl polyethylene glycol (mPEG) was conjugated to GOPEI through EDC chemistry, and siRNA molecules were allowed to bind GOPEI-mPEG electrostatically. B, C High-resolution TEM images of the surface morphological characterization of GO at different scales (5 nm, 200 nm, and 50 nm, respectively) showing a well-dispersed homogenous nanoparticle solution with varying sizes and dimensions. D Single GOPEI nanoparticle with rough and wrinkle surface characteristics. AFM analysis of GO and GOPEI surface morphology (E, F), lateral thickness (G, H), and topographical 3-D images (I, J) respectively. AFM analysis showed an even distribution of GO flakes with lateral dimensions in the 0.05–0.4 μm range and thickness of 1–2 layers of GO and GOPEI.
Graphene nanoparticle characterization
Next, we confirmed whether the conjugation of -COOH groups of GO to the primary amines of PEI occurred using FT-IR spectroscopy (Fig. 2A). A characteristic peak of amide bonding was observed at 1630–1695 cm−1 of GOPEI but was absent in GO samples [34]. A broad peak at 1705–1730 cm−1, characteristic of carboxyl groups, and further peaks at 1200–1300 cm−1 and 3200–3500 cm−1 corresponding to C=O and –OH groups were appreciable of GO. We further measured the composition of GOPEI using TGA. The thermogravimetric curve of GOPEI (Fig. 2B) showed a biphasic decomposition pattern relative to GO. Both samples exhibited rapid initial weight loss (<150 °C) attributed to remnant water evaporation. However, the mass loss of GOPEI in the 250–350 °C range increased considerably compared to GO due to PEI25 kDa decomposition (Td PEI25 kDa ~ 200 °C). Based on these results we concluded that PEI25 kDa makes up ~70% of the total mass of GOPEI. Next, the absorbance spectrum (Fig. S1A) and electrostatic adsorption of siSCR cargo onto GOPEI (Fig. S1B) were evaluated with UV–Vis spectroscopy. The siSCR complexed GOPEI showed higher and superimposed (GOPEI + siSCR) absorbance spectrum compared to their individual peaks indicating electrostatic complexation of the siSCR cargo with the GOPEI carrier (Fig. S1B). In addition to conventional TEM- and AFM-based graphene characterisation, the nanoparticle size distribution was periodically monitored before in vitro assays for potential aggregation using DLS to lower batch-to-batch variations and to increase experimental reproducibility. Next, for our reference, we established that the prepared GO had an average size distribution of 2354.6 ± 423 nm (Fig. 2C) and a low ζ-potential of −40.4 ± 0.4 mV (Fig. 2D), showing high size heterogeneity in solution with a PDI of 0.81 ± 0.11 (Fig. 2E). Following the activation of the –COOH groups with NaOH, GO exhibited lower hydrodynamic size distribution (318.7 ± 2.7 nm), and lower ζ-potential (−48.1 ± 0.4 mV), and the carboxylation lowered (p < 0.05) the polydispersity of the graphene sheets (0.263 ± 0.04). Different batches of GOPEI corresponding to various PEI concentrations showed increasing hydrodynamic sizes (p < 0.01) (206.0 ± 2.7 nm, 274.8 ± 7.2 nm and 347.4 ± 1.2 nm) and stable ζ-potential between +50–60 mV (55.2 ± 2.4 mV, 50.1 ± 5.1 mV and 57.3 ± 0.8 mV) and PDI (0.159 ± 0.01, 0.267 ± 0.01 and 0.247 ± 0.01) (Fig. 2C–E). GOPEI-mPEG exhibited similar physicochemical properties as GOPEI with a reduced lateral size of 129.3 ± 0.1 nm (p < 0.0001), and decreased ζ-potential (16.8 mV, p < 0.0001), and no significant change (p > 0.05) in PDI was appreciable following mPEG conjugation. Further, we confirmed that the hydrodynamic diameter of the graphene-siRNA polyplexes changed by modulating the GOPEI-mPEG to siRNA weight-to-weight (w/w) complexation ratios (Fig. 2F). Finally, the loading capacity, electrostatic adsorption, and retention of siSCR of the nanoparticles were assessed by agarose gel retardation assay (Fig. 2G). Relative to the control naked siSCR bands, the nanoparticle complexed siSCR groups showed lower band intensities indicating partial or total siSCR loading and retention at different w/w ratios. GOPEI nanoparticles containing increased amounts of PEI fully complexed siSCR at w/w ratios of 2:1, 1.25:1, and 0.75:1, respectively, while GOPEI-mPEG fully complexed the siSCR cargo at 1.25:1 w/w ratio. However, non-functionalized GO failed to completely retain siSCR, as evidenced by the presence of free siSCR bands at high w/w ratios. However, GO showed partial retention of siSCR, which is consistent with prior literature [35]. These findings were further validated in the last well of each nanoparticle group, where the nanoparticle to siSCR complexation ratio was 1:2, and the band intensity was comparable to the 0.5:1 group.
A FT-IR spectra of GO and increasing concentrations of cationic polymer functionalized GOPEI1x, GOPEI2x, GOPEI3x. Conjugation of the carboxyl group of GO and the amino group of PEI was confirmed by FT-IR spectroscopy by measuring the characteristic peak of the amide bonds at 1630–1695 cm−1. B TGA spectra of GO and GOPEI was measured in the temperature range of 25-800 ˚C. The decomposition of components of the nanoformulation brought about by the mass loss at increasing temperatures was used to estimate the PEI content in GOPEI. (C–E) DLS analyses of (C) hydrodynamic size, (D) ζ-potential, and (E) PDI of the nanoparticles GO, GOPEI1x, GOPEI2x, GOPEI3x, and GOPEI-mPEG. The particle sizes were 2354.6 ± 423, 318.7 ± 2.7, 206.0 ± 2.7, 274.8 ± 7.2, 347.4 ± 1.2, and 112 ± 1.2 nm, respectively. The ζ-potential were −40.4 ± 0.4, −48.1 ± 0.4, 55.2 ± 2.4, 50.1 ± 5.1, 57.3 ± 0.8, and 16.1 ± 0.6 mV, respectively. F In vitro evaluation of GOPEI-siSCR aggregation at different ratios. A DLS study measuring the hydrodynamic size of GOPEI-siSCR at different w/w ratios was carried out (n = 3) at the following ratios: 0.2:1, 0.5:1, 1:1, 2:1, 3:1, and 5:1. G Agarose gel retardation assay of the siSCR with GO and Cadmium quantum dots (CdQD) nanoparticles, respectively. GO and CdQD at various weight ratios of 2:1, 4:1, 6:1, 8:1, 10:1, 12:1, 14:1, 16:1, 18:1 and 20:1. Lipo2000, GOPEI1x, GOPEI2x, GOPEI3x and GOPEI-mPEG used w/w ratios of 0.25:1, 0.5:1, 0.75:1, 1:1, 1.25:1, 1.5:1, 1.75:1, 2:1, 2.25:1 and 2.5:1. Naked siSCR was used as negative control. The red frame represents the best complexation w/w proportion. All values are expressed as mean ± SD. ns- not significant and **p < 0.01 as analyzed using a two-tailed unpaired t-test.
Evaluation of in vitro biocompatibility
The aggregation potential of nanoparticles in biological solutions was observed and measured over 1 h, 24 h, 1, 3, and 6 months (Fig. 3A). We found that GO immediately precipitated and formed visible aggregates upon mixing with 0.9% NaCl, PBS, RPMI (10% FBS), and FBS (50%) solutions that persisted during the observation period (6 months) (Fig. S1C). GOPEI, however, did not show any observable aggregation and remained a homogenous solution even after 1 month of incubation in DIW, PBS, RPMI (10% FBS), FBS (50%), NaCl (0.9%) or RPMI (10% FBS, pH: 6.4) at RT that aimed to simulate the microenvironment of solid tumours. Further, the serum stability of GOPEI complexed siSCR evaluated against RNase degradation showed adequate complexation and protection of the siRNA cargo by the prepared nanoparticles in high serum-containing solutions (Fig. 3B, C). While naked siSCR completely degraded and its band disappeared after 6 h, GOPEI nanoparticles showed no bands after complexation with siSCR throughout the study even after 48 h in 50% FBS-containing solution, which simulated human plasma compared to negative control samples (Fig. S1D).
A Colloidal stability of GO, GOPEI, and GOPEI-mPEG were assessed in DIW, 0.9% NaCl, and RPMI + 10% FBS, 1 h after treatment. Nanoparticle aggregation was observed in all three biological solutions containing non-functionalized GO. B, C Gel-electrophoresis-based serum stability was carried out in GOPEI1x, GOPEI2x, and GOPEI3x. The susceptibility of siRNA to RNase degradation in serum was assessed for 48 h. Naked siSCR was used as a control. D Kinetic study of nanocarrier-siRNA serum stability was measured using mouse serum in siSCRFAM conjugated GOPEI, GOPEI-mPEG, and Lipo2000 relative to control naked siSCRFAM. The fluorescence of siSCRFAM was measured over a time of up to 8 h. E Kinetic study of nanoparticle behaviour was conducted in vitro using siSCRFAM complexed GO, GOPEI and GOPEI-mPEG. The degree of quenching induced by GOPEI upon binding siSCRFAM was measured by a Varioskan fluorescent plate reader. A 10 h time-course study of the live cell uptake of the nanoparticles was measured by recording their fluorescent emission. Samples containing only cells and RPMI media were used as negative controls. F Confocal laser scanning images showing the cellular uptake of GOPEI- siSCRFAM. Merged fluorescence images of cells treated with GOPEI- siSCRFAM with a magnified image of a single A2780 cell showing successful transfection of GOPEI as aggregations were compared with Lipo2000 as the positive control, and naked siSCRFAM was used as a negative control. G Quantitative assessment of nanoparticle: siRNA cellular uptake GOPEI-siRNA transfection efficiency was analyzed using flow cytometry. Two siSCRFAM concentrations of 60 nM and 90 nM complexed with Lipo2000 served as positive control. H Image showing intracellular graphene accumulation in cell pellets following three washing steps and centrifugation prior to protein isolation. I, J Time-course study showing SOD1 mRNA and protein levels following SOD1 knockdown. K, L The effect of concentration dependent SOD1 knockdown on protein levels was determined by Western blot and mRNA expression levels were measured using RT-qPCR. All values are expressed as mean ± SD. ns- not significant, *p < 0.05, **p < 0.01, and ****p < 0.0001 as analyzed using two-tailed unpaired t-test.
The agarose gel-based serum stability study results were further validated with a fluorescent kinetic study using mouse serum (Fig. 3D). The emitted fluorescence of nanoparticles complexed with siSCRFAM samples were recorded for 8 h. The negative control naked siSCRFAM group showed a rapid drop in fluorescence after 3 h and completely disappeared after 6 h, attributed mainly to degradation by RNAses of the mouse serum, which is in line with the results of the gel-based assay stability study. The GOPEI and GOPEI-mPEG complexed siSCRFAM samples steadily showed low fluorescence during the 8 h long study, indicating that a negligible amount of siSCRFAM was released from the nanoparticles during incubation compared to the positive control Lipo2000 group. Overall, we established that the prepared GOPEI and GOPEI-mPEG nanoparticles could provide sufficient protection for the siRNA cargo for a minimum of 4 h and were deemed suitable for further in vivo experiments.
Next, we evaluated the kinetic cellular uptake of the nanoparticles in a kinetic in vitro assay (Fig. 3E). During a 10 h time-course study using live cells, we observed increased siSCRFAM fluorescence after 6 h for GOPEI and GOPEI-mPEG nanoparticles but not for naked siSCRFAM. We attributed this increase in siSCRFAM fluorescence to the endosomal escape of the nanoparticles mediated by the proton sponge effect of PEI25 kDa in the acidic lysosomal environment, since under physiological and buffered pH conditions, our serum stability assay showed (Fig. S1D), that GOPEI managed to fully complex and prevent releasing the siSCR cargo for an extended period of 48 h [36]. The qualitative intracellular uptake of GOPEI-siSCRFAM complexes was also investigated by laser confocal microscopy (Fig. 3F, Fig. S1E). A2780 cells, in addition to control groups, were transfected naked siSCR, GOPEI, Lipo2000, GOPEI-siSCRFAM, and Lipo2000-siSCRFAM to measure the intracellular fluorescence following successful siRNA cargo delivery by GOPEI to the cells. GOPEI-siSCRFAM and Lipo2000-siSCRFAM showed evident green fluorescence in the cytoplasm, while for naked siSCRFAM no fluorescence was detectable. Similarly, cells treated with media or nanoparticles only (GOPEI or Lipo2000) showed no fluorescence. Finally, the quantitative cellular uptake of nanoparticle complexed siSCRFAM polyplexes were measured with flow cytometry (Fig. 3G and S1F). Naked siSCRFAM was partially taken up by A2780 cells, and unfunctionalized GO showed minimal siRNA delivery capability. In contrast, GOPEI complexed siSCRFAM (60, 90 nM) at a 3:1 w/w ratio showed significantly increased fluorescence (43.89% and 63.21%, p < 0.0001) comparable to the positive control Lipo2000 group with 66.00% and 66.20% using 60 and 90 nM siSCR, respectively. In addition, cellular uptake of the graphene-based nanoparticles was evident during protein isolation, as cell pellets showed evident dark discolouration attributed to the presence of intracellular graphene nanoparticles (Fig. 3H). GOPEI complexed with 90 nM siSCRFAM showed the highest fluorescence and thus was selected for further experiments. Finally, we determined that GOPEI-mPEG-siSOD1 can efficiently silence SOD1 expression on both mRNA (Fig. 3I) and protein (Fig. 3J) levels. In addition, time-course evaluation of SOD1 silencing showed potent downregulation of mRNA (Fig. 3K) and protein levels (Fig. 3L) after 24 and 72 h, respectively. We have also used the other three different cell lines, HeLa, A549, and Hep3B, to confirm the siRNA’s ability to downregulate SOD1 expression at the protein level (data not shown).
Evaluation of in vitro characteristics in cell-based assays
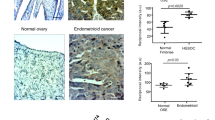
In the present study, to model the development of acquired cisplatin resistance, we first exposed human epithelial A2780 ovarian cancer cells to low-dose cisplatin (1 µM) for 3 months (Fig. 4A). Next, the degree of acquired cisplatin resistance induction was determined in the A2780 and resistant pair of A2780DDP cells (Fig. 4B, C), and we observed a ~3-fold increase in cisplatin IC50 with MTT assay (p = 0.0007), that we confirmed with clonogenic assay (Fig. S1G, H). To establish causality between acquired cisplatin resistance and SOD1 overexpression in vitro, we first determined the baseline SOD1 mRNA and protein levels between the cell lines (Fig. 4D, E). Consistent with our previous findings, A2780DDP exhibited a 1.61-fold increase in SOD1 mRNA (Fig. 4D) and a 1.3-fold increase in protein levels in vitro (Fig. 4E). The same cell line pair showed consistently increased (p > 0.001) SOD1 mRNA (Fig. 4F) while a notable 1.9-fold increase in protein levels in vivo (Fig. 4G) in four independent tumour quadrants (Fig. 4H). Interestingly, however, when measured in two outers and two inner tumour quadrants, we observed inhomogenous and fluctuating SOD1 mRNA expression levels between the respective quadrants, that we mechanistically validated on the protein levels in an independent study (unpublished observation). Based on these findings, we concluded that in our model, acquired cisplatin resistance is associated with SOD1 overexpression both in vitro and in vivo. To establish causality between cisplatin treatment and SOD1 overexpression, we measured the change of SOD1 mRNA levels upon a range of 0–30 µM cisplatin treatment. We found that cisplatin treatment elicited significant SOD1 mRNA overexpression (p < 0.0001) (Fig. 4I). In addition, we observed that GOPEI solely (1–8 µg/mL) could significantly (p < 0.001) increase SOD1 mRNA levels above 4 µg/mL attributed to the combined effect of oxidative stress induction by PEI and mechanical shear stress elicited by the GO sheets (Fig. 4J). Next, we evaluated the in vitro cytotoxicity of the nanocarriers and their effect on cell viability using the MTT assay (Fig. 4K). GO treatment showed relatively high cell viability (80%) even at 40 µg/ml concentration. In contrast, free PEI25 kDa, as anticipated, showed concentration-dependent toxicity and caused complete cell death (p < 0.0001) at or above 20 µg/mL concentration. As per our previous findings, the conjugation of GO with PEI significantly increased cell viability compared to free PEI alone (p < 0.0001) [34], while subsequent PEGylation further decreased cellular toxicity. In addition, combinational treatment with GOPEI (9 µg/mL) with cisplatin (2–28 µg/mL) resulted in a considerable decrease in cisplatin IC50 (7.71–6.24 µg/ml) (Fig. 4L) relative to the GOPEI (9 µg/mL) treated control. However, we attributed this chemosensitization effect not to SOD1 knockdown but rather the toxicity of PEI, which evidently necessitated the PEGylation of the nanoparticle.
A Schematic diagram of the in vitro induction of acquired platinum resistance. B Baseline cisplatin sensitivity of A2780 and A2780DDP cell lines. C The IC50 of A2780 and A2780DDP were 4.71 ± 0.26 and 13.95 ± 1.18 µg/ml, respectively. D Bar graph showing normalized baseline SOD1 mRNA levels (n = 6). E Western blot showing the SOD1 protein levels in A2780 and A2780DDP cell lines in vivo in Q1 and Q3. F Bar graph showing normalized baseline SOD1 mRNA (n = 3). G Schematic illustration of xenograft tumour sample preparation for immunoblotting and qRT-PCR. H Western blot showing the SOD1 protein levels in A2780 and A2780DDP cell-derived xenograft tumour tissue samples. I Cisplatin treatment of A2780DDP cells (n = 6) induced SOD1 mRNA overexpression in a concentration-dependent manner measured by RT-qPCR. J Time-course evaluation SOD1 mRNA induction by GOPEI treatment in A2780DDP cells (n = 6) measured by qRT-PCR. K Cytotoxicity of GO, PEI, GOPEI and GOPEI-mPEG were determined at the following concentrations: 2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 25, 30, 35 and 40 µg/mL, respectively. L MTT assay showing relative cell viability after A2780 cells were treated with 9 µg/mL of GOPEI for 48 h followed by cisplatin treatment at 14 different concentrations (2, 4, 6, 8, 10, 12, 14, 16, 18, 20, 22, 24, 26 and 28 µg/mL respectively). IC50 values were compared with that of A2780 cells (n = 6). M Volcano plot showing the transcriptional activation of the mitochondrial unfolded protein response by 15 µM cisplatin in A2780 cell line. N, O GOPEI and GOPEI-mPEG treatment-induced differential in vitro activation of the UPRmt in (N) A2780 and (O) A2780DDP cell lines. P Schematic diagram illustrating UPRmt activation by cisplatin, graphene and cationic polymers leading to mitochondrial dysfunction and subsequent mito-nuclear signalling.
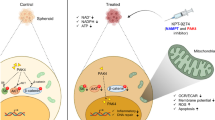
Next, we aimed to further understand the potential mechanism governing cisplatin and GOPEI elicited SOD1 mRNA upregulation and hypothesized the involvement of oxidative stress induction and mitochondrial damage. Cisplatin, GO, and PEI have been reported as causative agents of mitochondrial dysfunction [37]. In the mitochondria, SOD1 is located only in the intermembrane space. In healthy non-pathologic cells, it neutralizes superoxide anions generated by complex III of the electron transport chain, and it is also part of the executive branch of the UPRmt mediating mitoprotective functions primarily upon mitochondrial oxidative stress [38]. Thus, to explore the relationship between cisplatin or nanoparticle-mediated toxicity and the proposed activation of the UPRmt, using a set of UPRmt-specific primers, we monitored the transcriptional activation of the respective signalling pathways and found that cisplatin-induced mitochondrial damage can activate the UPRmt (Fig. 4M) in A2780 ovarian cancer cells on the mRNA and protein levels (unpublished observation). Further, we concluded that both GOPEI and GOPEI-mPEG had similar effect in the same cell line (Fig. 4N), while PEGylation did not entirely prevent the ROS-mediated activation of the UPRmt signalling. In addition, GOPEI and GOPEI-mPEG treatment of A2780DDP cells showed partial UPRmt activation (Fig. 4O), however, at a significantly lower magnitude. Further evaluation of additional mechanistic links between cisplatin or graphene nanoparticle-mediated UPRmt activation is beyond the scope of this study. Although, evidently different compounds can elicit varying degrees of UPRmt activation (Fig. 4P), implying specificity in the underlying mechanism and potential correlation with cisplatin-sensitivity, that merits further investigation in the future. Overall, based on the in vitro study results, we concluded that the prepared GOPEI-mPEG nanoparticle has acceptable biosafety properties for subsequent in vivo studies.
SOD1 knockdown reverses cisplatin resistance in vitro
The in vitro chemo-sensitizing effect of GOPEI-mPEG-siSOD1 was first evaluated by co-treating A2780DDP cells with siSOD1 (65 and 130 nM, respectively) (Fig. 5A). We observed that GOPEI-mPEG-siSOD1 treatment lowered cisplatin IC50 values (p < 0.0001) from a baseline 18.31 ± 0.53 µg/mL to 7.42 ± 0.79 upon treatment with 130 nM siSOD1 complexed GOPEI-mPEG relative to the GOPEI-mPEG-siSCR control group (Fig. 5B). Further, we observed that SOD1 knockdown caused considerable cell growth inhibition, evident from the drop in population doubling times in both examined cell lines (Fig. 5C, D). In addition, the in vitro therapeutic efficiency of SOD1 knockdown was measured using ultra-low attachment plates, clonogenic assay and MTT assay using a fixed concentration of 15 µM cisplatin. We concluded that under all of these conditions, SOD1 knockdown sensitized the A2780DDP cells to cisplatin (Fig. 5E–G). Finally, we measured the total ROS, superoxide, and mitochondrial membrane potential using DCHF-DA, MitoSox-Red, and JC-1 fluorescent dyes (Fig. 5H). We found that SOD1 knockdown alone and in combination with cisplatin treatment resulted in elevated total ROS (p < 0.0001) and further caused mitochondrial membrane depolarization (p < 0.05). We concluded that SOD1 knockdown is associated with significant cellular oxidative stress and mitochondrial toxicity, which may contribute to the sensitization of cisplatin in the A2780DDP cisplatin-resistant cells.
A MTT assays showing reversal of cisplatin resistance by GOPEI-siSOD1. Relative cell viability of A2780DDP cells treated with 65 and 130 nM of GOPEI-siSOD1 at a w/w ratio of 0.2:1 and subsequently with cisplatin (2–28 µg/mL). The graph also shows the relative cell viability of A2780 and A2780DDP cells relative to cisplatin treatment only as a reference (n = 18). B Change of cisplatin IC50 values normalized to that of A2780DDP following treatment. C Population doubling curves of the A2780 cell line (0–96 h) under various treatment conditions. D Population doubling curves of the A2780DDP cell line (0–96 h) under various treatment conditions. E Therapeutic efficacy and antiproliferation effect of GOPEI-mPEG-siSOD1 in A2780 and A2780DDP cells evaluated in ultra-low attachment plates, clonogenic and MTT assays. All values have been normalized to their respective untreated cell line control, and presented as mean ± SD (n = 3). F, G Representative images of ultra-low attachment and clonogenic assay cell culture plates. H Evaluation of total ROS, superoxide and mitochondrial membrane potential with plate reader under siSCR control and siSOD1 knockdown conditions confirmed with Western blot.
Evaluation of in vivo biodistribution, biosafety, and therapeutic efficacy of graphene-based nanoparticles
The in vivo pharmacokinetic and pharmacodynamic properties of GOPEI-mPEG were evaluated using whole-body fluorescent imaging (Fig. 6A). Using normal saline and naked siSCRCy3 as controls, mice bearing A2780DDP xenografts were administered GO, GOPEI, and GOPEI-mPEG complexed with siSCRCy3 via the tail vein and the nanoparticle distribution was monitored at different times points (0, 1, 4, and 8 h). The fluorescent intensity of naked siSCRCy3, GOPEI, and GOPEI-mPEG groups gradually increased during the study. However, no significant change in fluorescence was observed with non-functionalized GO, consistent with previous reports [35, 39]. The recorded fluorescent intensity was comparable between GOPEI- siSCRCy3 and GOPEI-mPEG-siSCRCy3 groups compared to their respective siSCR control groups. We concluded that GOPEI and GOPEI-mPEG could accumulate in tumour tissues and deliver the siSCRCy3 cargo. In addition, these results suggested that PEGylation of GOPEI did not significantly alter the drug loading and delivery capacity of the nanoparticle, a common issue encountered following PEGylation of nanoparticles [40].
A In vivo fluorescence imaging and biodistribution siSCR and siSCRCy3 complexed nanoparticle formulations administered via tail vein injection of female BALB/c nude mice bearing A2780DDP subcutaneous xenografts. Whole body fluorescence images were taken at different time points (0, 1, 4, 8 h) following tail vein injection. B–J In vivo blood biochemistry analysis of (B) ALT (C) AST (D) TP (E) ALB (F) ALP (G) BUN (H) CREA (I) UA (J) LDH of mouse samples following systemic administration of nanoparticles. K In vitro hemolysis activity of naked siSCR, GO, GOPEI and GOPEI-mPEG nanoparticles. Hemolysis rate was measured following incubation of nanoparticle polyplexes with purified red blood cells in PBS. L H&E staining of the heart, kidney, liver, spleen and lungs harvested from mice recieved intravenous naked siSCR, and siSCRCy3 complexed GO, GOPEI and GOPEI-mPEG respectively. Images of collected organs from each group was taken at 8 h after tail vein injection. M In vivo tumour therapeutic efficiency of GOPEI-mPEG-siSOD1. During a 14-day long therapeutic window, mice received 6 doses of respective siSCR or siSOD1-based treatments with or without cisplatin treatment. N H&E staining of A2780 and A2780DDP cell-derived tumour cross-sections in their respective treatment groups as in (Fig. 6M) harvested from mice three days after the last injection.
In addition, blood biochemistry parameters (Fig. 6B–J) were obtained to monitor acute systemic toxicity following intravenous nanoparticle administration via the tail vein. We found that GOPEI-mPEG exhibited lower liver function indices, including ALT (p < 0.05) and AST (p < 0.05), than GOPEI. However, both GOPEI and GOPEI-mPEG nanoparticles showed significantly higher levels of AST (p < 0.0001 and p < 0.001, respectively), while only GOPEI treatment showed ALT elevation (p < 0.001) compared to the blank saline control group. Further, only GOPEI-mPEG showed UA level elevation (p < 0.001), while both nanoparticles significantly raised serum LDH levels (p < 0.001 and p < 0.01, respectively). We concluded that both nanoparticles elicited toxicity to a certain degree in vivo. Furthermore, PEGylation of GOPEI did not entirely circumvent but still considerably decreased the toxicity induced presumably by GO or the PEI cationic polymer functional group.
Further, in vitro, hemocompatibility testing using whole mouse blood showed that GOPEI-complexed siSCR induced a significantly higher (p < 0.0001) hemolysis rate relative to naked siSCR and GO controls in a concentration-dependent manner (0–160 nM). This effect was considerably reduced by the PEGylation of GOPEI (p < 0.01) (Fig. 6K). It has been reported previously that cationic gene-drug carriers would cause blood toxicity by hemolysis and organic damage after administration in vivo [41]. As both GO and PEI have been known to induce hemolysis, we concluded that appropriate surface modification using PEG could effectively improve the biocompatibility of the prepared delivery carrier and moderate hemolytic toxicity in our study.
To establish causality between the potential acute toxicity observed with blood biochemistry testing and the associated pathological changes elicited by the nanoparticles in various organ systems, mice were sacrificed 8 h after intravenous administration of the formulations, and the heart, liver, spleen, kidneys, and lungs were subjected to standard hematoxylin and eosin (H&E) staining. As shown in Fig. 6L, no appreciable pathological findings were observed in the respective control saline/siSCRCy3 and GO/siSCRCy3 groups in agreement with our blood biochemistry results. However, non-PEGylated GOPEI nanoparticles elicited evident and visible structural damage to the liver and kidney by disrupting the physiological histology of the organ parenchyma. This effect was partially mitigated by nanoparticle PEGylation in the GOPEI-mPEG group, although a certain degree of organ damage was still visible in the liver. We concluded that nanoparticle PEGylation considerably decreased the off-target toxicity caused by the cationic polymer component of the nanoparticle; however, due to the propensity of the majority of systematically administered nanoparticles to accumulate in the liver, the IHC results indicated that a certain degree of hepatotoxicity and nephrotoxicity was associated with GOPEI-mPEG administration.
Next, the in vivo anti-tumour and cisplatin chemosensitization effect of GOPEI-mPEG-siSOD1 was evaluated in subcutaneous A2780 and A2780DDP cell-derived xenograft models (Fig. 6M). Relative to the A2780DDP group, identical 0.5 mg/kg intratumoral cisplatin dose induced a significantly higher tumour growth inhibition rate (276.3 ± 72.39 mm3, p = 0.0170) in the sensitive A2780 tumours, confirming that the cisplatin-resistant phenotype of A2780DDP was maintained in vivo as well. Further, we concluded that combined naked siSOD1 and cisplatin treatment had a negligible overall effect on tumour growth compared to cisplatin treatment only (770.81 ± 135.33 mm3, p = 0.9976). In addition, we found that intratumoral combined GOPEI-mPEG-siSCR and cisplatin treatment elicited significantly higher tumour growth inhibition relative to the 0.5 mg/kg naked siSOD1 plus cisplatin group (460.61 ± 59.32 mm3, p = 0.0046). This increased growth inhibition effect was attributed to the previously identified toxicity of the graphene nanoparticle on cancer cells observed with IHC and blood biochemistry testing, which despite the PEGylation, was not entirely circumvented. Finally, we found that relative to GOPEI-mPEG-siSCR group, 0.5 mg/kg siSOD1 complexed with GOPEI-mPEG at a 3:1 w/w ratio co-administered with cisplatin resulted in significant tumour growth inhibition (237.5799 ± 85.18 mm3, p = 0.0265) and structural tumour tissue damage (Fig. 6N) during the 14-day long therapeutic window. These results indicated that relative to the control siSCR group, GOPEI-mPEG-siSOD1 elicited partial re-sensitization of A2780DDP xenografts to cisplatin, which was attributed to the SOD1 knockdown via RNAi.
Discussion
Post-treatment disease recurrence and cisplatin-resistance development are major therapeutic obstacles for ovarian cancer patients [2]. Hence, in this study we aimed at addressing this clinical phenomenon and prepared a GOPEI-mPEG siRNA nanocarrier to selectively target SOD1 in vivo via RNAi, which we have previously identified as a potential target to reverse cisplatin-resistance [22]. GOPEI-mPEG exhibited favourable physicochemical characteristics for translational drug delivery applications regarding hydrodynamic diameter, ζ-potential, and PDI [24].
Following PEGylation, the cellular uptake studies confirmed the presence of adequate PEI surface charge to mediate endosomal escape via the intracellular proton sponge effect [42]. This mechanism was confirmed with fluorescent live cellular uptake studies, as an abrupt increase in fluorescence of siSCRFAM was detected after 5 h indicating the detachment and regained fluorescence of siSCRFAM following the endosomal escape. GOPEI-mPEG showed appropriate biocompatibility and formed homogenous solutions in various media modelling in vitro and in vivo conditions. At the same time, the gel- and fluorescence-based kinetic studies implied the complexed siSCR cargo was protected from RNase degradation. Further, all the prepared PEI-functionalized nanoparticles fully complexed siSCR at a 2:1 w/w ratio.
The cytotoxicity of the PEI cationic polymer component was significantly reduced with PEGylation; still, IHC imaging revealed that GOPEI-mPEG accumulation in the liver elicited structural damage and hepatotoxicity. We attributed the cytotoxicity produced by the PEI functional group to oxidative stress induction, which led to evident SOD1 mRNA overexpression by GOPEI in a concentration-dependent manner [43]. It can be concluded that non-PEGylated GOPEI can counteract the knockdown effect mediated by siSOD1, which, theoretically based on our original hypothesis, may increase cisplatin resistance in surviving cell populations and subsequent passages via clonal selection.
The nanocarrier’s size, shape, surface charge, and surface functionalization determine the net toxicity of a graphene nanoparticle [44]. The increased ROS generation by concentration-dependent graphene treatment can cause oxidative stress-mediated mitochondrial damage and stress-reactive transcriptional factor activation, altering the gene expression profile of the affected cell [45]. The resultant oxidative stress, thus, may upregulate vital antioxidant enzymes, including but not limited to CAT, PRX, SOD2, and SOD1, as a defense mechanism [46]. For instance, Mendonca et al. showed that graphene oxide caused significant SOD1 overexpression from 1 h up to 7 days following transfection [46]. A similar cellular effect was published by Jarosz et al., who found that both SOD1 and SOD2 were overexpressed upon treating HepG2 cells with GO in a concentration-dependent manner [47].
Further, we concluded that, like cisplatin, both GOPEI and GOPEI-mPEG nanoparticles elicited mitochondrial dysfunction, evident from the specific activation of various arms of the UPRmt pathways [48]. GO-based nanoparticles and PEI have been shown to impair tumour progression and suppress metastatic potential by inhibiting and disrupting mitochondrial respiration [49]. In addition, GO-induced ROS generation can stimulate mitochondrial dysfunction and subsequent apoptotic signalling pathways, which may explain the decrease in cisplatin IC50 values of A2780DDP cells following GOPEI treatment [47]. Through a series of immunohistochemistry analyses among various treatment groups to monitor the change in protein abundance of 7 UPRmt signature genes including FOXO3, HSP60, LC3B, NRF1, SIRT3, SOD1, and SOD2 according to previous literature [48]. We have concluded that both GOPEI-mPEG mediated in vivo SOD1 knockdown evoke UPRmt activation (detailed data not shown). Further, we observed the baseline upregulation of UPRmt markers upon cisplatin treatment. To the best of our knowledge, this is the first report of cisplatin-induced activation of all three arms of the UPRmt in ovarian cancer [49]. While further exploration of these pathways is beyond the scope of this study, based on the striking differences in the activation of UPRmt pathways between baseline and upon cisplatin treated cells, it would be intriguing to explore further the potential involvement of these pathways in intrinsic or acquired cisplatin resistance in other cell lines as well. Furthermore, as graphene-based nanomaterials tend to accumulate and intercalate in the mitochondrial outer membrane causing mitotoxicity, thus it would be reasonable to examine whether the activation of the UPRmt could be used as a reporter in graphene biosafety and biocompatibility assays [50].
Conclusion
Despite the observed toxicities mediated by the respective nanoparticles, the in vivo results showed that SOD1 knockdown has a partial chemo-sensitising effect in cisplatin resistance relative to the siSCR control group. This observed effect is per our previous in vitro results. We concluded that while these promising results exhibited the potential role of SOD1 as a therapeutic target for the reversal of cisplatin resistance, yet the full therapeutic potential of SOD1 knockdown has not been fully demonstrated in this study due to the SOD1 mRNA induction properties of the nanoparticles, which counteracted the effect of the siSOD1 dosing primarily arising from the toxicity mediated presumably by the cationic polymer component. Overall, the therapeutic applicability of SOD1 knockdown in reversing cisplatin resistance will be further investigated using other drug delivery platforms such as lipid nanoparticles (LNPs) and in other cell- and patient-derived xenograft models.
Data availability
Materials described in this manuscript including all relevant raw data will be freely available from the corresponding author for non-commercial use upon request.
References
Webb PM, Jordan SJ. Epidemiology of epithelial ovarian cancer. Best Pr Res Clin Obstet Gynaecol. 2017;41:3–14.
Jayson GC, Kohn EC, Kitchener HC, Ledermann JA. Ovarian cancer. Lancet. 2014;384:1376–88.
Prestayko AW, D’Aoust JC, Issell BF, Crooke ST. Cisplatin (cis-diamminedichloroplatinum II). Cancer Treat Rev. 1979;6:17–39.
Lebwohl D, Canetta R. Clinical development of platinum complexes in cancer therapy: an historical perspective and an update. Eur J Cancer. 1998;34:1522–34.
Dasari S, Tchounwou PB. Cisplatin in cancer therapy: molecular mechanisms of action. Eur J Pharmacol. 2014;740:364–78.
Galanski M. Recent developments in the field of anticancer platinum complexes. Recent Pat Anticancer Drug Discov. 2006;1:285–95.
Kelland L. The resurgence of platinum-based cancer chemotherapy. Nat Rev Cancer. 2007;7:573–84.
Kartalou M, Essigmann JM. Mechanisms of resistance to cisplatin. Mutat Res. 2001;478:23–43.
Pokhriyal R, Hariprasad R, Kumar L, Hariprasad G. Chemotherapy resistance in advanced ovarian cancer patients. Biomark Cancer. 2019;11:1179299x19860815.
van Zyl B, Tang D, Bowden NA. Biomarkers of platinum resistance in ovarian cancer: what can we use to improve treatment. Endocr Relat Cancer. 2018;25:R303–r18.
Galluzzi L, Vitale I, Michels J, Brenner C, Szabadkai G, Harel-Bellan A, et al. Systems biology of cisplatin resistance: past, present and future. Cell Death Dis. 2014;5:e1257.
Eckstein N. Platinum resistance in breast and ovarian cancer cell lines. J Exp Clin Cancer Res. 2011;30:91.
Davis A, Tinker AV, Friedlander M. “Platinum resistant” ovarian cancer: what is it, who to treat and how to measure benefit? Gynecol Oncol. 2014;133:624–31.
Januchowski R, Sterzyńska K, Zaorska K, Sosińska P, Klejewski A, Brązert M, et al. Analysis of MDR genes expression and cross-resistance in eight drug resistant ovarian cancer cell lines. J Ovarian Res. 2016;9:65.
Battistella C, Klok HA. Reversion of P-gp-mediated drug resistance in ovarian carcinoma cells with PHPMA-zosuquidar conjugates. Biomacromolecules. 2017;18:1855–65.
Fitzpatrick DPG, You J-S, Bemis KG, Wery J-P, Ludwig JR, Wang M. Searching for potential biomarkers of cisplatin resistance in human ovarian cancer using a label-free LC/MS-based protein quantification method. PROTEOMICS – Clin Appl. 2007;1:246–63.
Eleutherio ECA, Silva Magalhães RS, de Araújo Brasil A, Monteiro Neto JR, de Holanda Paranhos L. SOD1, more than just an antioxidant. Arch Biochem Biophys. 2021;697:108701.
Phillips JP, Campbell SD, Michaud D, Charbonneau M, Hilliker AJ. Null mutation of copper/zinc superoxide dismutase in Drosophila confers hypersensitivity to paraquat and reduced longevity. Proc Natl Acad Sci USA. 1989;86:2761–5.
Blander G, de Oliveira RM, Conboy CM, Haigis M, Guarente L. Superoxide dismutase 1 knock-down induces senescence in human fibroblasts. J Biol Chem. 2003;278:38966–9.
Noman ASM, Parag RR, Rashid MI, Islam S, Rahman MZ, Chowdhury AA, et al. Chemotherapeutic resistance of head and neck squamous cell carcinoma is mediated by EpCAM induction driven by IL-6/p62 associated Nrf2-antioxidant pathway activation. Cell Death Dis. 2020;11:663.
Brown DP, Chin-Sinex H, Nie B, Mendonca MS, Wang M. Targeting superoxide dismutase 1 to overcome cisplatin resistance in human ovarian cancer. Cancer Chemother Pharmacol. 2009;63:723–30.
Kim JW, Sahm H, You J, Wang M. Knock-down of superoxide dismutase 1 sensitizes cisplatin-resistant human ovarian cancer cells. Anticancer Res. 2010;30:2577–81.
Kim HJ, Kim A, Miyata K, Kataoka K. Recent progress in development of siRNA delivery vehicles for cancer therapy. Adv Drug Deliv Rev. 2016;104:61–77.
Tatiparti K, Sau S, Kashaw SK, Iyer AK. siRNA delivery strategies: a comprehensive review of recent developments. Nanomaterials. 2017;7:77.
Li J, Liang H, Liu J, Wang Z. Poly (amidoamine) (PAMAM) dendrimer mediated delivery of drug and pDNA/siRNA for cancer therapy. Int J Pharm. 2018;546:215–25.
Geim AK, Novoselov KS. The rise of graphene. Nat Mater. 2007;6:183–91.
Daniyal M, Liu B, Wang W. Comprehensive review on graphene oxide for use in drug delivery system. Curr Med Chem. 2020;27:3665–85.
Singh DP, Herrera CE, Singh B, Singh S, Singh RK, Kumar R. Graphene oxide: an efficient material and recent approach for biotechnological and biomedical applications. Mater Sci Eng C Mater Biol Appl. 2018;86:173–97.
Zhang L, Lu Z, Zhao Q, Huang J, Shen H, Zhang Z. Enhanced chemotherapy efficacy by sequential delivery of siRNA and anticancer drugs using PEI-grafted graphene oxide. Small. 2011;7:460–4.
Ulasov AV, Khramtsov YV, Trusov GA, Rosenkranz AA, Sverdlov ED, Sobolev AS. Properties of PEI-based polyplex nanoparticles that correlate with their transfection efficacy. Mol Ther. 2011;19:103–12.
Hall A, Lächelt U, Bartek J, Wagner E, Moghimi SM. Polyplex evolution: understanding biology, optimizing performance. Mol Ther. 2017;25:1476–90.
Guo M, Huang J, Deng Y, Shen H, Ma Y, Zhang M, et al. pH-responsive cyanine-grafted graphene oxide for fluorescence resonance energy transfer-enhanced photothermal therapy. Adv Funct Mater. 2015;25:59–67.
Rao X, Huang X, Zhou Z, Lin X. An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat Bioinforma Biomath. 2013;3:71–85.
Chen B, Liu M, Zhang L, Huang J, Yao J, Zhang Z. Polyethylenimine-functionalized graphene oxide as an efficient gene delivery vector. J Mater Chem. 2011;21:7736–41.
de Lázaro I, Vranic S, Marson D, Rodrigues AF, Buggio M, Esteban-Arranz A, et al. Graphene oxide as a 2D platform for complexation and intracellular delivery of siRNA. Nanoscale. 2019;11:13863–77.
Boussif O, Lezoualc'h F, Zanta MA, Mergny MD, Scherman D, Demeneix B, et al. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: polyethylenimine. Proc Natl Acad Sci USA. 1995;92:7297–301.
Zhu L, Yuan Y, Yuan L, Li L, Liu F, Liu J, et al. Activation of TFEB-mediated autophagy by trehalose attenuates mitochondrial dysfunction in cisplatin-induced acute kidney injury. Theranostics. 2020;10:5829–44.
Kenny TC, Craig AJ, Villanueva A, Germain D. Mitohormesis primes tumor invasion and metastasis. Cell Rep. 2019;27:2292–303.e6.
Reina G, Chau NDQ, Nishina Y, Bianco A. Graphene oxide size and oxidation degree govern its supramolecular interactions with siRNA. Nanoscale. 2018;10:5965–74.
Hattori Y, Tamaki K, Sakasai S, Ozaki KI, Onishi H. Effects of PEG anchors in PEGylated siRNA lipoplexes on in vitro gene‑silencing effects and siRNA biodistribution in mice. Mol Med Rep. 2020;22:4183–96.
Lv Y, Chen B, Chen J, Dong Y, Liu JH, Xu L. Effect of graphene oxide on the function of erythrocytes. Sheng Wu Gong Cheng Xue Bao. 2021;37:4047–55.
Smith SA, Selby LI, Johnston APR, Such GK. The endosomal escape of nanoparticles: toward more efficient cellular delivery. Bioconjug Chem. 2019;30:263–72.
Calarco A, Bosetti M, Margarucci S, Fusaro L, Nicolì E, Petillo O, et al. The genotoxicity of PEI-based nanoparticles is reduced by acetylation of polyethylenimine amines in human primary cells. Toxicol Lett. 2013;218:10–7.
Tabish TA, Zhang S, Winyard PG. Developing the next generation of graphene-based platforms for cancer therapeutics: The potential role of reactive oxygen species. Redox Biol. 2018;15:34–40.
Gurunathan S, Arsalan Iqbal M, Qasim M, Park CH, Yoo H, Hwang JH, et al. Evaluation of graphene oxide induced cellular toxicity and transcriptome analysis in human embryonic kidney cells. Nanomaterials. 2019;9:969.
Mendonça MC, Soares ES, de Jesus MB, Ceragioli HJ, Irazusta SP, Batista ÂG, et al. Reduced graphene oxide: nanotoxicological profile in rats. J Nanobiotechnology. 2016;14:53.
Jarosz A, Skoda M, Dudek I, Szukiewicz D. Oxidative stress and mitochondrial activation as the main mechanisms underlying graphene toxicity against human cancer cells. Oxid Med Cell Longev. 2016;2016:5851035.
Kenny TC, Gomez ML, Germain D. Mitohormesis, UPR(mt), and the complexity of mitochondrial DNA landscapes in cancer. Cancer Res. 2019;79:6057–66.
Chatterjee N, Eom HJ, Choi J. A systems toxicology approach to the surface functionality control of graphene-cell interactions. Biomaterials. 2014;35:1109–27.
Ou L, Song B, Liang H, Liu J, Feng X, Deng B, et al. Toxicity of graphene-family nanoparticles: a general review of the origins and mechanisms. Part Fibre Toxicol. 2016;13:57.
Acknowledgements
We express our gratitude to the XJTLU research facilities to allow us to use all the required instruments for nanoparticle characterization.
Funding
This work was supported by XJTLU Key Programme Special Fund – KSF Exploratory Research KSF-E-14.
Author information
Authors and Affiliations
Contributions
AS and ES contributed to the writing of the manuscript. AS and MW contributed to the conception and experimental design of the study; AS, ES, JH, and ZZ contributed to the nanoparticle preparation and characterization; AS, HD, and PZ contributed significantly to the in vivo study. MW and SR helped with manuscript revising. All authors approved the final version and agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Animal Study Ethics approval of this research was granted by the Ethics Committee of Xi’an Jiaotong-Liverpool University. The animal study was carried out in Suzhou GenePharma with a certificate of government approved Animal Study Facility.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Szénási, A., Sivasudhan, E., Du, H. et al. Targeting SOD1 via RNAi with PEGylated graphene oxide nanoparticles in platinum-resistant ovarian cancer. Cancer Gene Ther 30, 1554–1568 (2023). https://doi.org/10.1038/s41417-023-00659-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-023-00659-2