Abstract
YAP is a transcriptional co-activator with critical roles in tumorigenesis. However, its upstream regulatory mechanism, especially how its mRNA stability is regulated, remains to be further studied. Here, we validated that YAP expression was higher in lung adenocarcinoma (LUAD) tissues compared to adjacent normal tissues, and found that YAP m5C modification occurred in its 328–331 3′ UTR region under the promotion NSUN2 and ALYREF, and increased the stability of YAP mRNA. This m5C modification also inhibited miR-582-3p binding and m6A modification in the nearby region. In addition, YAP m5C modification enhanced the exosome secretion effect, which was caused by two YAP-dependent transcription factors, Mycn and SOX10, and then stimulating the transcription of seven downstream exosome-promoting genes. Furthermore, we found that YAP m5C modification and its exosome-secretion-promoting function contributed to the malignant phenotype and AZD9291 (a third-generation EGFR-TKI) resistance of LUAD cells. Collectively, YAP is promoted by its m5C modification, and blocking YAP m5C modification will be helpful for future LUAD treatment.
Similar content being viewed by others
Introduction
YAP is a cancer-promoting transcription co-factor downstream of Hippo pathway [1]. Transcription factors related to YAP include TEAD family, SMAD family, RUNX family, etc [2, 3]. Our Lab found that transcription factor TFCP2 regulated by YAP to promote the transcription of ferritin [4, 5]. Additionally, TFCP2 also recruits transcription factors for YAP, including FOXC1, FOXA1, etc., to promote cancer [6]. We also found that transcription-promoting activity of YAP is controlled by post-translational modifications such as O-GlcNAcylation, which can inhibit YAP phosphorylation [7]. In lung adenocarcinoma (LUAD), YAP can promote tumor cell proliferation and stemness [8]. High expression of YAP predicts poor prognosis in EGFR-mutant patients [9]. YAP 331 arginine to tryptophan (R331W) mutation is an allele predisposed for LUAD with high familial penetrance [10]. The above findings suggest that YAP is a promoting factor for LUAD. However, the mechanism by which YAP activates and promotes LUAD is still not fully understood.
The main function of YAP is to control organ size by regulating cell proliferation and contact inhibition [11]. In tumors, high expression of YAP causes cells to acquire stemness, and echoes the effect of epithelial-to-mesenchymal transition (EMT) [12]. Therefore, tumors with high expression of YAP have a strong ability of invasion and metastasis [13]. Under hypoxia, YAP can bind and maintain the stability of HIF1α, and further promote the transcription of PKM2 gene to stimulate glycolysis [14]. Our Lab elucidated that YAP promotes NUDT9 and SLC5A3 transcription and activates the hexosamine biosynthesis pathway (HBP) [7]. YAP also promotes tumor progression by regulating tumor microenvironment, for example by promoting transcription of CCL2, a macrophage-specific chemokine, to recruit tumor-associated macrophages [15]. In tumor microenvironment, exosome is a type of extracellular vesicle that originated from endocytosis and contains a broad repertoire of cargoes, including nucleic acids (i.e., DNA, mRNA, microRNA, long and short noncoding RNA) and proteins (i.e., cytoskeletal proteins, transmembrane proteins, heat shock proteins and ATPase) [16, 17]. As a master regulator of cellular signaling, exosome orchestrates various autocrine and paracrine functions to alter tumor microenvironment, growth and progression [17]. Tumor cells can transfer malignant phenotype to normal cells via exosome, and establish a fertile local and distant microenvironment to help cancer cell growth [18]. However, studies on the role of YAP in promoting exosomes are limited. YAP is also an important factor promoting drug resistance in cancer [19]. Targeted therapy for LUAD has achieved promising effects in the treatment of some driver gene mutations such as EGFR mutation [20]. Currently, third-generation EGFR-tyrosine kinase inhibitor (TKI) has been used in first-line treatment for LUAD, which can target T790M mutation resistance caused by first generation EGFR-TKI, and other EGFR oncogenic mutations such as exon 19 deletion [21, 22]. However, third-generation EGFR-TKI further generates new drug resistance such as C797S mutation [23]. Therefore, whether other proto-factors such as YAP can lead to third-generation EGFR-TKI resistance remains to be studied.
YAP can be phosphorylated by the core kinase in Hippo pathway, LATS1/2, to induce cytoplasmic localization, and subsequent ubiquitin mediated degradation driven by the E3-ligase βTrCP [24]. However, some regulation patterns to YAP are independent of Hippo pathway. For example, our Lab also revealed that YAP can be O-GlcNAcylated at Thr241 and this O-GlcNAcylation event antagonizes Hippo pathway-mediated phosphorylation of YAP [7]. 3′ untranslated region (3′UTR) is an essential region in mRNA and have important functions in regulating mRNA localization, mRNA stability, and translation [25]. Most functions of 3′ UTRs are mediated by RNA-binding proteins [25]. For example, YAP 3′UTR can be alternative spliced in a splicing factor hhnRNP F-dependent manner, and further lead to entire mRNA degradation [26]. However, microRNA binding and RNA modification (such as N6-methyladenosine (m6A) modification, the most abundant internal modification of RNA in eukaryotic cells which occurs at the consensus motif RRACH (R = A or G, H = A, C or U) [27]) also play independent regulatory roles in 3′UTR [28]. For instance, miR-195-5p and miR-381 are two microRNAs that are identified to directly bind to YAP 3′UTR to induce YAP mRNA degradation [29, 30]. Our Lab also found that both miR-7 and miR-582-3p binds to YAP 3′ UTR to result in degradation of YAP mRNA, while m6A modification near the binding site of miR-582-3p on YAP 3′ UTR can inhibit the miR-582-3p binding [31, 32]. 5-methylcytosine (m5C) modification, another type of abundant RNA methylation, is catalyzed by 5-methylcytosine RNA methyltransferase to transfer a methyl group to the fifth carbon of a cytosine base in RNA sequences [33]. m5C has been frequently associated with tRNA, but also found in mRNAs most often in their 5′ and 3′ UTR regions [34], and has emerged as an important regulator of many aspects of gene expression, including RNA export, ribosome assembly, translation, and RNA stability [35]. NSUN2, a major m5C mRNA methyltransferase, has been found overexpressed and activated in many types of cancer including LUAD [36]. However, how m5C modification and NSUN2 promote cancer, and how they function through YAP still remains largely unknown.
In this study, we elucidated YAP mRNA stability was increased by its m5C modification, which was NSUN2- and ALYREF-dependent. Moreover, m5C modification of YAP enhanced its promoting effect of exosome secretion. YAP m5C modification and its function to promote exosome secretion contributed to the transformation phenotype and AZD9291 resistance of LUAD cells. Targeting YAP m5C might be helpful for treating LUAD in the future.
Materials and methods
Tissue samples
All the tissue and plasma specimens (Shanghai, China, mean age ± SD, 60.05 ± 8.37 years; male: female ratio, 1.04:1) were recruited in Shanghai Chest Hospital (Shanghai, China) from March 2015 to December 2021. The diagnostic power was analyzed by receiver operating characteristic (ROC) curve analysis.
Immunoblotting (IB) and enzyme linked immunosorbent assay (ELISA)
Protein levels were analyzed using IB and ELISA following conventional protocols. For IB, the primary antibodies used were anti-YAP (Abcam, Cambridge, MA, USA, #ab52771), anti-GAPDH (CST, Boston, MA, USA, #5174 and #51332), anti-p-127-YAP (Abcam, #ab76252), anti-p-381-YAP (CST, #13619), anti-O-Glc-YAP (developed by Biolynx, Hangzhou, China), anti-HRS (Abcam, #ab72053), anti-PLD2 (CST, #13904), anti-RAB2B (Novus, Littleton, CO, USA, #NBP1-31631), anti-RAB27A (Abcam, #ab55667), anti-RAB27B (Abcam, #ab103418), anti-VAMP (Abcam, #ab36195), anti-ATG7 (Abcam, #ab52472), anti-SOX10 (Abcam, #ab227680), anti-Mycn (Abcam, #ab227822), anti-ALYREF (Abcam, #ab202894), anti-NSUN2 (Abcam, #ab259941), anti-DNMT2 (Abcam, #ab220175), anti-NSUN1 (Abcam, #ab270175), anti-NSUN3 (Abcam, #ab272616), anti-NSUN4 (Abcam, #ab235430), anti-NSUN5 (Abcam, #ab121633), anti-NSUN6 (Novus, #NBP1-92202), anti-NSUN7 (Biorbyt, Cambridge, UK, #orb258175), anti-YBX1 (Abcam, #ab255606), anti-EXOSC10 (Abcam, #ab94981), anti-SPI1 (Abcam, #ab227835), anti-CD63 (Abcam, #ab271286), anti-TSG101 (Abcam, #12501), anti-ALIX (Abcam, #ab275377), anti-CD9 (Abcam, #ab92726), and anti-calnexin (Abcam, #ab133615). For ELISA, YAP and NSUN2 levels were measured using kits from Yingxin Biotech Ltd. (Shanghai, China) as per manufacturer’s instructions.
Cell culture
Established A549, H1299, H1975, Wi-38, MRC-5 and BEAS-2B cell lines were purchased from Fuheng Biotechnology (Shanghai, China). H1975- and HCC827-based AZD9291 resistant cell lines were acquired from previous studies [5]. For 3D spheroid culture, basement membrane extract (BME) (Trevigen, Gaithersburg, MD, USA) was seeded in a 96-well plate at 50 μl/well and plated at 37 °C for 30 min. Then, cells with indicated genes overexpressed or knocked out were seeded on top of BME at a density of 1 × 105 cells per well. After formation of spheroids, they were treated with AZD9291 (MedChemExpress, Monmouth, NJ, USA) with or without GW4869 (Sigma, St Louis, MO, USA) for 24 h. Images were captured after staining with SYTOX green (Invitrogen, Carbsland, CA, USA) [37].
RNA-immunoprecipitation (RNA-IP)
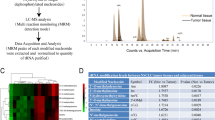
For RIP assay, a Magna RIP Kit (Merck Millipore, Billerica, MA, USA) was used. Briefly, a total of 1 × 107 cells were lysed in complete RNA lysis buffer, then cell lysates were incubated with RIP immunoprecipitation buffer containing magnetic beads conjugated with 5 μg anti-m5C (Abcam, #ab10805), anti-DNMT2 (Abcam, #ab220175), anti-NSUN1 (Abcam, #ab270175), anti-ALYREF (Abcam, #ab202894), anti-NSUN2 (Abcam, #ab259941), anti-DNMT2 (Abcam, #ab220175), anti-NSUN3 (Abcam, #ab272616), anti-NSUN4 (Abcam, #ab235430), anti-NSUN5 (Abcam, #ab121633), anti-NSUN6 (Novus, #NBP1-92202), anti-NSUN7 (Biorbyt, #orb258175), anti-YBX1 (Abcam, #ab255606) antibodies or control IgG (CST #3900) overnight at 4 °C. Next, the beads were washed with washing buffer and incubated with proteinase K at 55 °C for 30 min to remove the proteins. The remaining RNA was extracted using Trizol (Ambion) and reverse-transcribed into complementary DNA using the PrimeScriptTM RT reagent kit (Takara). The SYBR premix Ex Taq (Takara) kit was used for real-time qPCR [31].
Regents and plasmids
For regents, ActD (MedChemExpress), CHX (Sigma), GW4869 (Sigma), AZD9291 (MedChemExpress). Plasmids DNMT2, NSUN1-7, ALYREF, YBX1, EXOSC10, Mycn, SOX10, SPI1, YAP-(-1k)-promoter, YAP-(-2k)-promoter, Mut4-, Mut5-, Mut6-YAP-3’ UTR overexpression plasmids were purchased from Biolink (Shanghai, China). miR-582-3p mimics, miR-582-3p inhibitors, WT-, Mut1-, Mut2- and Mut3-YAP-3′ UTR luciferase reporters, YAP knockout and YAP overexpression plasmids were acquired from previous studies [5, 7, 31]. Short hairpin RNAs (shRNAs) and lentiCRISPR v2-based constructs were used for knocking down and knocking out Mycn, SOX10, NSUN2 and ALYREF, respectively. The sequences for gRNAs and shRNAs were summarized in Table S1.
Quantitative RT-PCR (qPCR)
Total RNA was extracted using Trizol (Ambion, Carlsbad, CA, USA) and reverse-transcribed into complementary DNA using the PrimeScriptTM RT reagent kit (Takara, Dalian, China). The SYBR premix Ex Taq (Takara) kit was used for real-time qPCR. The primers are listed in Table S1.
Luciferase activity measurement
Luciferase activities were measured using a dual-luciferase kit (Promega, Madison, WI, USA) according to the manufacturer’s instructions. Firefly reporters (including WT-, Mut1- Mut2-, Mut3-, Mut4- Mut5- and Mut6-YAP-3′ UTR, as well as YAP-(-1k)-promoter, YAP-(-2k)-promoter luciferase reporters) were co-transfected into lung cancer cells with Renilla plasmids using Lipofectamine™ 2000 transfection reagent (Invitrogen). After incubation for 48 h, the cells were harvested and then lysed in the passive lysis buffer from the kit. The fluorescence intensity of luciferase reporters was then examined and normalized to the Renilla luciferase activity.
Co-immunoprecipitation (co-IP)
co-IP was performed as described previously. Cell lysates were mixed with 50 ul protein A/G-magnetic beads (Novex, Oslo, Norway) and incubated at 4 °C overnight with the indicated antibodies. The beads were washed with Western/IP lysis buffer (Beyotime, Haimen, China), suspended in SDS-PAGE loading buffer and then detected by IB. The antibodies used for co-IP were: anti-YAP (Santa Cruz Biotechnology, Santa Cruz, CA, USA, #sc-101199).
RNA probe precipitation
Biotin-labeled probe was synthesized by Sangon Biotech (Shanghai, China). Cells were fixed using 4% formaldehyde, then lysed, sonicated, and centrifuged. Next, the supernatant was extracted as input, and the remaining amount was incubated with indicated probe and Dynabeads M-280 Streptavidin (Thermo Scientific, Waltham, MA, USA) overnight at room temperature. The next day, the mixture was washed and incubated with lysis buffer and proteinase K to reverse the cross-linking. Finally, the RNA mixture was extracted using Trizol and was detected via qPCR [5,6,7, 31]. Probe sequence was listed in Table S1.
Isolation and measurement of exosome
Exosome was isolated from cultured media of cells or plasma of human via three sequential centrifugation steps at 4 °C, (1) 15 min at 500 g to remove cells; (2) 30 min at 10,000 g to remove cell debris; and (3) ultracentrifugation at 110,000 g for 70 min to pellet exosome. The pellet was finally resuspended in PBS and centrifuged at 110,000 g for another 70 min to remove soluble and secreted proteins. The concentration and size of exosome were analyzed using nanoparticle tracking analysis (NTA) by a Nanosight NS 300 system (NanoSight Technology, Malvern, UK) [37].
Immunofluorescence (IF), immunohistochemistry (IHC),
IF and IHC were performed according to the conventional protocols. For IF, PKH67 (Sigma, MKCG5294) was probed for marking exosome packaged in H1299 cells. For IHC, the primary antibodies used in the study were anti-YAP (Abcam, #ab52771).
TEM analysis
Exosome was resuspended in 4% paraformaldehyde, and images were captured using the JEM1230 TEM (JEOL, Tokyo, Japan).
Measurements of cell viability, caspase 3/7 activity and anchorage-independent colony formation
Cell viability was detected using a CCK8 kit (Beyotime). Caspase3/7 activity was measured using Caspase 3/7 Glo luciferase reagent (Promega). As for anchorage-independent colony formation assay, LUAD cells were seeded in a six-well plate containing 0.3% agarose at a density of 6 × 103 cells per well. Two weeks later, the numbers of colonies were calculated under microscope.
Mouse experiments
A549 cells with indicated genes overexpressed or knocked out (initial 5 × 106) were subcutaneously injected into 6-week-old athymic nude mice (Jiesijie, Shanghai, China) with or without GW4869 treatment to acquire cell-derived xenograft mouse models. Fresh tissues in a size of 2 mm3 were subcutaneously implanted into six-week-old athymic nude mice to generate PDX mouse models. After successful passage, the PDX model could be used for further analysis. Tumor volumes were calculated as 0.5 × L × W2 (L indicating length while W indicating width) [5]. Histological sections were prepared using a microtome and subsequently stained with IHC or hematoxylin and eosin.
Statistical analysis
Tests used in this study included student’s t test, one-way, two-way ANOVA, χ2 test, and the Spearman rank-correlation analysis. A p < 0.05 was considered statistically significant.
Results
YAP was highly expressed in LUAD
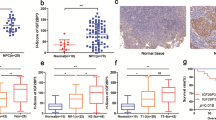
Firstly, we evaluate YAP expression in LUAD. After analyzed paired of LUAD and adjacent normal tissues, we found that YAP protein level was increased in LUAD samples (Fig. 1A). Also, YAP protein level was upregulated in stage III LUAD tissues compared to stage I and II LUAD tissues, as well as in >3 cm diameter LUAD tissues compared to <3 cm LUAD tissues (Fig. 1B, C). The diagnostic power was calculated by ROC curve, and the AUC-ROC of YAP was 0.8331, 95% CI was 0.7765–0.8897, the best sensitivity was 81.73%, and the best specificity was 81.73% (Fig. 1D). We also found YAP expression was not associated with patient age, gender and smoking habit (Fig. S1A). These data suggested YAP was highly expressed in LUAD.
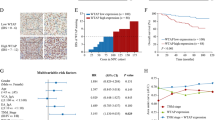
A YAP protein levels were measured by ELISA in LUAD and adjacent normal tissues. B, C YAP protein levels in different stage (B) and tumor diameter (C) LUAD tissues. D ROC curves of YAP for discriminating LUAD from non-tumor tissues. E The enrichment of m5C in A549, H1299, Wi-38, MRC-5 and BEAS-2B cells at indicated regions (P1: 1–700, P2: 701–1400, P3: 1401–2100, P4: 2101–2800 P5: 2801–3493) of YAP 3′ UTR was calculated as the percentage of the input RNA via RNA-IP using the anti-m5C antibodies. F The enrichment of indicated m5C writers in A549 and H1299 cells at P1 region of YAP 3′ UTR was calculated as the percentage of the input RNA via RNA-IP using the indicated antibodies. G YAP mRNA level was measured by qPCR in A549 and H1299 cells with indicated plasmids overexpressed. H YAP protein level was measured by IB in A549 cells with indicated plasmids overexpressed. The level of proteins was normalized to that of GAPDH, and the normalized level of proteins in negative control groups were arbitrarily set to 1. I NSUN2 protein levels were measured in LUAD and adjacent normal tissues. J Correlation between plasma exosome concentration and tissue YAP protein level in LUAD samples. K YAP m5C levels were measured in LUAD and adjacent normal tissues. L, M NSUN2 protein levels in different stage (L) and tumor diameter (M) LUAD tissues. N, O ROC curves of NSUN2 and YAP m5C for discriminating LUAD from non-tumor tissues. The data are shown as the means ± SD from three independent experiments. Images of IB are representative ones of three independent experiments. **p < 0.01 indicate statistical significance. The data from (A, C, I, K, M) were analyzed by a student’s t test. The data from (B, F, G, L) were analyzed by a one-way ANOVA test. The data from (E) were analyzed by a two-way ANOVA test. The data from (J) were analyzed by a Spearman rank-correlation analysis.
m5C modification of YAP depends on NSUN2
In our previous research, we found that RNA m6A modification exists at YAP 3′ UTR region [31]. We further explored whether m5C modification, another common RNA modification, exists at YAP 3′ UTR region. We divided YAP 3′ UTR region into 5 segments, and then found that m5C modification existed in all 5 segments, and the m5C modification abundance of P1 region (1–700) was the highest (Fig. 1E, Fig. S1B) in A549 and H1299 cells. We also tested YAP m5C modification in lung fibroblast cell lines Wi-38 and MRC-5, and bronchial epithelial cell line BEAS-2B, and found that m5C modification levels in Wi-38, MRC-5 and BEAS-2B were significantly lower than those in A549 and H1299 cells (Fig. 1E) Through the screening of m5C methyltransferase (including DNMT2, NSUN1-7 [35]), we found that NSUN2 bound to the P1 region more significantly (Fig. 1F, Fig. S1C). Moreover, NSUN2 increased YAP mRNA and protein expression, whereas other m5C methyltransferases did not have similar capabilities (Fig. 1G, H and Fig. S1D). In addition, NSUN2 protein level was increased in LUAD samples compared to adjacent normal tissues (Fig. 1I) and positively associated with YAP protein level (Fig. 1J). Also, increasing YAP m5C modification were found in LUAD samples (Fig. 1K). Like YAP, NSUN2 protein levels were upregulated in stage III LUAD tissues (Fig. 1L), as well as in >3 cm diameter LUAD tissues (Fig. 1M). The AUC-ROC of NSUN2 was 0.8238, 95% CI was 0.7666–0.8809, the best sensitivity was 80.77%, and the best specificity was 74.04% (Fig. 1N). The AUC-ROC of YAP m5C was 0.8133, 95% CI was 0.6970–0.9296, the best sensitivity was 90.00%, and the best specificity was 76.67% (Fig. 1O). These data implied that YAP m5C depends on NSUN2.
m5C modification of YAP relies on ALYREF and increases YAP mRNA stability
We subsequently investigated the reader that is responsible for YAP m5C. We found ALYREF, but not another reader YBX1, bound to the P1 region (Fig. 2A, Fig. S2A). ALYREF increased YAP mRNA and protein expression, whereas this effect was abolished by NSUN2 knockdown; meanwhile, YBX1 had no effect on up-regulating YAP (Fig. 2B–D, Fig. S2B). NSUN2 and ALYREF overexpression both upregulated, while NSUN2 and ALYREF knockdown down-regulated the stability of YAP mRNA. The effects of NSUN2sh2 and ALYREFsh2 which are designed for the 3′ UTR region could be reversed by their overexpression vectors, indicating that shRNAs do not have off-target effects (Fig. 2E–H). Overexpression of RNA exonucleases EXOSC10 significantly down-regulated the mRNA stability of YAP, but this effect was reversed by overexpression of NSUN2 and ALYREF (Fig. 2I, J and Fig. S2C), suggesting that YAP m5C modification repressed its 3′-5′mRNA degradation. The above results prove that m5C modification can occur in the 3′ UTR region of YAP, which can increase the stability of YAP mRNA.
A The enrichment of ALYREF and YBX1 in A549 and H1299 cells at P1 region of YAP 3′ UTR was calculated as the percentage of the input RNA via RNA-IP using the anti-ALYREF and anti-YBX1 antibodies. B YAP mRNA level was measured by qPCR in A549 and H1299 cells with ALYREF or YBX1 overexpressed with or without NSUN2 knockdown. C–D YAP protein level was measured by IB in A549 (C) and H1299 (D) cells with ALYREF or YBX1 overexpressed with or without NSUN2 knockdown. The level of proteins was normalized to that of GAPDH, and the normalized level of proteins in negative control groups were arbitrarily set to 1. E–J YAP mRNA stability was measured in A549 and H1299 cells with NUSN2 (E–F), ALYREF (G–H) and EXOSC10 (I–J) overexpressed or knocked down at indicated time after ActD treatment. The data are shown as the means ± SD from three independent experiments. Images of IB are representative ones of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance. The data from (A–B) were analyzed by a one-way ANOVA test. The data from (E–J) were analyzed by a two-way ANOVA test.
m5C modification of YAP inhibits miR-582-3p binding
We further explored the mechanism by which YAP m5C modification affects YAP expression. Firstly, we found that although NSUN2 and ALYREF knockdown significantly inhibited YAP m5C modification (Fig. S3A, B), they had no influence on the protein stability, promoter activity, phosphorylation and O-GlcNAcylation of YAP (Fig. 3A–C, Fig. S3C–D). In previous studies, we have found that miR-582-3p can bind to the 3′ UTR region of YAP to promote YAP mRNA degradation [31]. Here, we observed that when the predicted binding motif close to the 5′end (315–321 region in YAP 3′ UTR) was mutated (shown as Mut1-YAP-3′ UTR), the miR-582-3p mimics and miR-582-3p-inhibitors lost their abilities to regulate the activity of YAP 3′ UTR. However, miR-582-3p still significantly regulated the luciferase activities of the other two reporters with mutated 1002–1008 or 2528–2534 region in YAP 3′ UTR (two predicted miR-582-3p binding motifs in 3′ UTR), validating that the 315–321 region in YAP 3′ UTR might be the miR-582-3p binding motif in both liver cancer and LUAD cell lines (Fig. 3D, E and Fig. S3E) [31]. Overexpression of NSUN2 and ALYREF could significantly inhibit the binding of YAP 3′ UTR to miR-582-3p (Fig. 3F, Fig. S3F), enhance the m5C modification while inhibiting the m6A modification (which can promote the binding of YAP 3′ UTR and miR-582-3p [31]) (Fig. 3G, Fig. S3G). Since m5C modification occurs in GC-rich sequences [38,39,40], and m5C modification of YAP 3′ UTR P1 segment was closely related to miR-582-3p binding and m6A modification, we searched for three GC-rich motifs near the miR-582-3p binding m6A modification sites, and mutated G and C to A (Fig. 3H). Compared to WT-YAP-3′ UTR, Mut6-YAP 3′ UTR (328~331 region mutated) significantly inhibited YAP m5C modification, whereas Mut4 (301–308 region mutated) and Mut5 (313–318 region mutated) had no influence (Fig. 3I, J). The above data suggested that YAP m5C modification inhibited the binding between miR-582-3p and YAP 3′ UTR to prevent YAP degradation, and 328–331 motif might be the key motif of YAP m5C modification.
A YAP protein stability was measured in A549 cells with NSUN2 or ALYREF knockdown at indicated time after CHX treatment. B YAP -1k promoter activity was measured in A549 and H1299 cells with NUSN2 or ALYREF overexpressed or knocked down. C YAP phosphorylation and O-GlcNAcylation was measured in A549 and H1299 cells with NUSN2 or ALYREF overexpressed or knocked down. The level of proteins was normalized to that of co-immunoprecipitated YAP, and the normalized level of proteins in negative control groups were arbitrarily set to 1. The YAP level in each co-IP samples was adjusted to the same content. D WT and potential miR-582-3p binding motif mutation YAP 3′ UTR sequences were presented. Red fonts represented the mutant bases. E Activities of WT-YAP-3′ UTR, Mut1-YAP-3′ UTR, Mut2-YAP-3′ UTR, or Mut3-YAP-3′ UTR were measured in A549 cells with miR-582-3p overexpressed or knocked down. F miR-582-3p in NSUN2 with or without ALYREF overexpressed A549 cell lysis was pulled down and enriched with a YAP 3’ UTR P1 region probe and then detected by qPCR. G The enrichment of m6A and m5C in NSUN2 with or without ALYREF overexpressed A549 cells at P1 region of YAP 3′ UTR was calculated as the percentage of the input RNA via RNA-IP using the anti-m6A and anti-m5C antibodies. H WT and potential m5C modification motif mutation YAP 3′ UTR sequences were presented. Red fonts represented the mutant bases. I, J WT-YAP-3′ UTR, Mut4-YAP-3′ UTR, Mut5-YAP-3′ UTR, or Mut6-YAP-3′ UTR was overexpressed in YAP knockdown A549 (I) or H1299 (J) cells. The enrichment of m5C at P1 region of YAP 3′ UTR was calculated as the percentage of the input RNA via RNA-IP using the m5C antibodies. The data are shown as the means ± SD from three independent experiments. Images of IB are representative ones of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance. NS, non-significant. The data from (A) were analyzed by a two-way ANOVA test. The data from (B, E–G) were analyzed by a one-way ANOVA test. The data from (I, J) were analyzed by a student’s t test.
YAP stimulates exosome secretion via Mycn- and SOX10-dependent transcription
Whether YAP and its m5C modification regulate exosome? We found that YAP promoted exosome secretion of A549 and H1299 cells, and NSUN2 and ALYREF could further promote this ability (Fig. 4A). When YAP was knocked out, the exosome secretion of A549 and H1299 cells decreased, and this effect could not be reversed by NSUN2 and ALYREF, suggesting that the ability of NSUN2 and ALYREF to promote exosome secretion depends on YAP (Fig. 4B). PKH67 staining experiments further confirmed the above suggestions (Fig. 4C, Fig. S4A). We also found that YAP protein level was positively associated with exosome concentrations in paired of LUAD plasma and tissue samples (Fig. S4B). The exosomes were examined by transmission electron microscope (TEM) analysis (Fig. 4D). The exosome biomarkers including CD63, TSG101, ALIX and CD9 were detected, and endoplasmic reticulum (ER) biomarker calnexin was used as the negative control to validate the quality of the exosome (Fig. S4C). However, the size and concentration of exosomes in A549 and H1299 cells were close, and YAP, NSUN2 and ALYREF could not significantly change the size of exosomes (Fig. 4E–G).
A–B Exosome concentration was measured in A549 and H1299 cells with YAP overexpressed (A) or knocked out (B), with or without NSUN2 and ALYREF overexpressed. C The exosome from H1299 cells with YAP, NSUN2 with or without ALYREF overexpression was marked by PKH67 (green) and incubated with A549 cells. Afterwards, IF was performed for the detection of exosome. Scale bar, 50 µm. D Representative TEM images of plasma exosome in A549 and H1299 cells. Scale bar, 300 nm. E Relationship between exosome concentration and size in A549 and H1299 cells. The mean and mode size of exosome were shown. F Exosome concentration in A549 and H1299 cells. G Relationship between exosome concentration and size in A549 cells with YAP, NSUN2 with or without ALYREF overexpression. The mean and mode size of exosome were shown. The data are shown as the means ± SD from 3 to 5 independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance. NS, non-significant. The data from (A–C) were analyzed by a one-way ANOVA test. The data from (E, G) were analyzed by a two-way ANOVA test. The data from (F) were analyzed by a student’s t test.
We further explored how YAP stimulates exosome secretion. We screened a series exosome biogenesis, transport and release-related genes to evaluate whether they were regulated by YAP [41,42,43]. We found YAP overexpression increased, while YAP knockout decreased mRNA level of HRS, PLD2, RAB2B, RAB27A, RAB27B, VAMP and ATG7 (Fig. 5A, B and Fig. S5A). Using LASAGNA database, we found that there were binding motifs of Mycn, SOX10 and SPI1 in the promoter regions of the above 7 genes (Fig. 5C). Compared to SPI1, Mycn and SOX10 could significantly promote exosome secretion, and the mRNA levels of 7 exosome secreting genes (Fig. 5D, E, Fig. S5B). In addition, Mycn or SOX10 knockout partially reversed, while Mycn and SOX10 simultaneous knockout almost completely reversed the promotion effect of YAP on exosome secretion, as well as mRNA and protein expression of 7 exosomal secreting genes (Fig. 5F, G, Fig. S5C–E). Also, NSUN2 single knockout partially reversed, while NSUN2 and ALYREF simultaneous knockout almost completely reversed the promotion effect of YAP, SOX10 and Mycn (Fig. 5H, I, Fig. S5F–G). In all, these data revealed a YAP-mediated of SOX10 and Mycn exosome secretion-promoting transcription system.
A Genes related to exosome biogenesis, transport and release were listed. B Indicated mRNA levels were measured by qPCR in YAP overexpressed or knocked out A549 cells. C Mycn, SOX10 and SPI1 binding motif found from JASPAR database. D, E Exosome concentration (D) and HRS, PLD2, RAB2B, RAB27A, RAB27B, VAMP and ATG7 mRNA levels (E) in A549 cells with Mycn, SOX10 or SPI1 overexpressed. F, G Exosome concentration (F) and HRS, PLD2, RAB2B, RAB27A, RAB27B, VAMP and ATG7 protein levels (G) in A549 cells with YAP overexpressed with or without Mycn or SOX10 knockout. The level of proteins was normalized to that of GAPDH, and the normalized level of proteins in negative control groups were arbitrarily set to 1. H, I Exosome concentration (H) and HRS, PLD2, RAB2B, RAB27A, RAB27B, VAMP and ATG7 protein levels (I) in A549 cells with indicated genes overexpressed or knocked out. The level of proteins was normalized to that of GAPDH, and the normalized level of proteins in negative control groups were arbitrarily set to 1. The data are shown as the means ± SD from three independent experiments. Images of IB are representative ones of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance. The data from (B) were analyzed by a student’s t test. The data from (D–F, H) were analyzed by a one-way ANOVA test.
YAP m5C modification and downstream exosome secretion-promoting transcription system are essential for LUAD transformation phenotypes
Next, we investigated how YAP-mediated exosome secretion-promoting transcription system regulated transformation phenotypes of LUAD cells. We observed that YAP could induce cell viability, colony formation while reduce caspase 3/7 activity, and these effects were further enhanced by SOX10 and Mycn overexpression and reversed by knockout of NSUN2 and ALYREF. Similarly, NSUN2 and ALYREF could further enhance YAP-mediated increasing cell viability, colony formation and decreasing caspase 3/7 activity, whereas these effects could be reversed by an exosome inhibitor GW4869 treatment (Fig. 6A–D, Fig. S6A–D). We used CCK8 experiments to measure the cell viability in control, YAP overexpression, YAP, SOX10 and Mycn overexpression, as well as YAP, NSUN2 and ALYREF overexpression cells, and added GW4869 to each group to eliminate the influence of exosome. Interestingly, we found that there was no statistically significant difference in cell viability among the four groups (Fig. S6E), which excluded the influence of other factors except for exosome on cell proliferative capacity. Xenograft experiments revealed that YAP co-expressed with Mycn and SOX10 significantly increased tumor volume, whereas this effect could be reversed by NSUN2 and ALYREF knockout; also, YAP co-expressed with NSUN2 and ALYREF significantly raised tumor volume, and this effect could be reversed by GW4869 treatment (Fig. 6E–H). Next, we established two series of LUAD patient derived xenograft (PDX) models, and the morphology of the tissue was verified by H&E staining. (Fig. S6F). PDX1 highly expressed YAP and NSUN2, while PDX2 lowly expressed YAP and NSUN2 (Fig. 6I–K). Compared to PDX2, the exosome concentration and YAP m5C level were much higher in PDX1 (Fig. S6G, H), and the transplanted tumor size in PDX1 was significantly larger (Fig. 6L). These data suggested YAP m5C modification and its downstream exosome secretion-promoting transcription system are both important for transformation phenotypes of LUAD.
A, B Cell viability was measured in A549 or H1299 cells with indicated genes overexpressed or knocked out. C, D Colony formation was measured in A549 or H1299 cells with indicated genes overexpressed or knocked out, and represented images were shown, scale bar, 100 µm. E–H Xenografts formed by A549 cells with indicated treatment (Group 1: YAP overexpression in WT cells with DMSO treatment. Group 2: YAP, SOX10 and Mycn overexpression in WT cells with DMSO treatment. Group 3: YAP, SOX10 and Mycn overexpression in ALYREF−/− and NSUN2−/− cells with DMSO treatment. Group 4: YAP, NSUN2 and ALYREF overexpression in WT cells with DMSO treatment. Group 5: YAP, NSUN2 and ALYREF overexpression in WT cells with GW4869 treatment (2 mg/kg, 3 weeks). Legends for each group (n = 5) were summarized in (E). Represented xenograft images were shown in (F). YAP, SOX10, Mycn, ALYREF and NSUN2 expressions were validated by IB (G). Tumor volume was monitored for 36 days (H). The level of proteins was normalized to that of GAPDH, and the normalized level of proteins in negative control groups were arbitrarily set to 1. I YAP expression in PDX1 and PDX2 tissues as measured by IHC, scale bar, 100 µm. J, K YAP (J) and NSUN2 (K) protein level in PDX1 and PDX2 tissues. L Tumor volume of PDX1 and PDX2 as monitored in 14 days. The data are shown as the means ± SD from 3 to 5 independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance. The data from (A, B) were analyzed by a one-way ANOVA test. The data from (H, L) were analyzed by a two-way ANOVA test. The data from (J, K) were analyzed by a student’s t test.
YAP m5C modification and downstream exosome secretion-promoting transcription system lead to AZD9291 resistance
We also explored whether YAP-mediated exosome secretion-promoting transcription system would cause resistance of third generations of EGFR-TKI. We used AZD9291-resistant H1975 and HCC827 cell lines from previous studies to investigate this question [5]. Through IC50 value determination and 3D cell culture, we observed that knockout of NSUN2 and ALYREF significantly inhibited the AZD9291 resistance of LUAD cells, and GW4869 treatment further aggravated this phenomenon. On the contrary, the overexpression of YAP stimulated the resistance of LUAD cells to AZD9291, while the overexpression of Mycn and SOX10 further promoted this phenomenon (Fig. 7A–F, Fig. S7A–B). We also found that protein levels of YAP and its related exosome secreting genes, YAP m5C modification level and exosome concentration in AZD9291 drug-resistant cells were significantly higher than those in non-drug-resistant LUAD cells (Fig. 7G–I). Therefore, we suggested that YAP-mediated exosome secretion-promoting transcription system would lead to resistance of third generations of EGFR-TKI in LUAD.
A–D IC50 was calculated in AZD9291 (24 h) treated H1975 AZD9291 resistant (H1975Res, A), normal H1975 (B), HCC827Res (C), normal HCC827 (D) cells with indicated treatment. GW4869 (10 µM, 24 h). E 3D spheroids were generated by NSUN2 with or without ALYREF knocked out H1975Res and HCC827Res cells. After the formation of 3D spheroids, AZD9291 (10 µM, 24 h) with or without GW4869 (10 µM, 24 h) was treated. Then cells were stained with SYTOX green. Representative images and graphed data are shown, Scale bar, 50 µm. F 3D spheroids were generated by YAP with or without Mycn and SOX10 overexpressed H1975 and HCC827 cells. After the formation of 3D spheroids, AZD9291 (10 µM, 24 h) was treated. Then cells were stained with SYTOX green. Representative images and graphed data are shown, Scale bar, 50 µm. G–I Indicated protein levels (G), YAP m5C level (H), exosome concentration (I) were measured in H1975, H1975Res, HCC827 and HCC827Res cells. The level of proteins was normalized to that of GAPDH, and the normalized level of proteins in negative control groups were arbitrarily set to 1. The data are shown as the means ± SD from 3 to 5 independent experiments. Images of IB are representative ones of three independent experiments. *p < 0.05, **p < 0.01 indicate statistical significance. The data from (A–D) were analyzed by a two-way ANOVA test. The data from (H, I) were analyzed by a student’s t test.
Exosome autocrine has no additional effect for LUAD cell viability and resistance to AZD9291
In order to evaluate the autocrine effect of the exosome, we first used cell culture medium with the same treatment to replace the original medium at 4, 8 and 16 h after finishing cell transfection (24 h after transfection) to weaken the autocrine effect of the exosome. However, we found that such treatment did not affect cell viability in the control group, YAP overexpression group, YAP, SOX10 and Mycn overexpression group, as well as YAP, SOX10 and Mycn overexpression combined with NSUN2 and ALYREF knockout group (Fig. S7C). Subsequently, we used cell culture medium with the same treatment to replace the original medium at 4, 8 and 16 h after AZD9291 treatment to weaken the autocrine effect of the exosome when we measure the IC50 in the control group, YAP overexpression group, YAP, SOX10 and Mycn overexpression group, as well as YAP, SOX10 and Mycn overexpression combined with NSUN2 and ALYREF knockout group. Similarly, we observed that the IC50 value of AZD9291 treated cells was not affected by the change of culture medium (Fig. S7D). These data suggested that exosome autocrine did not additionally affect LUAD cell viability and resistance to AZD9291.
Discussion
As an important signal node in cells, YAP can control organ size and promote tumor development by regulating transcription [44]. Therefore, it is possible to find a more effective way to inhibit YAP by analyzing its upstream regulation mechanism. In addition to Hippo phosphorylation kinase cascade [11], as well as ERBB2, CREB, and HBP [7, 45, 46], YAP is also regulated by Wnt, PI3K/AKT/mTOR, GPCR signaling, SFKs and VGLL4 [47,48,49]. These findings confirm that the mechanisms by which YAP is regulated are very broad. RNA modifications also regulate YAP by affecting the stability of YAP mRNA. For instance, FTO depletion increased YAP m6A modification at mRNA 3′ UTR, accelerating the degradation of YAP mRNA [50]. Our Lab also confirmed that m6A modification of YAP 3′ UTR leads to the degradation of YAP mRNA [31]. Here, we found a m5C modification motif near the m6A modification motif. The increase of m5C modification led to the decrease of m6A modification and reduced the binding of miR-582-3p to YAP 3′ UTR. It’s possible that in the 3′ UTR region of YAP, there is a dynamic balance of m5C and m6A modification, thereby regulating the stability of YAP mRNA.
In this study, we divided the 3′ UTR of YAP into five parts to explore its m5C modification. First of all, when exploring the m5C modification of YAP 3′UTR, it must be divided to investigate, otherwise it would be difficult to find the exact m5C site, and the long product would reduce the amplification efficiency of qPCR [51]. As for why the 3′UTR of YAP was further subdivided, the first reason is that the complexity of the experiment would be increased; the second reason is that there are more m5C sites besides our findings, and too much subdivision would reduce the statistical significance among each segment; and the third reason is that five divided segments were sufficient for subsequent exploration. Therefore, there was no further subdivision in our research. Of course, it could be seen from the results that the m5C-modified region of YAP 3′UTR is by no means limited to the 328–331 motif we found, and there might be different situations in different cell lines. In follow-up studies, researchers can further develop a more comprehensive discussion of YAP m5C modifications by exploring the status of m5C modifications in other YAP 3′UTR regions, as well as m5C modifications occurring in other YAP mRNA regions.
Recently, researchers found that YAP can promote cancer by regulating the tumor microenvironment [1]. For example, YAP stimulates the transcription of PDL1, and YAP-induced PDL1 expression protects tumor cells from T cell cytotoxicity, and treatment with PD1 inhibitors restores T cell effector functions, resulting to the killing of YAP-expressing tumor cells [52]. In addition, YAP is upregulated in Treg cells to enforce their capacity to suppress antitumor immunity [53]. Exosomes are master cellular regulators in tumor microenvironment and critical messengers that participate in cell-to-cell communications [54]. There are some reports on the regulation of YAP by components in tumor exosomes. For example, Wnt5a, Wnt5b, long non-coding ASMTL-AS1 in tumor cell exosomes all have the ability to activate YAP to promote tumor formation and development [55, 56]. However, studies of YAP regulating exosome generation are rare. In addition, no studies have shown that the 3′ UTR of YAP is related to the secretion of exosomes at present. Our study demonstrated that the 3′ UTR of YAP is modified by m5C, and this modification depends on NSUN2 and ALYREF. We made it clear that NSUN2 and ALYREF could promote exosome secretion only in the presence of YAP. Therefore, these data innovatively demonstrated that YAP 3′ UTR could play a role in promoting exosome secretion. Subsequently, by screening a series of exosome biogenesis, transport and release genes, we found that YAP simultaneously promoted the transcription of HRS, PLD2, RAB2B, RAB27A, RAB27B, VAMP, ATG7, and the transcription factors that played a direct role in promoting transcription were Mycn and SOX10, which were the first time found to act as YAP-related transcription factors.
High expression of YAP leads to resistance to various types chemotherapies, immunotherapies and targeted therapies. For example, taxane is a major class of anti-microtubule chemotherapy drugs widely used in treatment of ovarian, lung, breast, cervical and prostate cancer and acts to stabilize GDP-bound tubulin, thereby preventing microtubule disassembly during cell cycle [57]. miRNA-363, which is upregulated in taxane-resistant cancer cells, confers resistance by degrading LATS mRNA, which in turn dephosphorylates and activates YAP [58]. The transcriptional activation of multiple downstream genes of YAP can also lead to chemoresistance. For example, YAP activation upregulates PD-L1 transcription and leads to immunotherapy resistance [52], and is a major resistance mechanism to EGFR inhibitors [19]. Our study is the first to clarify that YAP is associated with resistance to third-generation EGFR-TKIs, and this resistance may be related to the YAP m5C modification and the its function of promoting exosome secretion. It has been reported that exosomes can lead to resistance of third-generation EGFR-TKI [59, 60], and other key downstream genes of YAP promoting resistance of third-generation EGFR-TKI remain to be explored.
In conclusion, YAP m5C modification increases its mRNA stability and promotes the Mycn- and SOX10-dependent transcription of exosome secretion-related genes in LUAD (Fig. 8). Targeting YAP m5C modification will be helpful for future LUAD treatment.
Briefly, in LUAD, YAP m5C modification on its 3′UTR depends on NSUN2 and ALYREF, and this m5C modification inhibits its m6A modification and miR-582-3p binding. Furthermore, YAP m5C modification increases YAP mRNA stability and stimulates Mycn- and SOX10-dependent exosome secretion-related genes including HRS, PLD2, RAB2B, RAB27A, RAB27B, VAMP and ATG7.
Data availability
All data generated or analyzed during this study are included in this paper and its supplementary files.
References
Zanconato F, Cordenonsi M, Piccolo S. YAP and TAZ: a signalling hub of the tumour microenvironment. Nat Rev Cancer. 2019;19:454–64.
Pearson JD, Huang K, Pacal M, McCurdy SR, Lu S, Aubry A, et al. Binary pan-cancer classes with distinct vulnerabilities defined by pro- or anti-cancer YAP/TEAD activity. Cancer Cell. 2021;39:1115–34.e12.
Jang JW, Kim MK, Lee YS, Lee JW, Kim DM, Song SH, et al. RAC-LATS1/2 signaling regulates YAP activity by switching between the YAP-binding partners TEAD4 and RUNX3. Oncogene. 2017;36:999–1011.
Wang Y, Qiu S, Wang H, Cui J, Tian X, Miao Y, et al. Transcriptional Repression of Ferritin Light Chain Increases Ferroptosis Sensitivity in Lung Adenocarcinoma. Front Cell Dev Biol. 2021;9:719187.
Zhang X, Yu K, Ma L, Qian Z, Tian X, Miao Y, et al. Endogenous glutamate determines ferroptosis sensitivity via ADCY10-dependent YAP suppression in lung adenocarcinoma. Theranostics. 2021;11:5650–74.
Zhang X, Sun F, Qiao Y, Zheng W, Liu Y, Chen Y, et al. TFCP2 Is Required for YAP-Dependent Transcription to Stimulate Liver Malignancy. Cell Rep. 2017;21:1227–39.
Zhang X, Qiao Y, Wu Q, Chen Y, Zou S, Liu X, et al. The essential role of YAP O-GlcNAcylation in high-glucose-stimulated liver tumorigenesis. Nat Commun. 2017;8:15280.
Zhou B, Flodby P, Luo J, Castillo DR, Liu Y, Yu FX, et al. Claudin-18-mediated YAP activity regulates lung stem and progenitor cell homeostasis and tumorigenesis. J Clin Investig. 2018;128:970–84.
Chaib I, Karachaliou N, Pilotto S, Codony Servat J, Cai X, Li X, et al. Co-activation of STAT3 and YES-Associated Protein 1 (YAP1) Pathway in EGFR-Mutant NSCLC. J Natl Cancer Inst. 2017;109:djx014.
Chen HY, Yu SL, Ho BC, Su KY, Hsu YC, Chang CS, et al. R331W Missense Mutation of Oncogene YAP1 Is a Germline Risk Allele for Lung Adenocarcinoma With Medical Actionability. J Clin Oncol. 2015;33:2303–10.
Moya IM, Halder G. Hippo-YAP/TAZ signalling in organ regeneration and regenerative medicine. Nat Rev Mol Cell Biol. 2019;20:211–26.
Qin S, Jiang J, Lu Y, Nice EC, Huang C, Zhang J, et al. Emerging role of tumor cell plasticity in modifying therapeutic response. Signal Transduct Target Ther. 2020;5:228.
Jin D, Guo J, Wu Y, Yang L, Wang X, Du J, et al. m(6)A demethylase ALKBH5 inhibits tumor growth and metastasis by reducing YTHDFs-mediated YAP expression and inhibiting miR-107/LATS2-mediated YAP activity in NSCLC. Mol Cancer. 2020;19:40.
Zhang X, Li Y, Ma Y, Yang L, Wang T, Meng X, et al. Yes-associated protein (YAP) binds to HIF-1alpha and sustains HIF-1alpha protein stability to promote hepatocellular carcinoma cell glycolysis under hypoxic stress. J Exp Clin Cancer Res. 2018;37:216.
Thomann S, Weiler SME, Wei T, Sticht C, De La Torre C, Toth M, et al. YAP-induced Ccl2 expression is associated with a switch in hepatic macrophage identity and vascular remodelling in liver cancer. Liver Int. 2021;41:3011–23.
Kamerkar S, LeBleu VS, Sugimoto H, Yang S, Ruivo CF, Melo SA, et al. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature. 2017;546:498–503.
Sung BH, von Lersner A, Guerrero J, Krystofiak ES, Inman D, Pelletier R, et al. A live cell reporter of exosome secretion and uptake reveals pathfinding behavior of migrating cells. Nat Commun. 2020;11:2092.
Jia Y, Chen Y, Wang Q, Jayasinghe U, Luo X, Wei Q, et al. Exosome: emerging biomarker in breast cancer. Oncotarget. 2017;8:41717–33.
Nguyen CDK, Yi C. YAP/TAZ Signaling and Resistance to Cancer Therapy. Trends Cancer. 2019;5:283–96.
Koba H, Kimura H, Yoneda T, Ogawa N, Tanimura K, Tambo Y, et al. NOTCH alteration in EGFR-mutated lung adenocarcinoma leads to histological small-cell carcinoma transformation under EGFR-TKI treatment. Transl Lung Cancer Res. 2021;10:4161–73.
Akamatsu H, Toi Y, Hayashi H, Fujimoto D, Tachihara M, Furuya N, et al. Efficacy of Osimertinib Plus Bevacizumab vs Osimertinib in Patients With EGFR T790M-Mutated Non-Small Cell Lung Cancer Previously Treated With Epidermal Growth Factor Receptor-Tyrosine Kinase Inhibitor: West Japan Oncology Group 8715L Phase 2 Randomized Clinical Trial. JAMA Oncol. 2021;7:386–94.
Tamiya M, Kunimasa K, Nishino K, Matsumoto S, Kawachi H, Kuno K, et al. Successful treatment of an osimertinib-resistant lung adenocarcinoma with an exon 18 EGFR mutation (G719S) with afatinib plus bevacizumab. Investig N Drugs. 2021;39:232–6.
Leonetti A, Sharma S, Minari R, Perego P, Giovannetti E, Tiseo M. Resistance mechanisms to osimertinib in EGFR-mutated non-small cell lung cancer. Br J Cancer. 2019;121:725–37.
Zhao B, Li L, Tumaneng K, Wang CY, Guan KL. A coordinated phosphorylation by Lats and CK1 regulates YAP stability through SCF(beta-TRCP). Genes Dev. 2010;24:72–85.
Mayr C. What Are 3’ UTRs Doing? Cold Spring Harb Perspect Biol. 2019;11:a034728.
Chu WK, Hung LM, Hou CW, Chen JK. Heterogeneous ribonucleoprotein F regulates YAP expression via a G-tract in 3’UTR. Biochim Biophys Acta Gene Regul Mech. 2019;1862:12–24.
Zhang C, Chen Y, Sun B, Wang L, Yang Y, Ma D, et al. m(6)A modulates haematopoietic stem and progenitor cell specification. Nature. 2017;549:273–6.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149:1635–46.
Sun M, Song H, Wang S, Zhang C, Zheng L, Chen F, et al. Integrated analysis identifies microRNA-195 as a suppressor of Hippo-YAP pathway in colorectal cancer. J Hematol Oncol. 2017;10:79.
Jin D, Guo J, Wu Y, Chen W, Du J, Yang L, et al. Metformin-repressed miR-381-YAP-snail axis activity disrupts NSCLC growth and metastasis. J Exp Clin Cancer Res. 2020;39:6.
Zhang X, Xu Y, Qian Z, Zheng W, Wu Q, Chen Y, et al. circRNA_104075 stimulates YAP-dependent tumorigenesis through the regulation of HNF4a and may serve as a diagnostic marker in hepatocellular carcinoma. Cell Death Dis. 2018;9:1091.
Chen R, Qian Z, Xu X, Zhang C, Niu Y, Wang Z, et al. Exosomes-transmitted miR-7 reverses gefitinib resistance by targeting YAP in non-small-cell lung cancer. Pharm Res. 2021;165:105442.
Wnuk M, Slipek P, Dziedzic M, Lewinska A. The Roles of Host 5-Methylcytosine RNA Methyltransferases during Viral Infections. Int J Mol Sci. 2020;21:8176.
Nombela P, Miguel-Lopez B, Blanco S. The role of m(6)A, m(5)C and Psi RNA modifications in cancer: Novel therapeutic opportunities. Mol Cancer. 2021;20:18.
Bohnsack KE, Hobartner C, Bohnsack MT. Eukaryotic 5-methylcytosine (m(5)C) RNA Methyltransferases: Mechanisms, Cellular Functions, and Links to Disease. Genes (Basel) 2019;10:102.
Chellamuthu A, Gray SG. The RNA Methyltransferase NSUN2 and Its Potential Roles in Cancer. Cells. 2020;9:1758.
Zhang X, Xu Y, Ma L, Yu K, Niu Y, Xu X, et al. Essential roles of exosome and circRNA_101093 on ferroptosis desensitization in lung adenocarcinoma. Cancer Commun (Lond). 2022;42:287–313.
Huang T, Chen W, Liu J, Gu N, Zhang R. Genome-wide identification of mRNA 5-methylcytosine in mammals. Nat Struct Mol Biol. 2019;26:380–8.
Meng B, Epp N, Wijaya W, Mrazek J, Hoover TR. Methylation Motifs in Promoter Sequences May Contribute to the Maintenance of a Conserved (m5)C Methyltransferase in Helicobacter pylori. Microorganisms 2021;9:2474.
Huo W, Adams HM, Zhang MQ, Palmer KL. Genome Modification in Enterococcus faecalis OG1RF Assessed by Bisulfite Sequencing and Single-Molecule Real-Time Sequencing. J Bacteriol. 2015;197:1939–51.
Qin X, Guo H, Wang X, Zhu X, Yan M, Wang X, et al. Exosomal miR-196a derived from cancer-associated fibroblasts confers cisplatin resistance in head and neck cancer through targeting CDKN1B and ING5. Genome Biol. 2019;20:12.
Isaac R, Reis FCG, Ying W, Olefsky JM. Exosomes as mediators of intercellular crosstalk in metabolism. Cell Metab. 2021;33:1744–62.
Kalluri R, LeBleu VS. The biology, function, and biomedical applications of exosomes. Science 2020;367:eaau6977.
Yu FX, Zhao B, Guan KL. Hippo Pathway in Organ Size Control, Tissue Homeostasis, and Cancer. Cell. 2015;163:811–28.
Aharonov A, Shakked A, Umansky KB, Savidor A, Genzelinakh A, Kain D, et al. ERBB2 drives YAP activation and EMT-like processes during cardiac regeneration. Nat Cell Biol. 2020;22:1346–56.
Wang J, Ma L, Weng W, Qiao Y, Zhang Y, He J, et al. Mutual interaction between YAP and CREB promotes tumorigenesis in liver cancer. Hepatology. 2013;58:1011–20.
Taniguchi K, Wu LW, Grivennikov SI, de Jong PR, Lian I, Yu FX, et al. A gp130-Src-YAP module links inflammation to epithelial regeneration. Nature. 2015;519:57–62.
Piccolo S, Dupont S, Cordenonsi M. The biology of YAP/TAZ: hippo signaling and beyond. Physiol Rev. 2014;94:1287–312.
Zhang W, Gao Y, Li P, Shi Z, Guo T, Li F, et al. VGLL4 functions as a new tumor suppressor in lung cancer by negatively regulating the YAP-TEAD transcriptional complex. Cell Res. 2014;24:331–43.
Li DQ, Huang CC, Zhang G, Zhou LL. FTO demethylates YAP mRNA promoting oral squamous cell carcinoma tumorigenesis. Neoplasma. 2021;69:71–9.
Rodriguez A, Rodriguez M, Cordoba JJ, Andrade MJ. Design of primers and probes for quantitative real-time PCR methods. Methods Mol Biol. 2015;1275:31–56.
Kim MH, Kim CG, Kim SK, Shin SJ, Choe EA, Park SH, et al. YAP-Induced PD-L1 Expression Drives Immune Evasion in BRAFi-Resistant Melanoma. Cancer Immunol Res. 2018;6:255–66.
Ni X, Tao J, Barbi J, Chen Q, Park BV, Li Z, et al. YAP Is Essential for Treg-Mediated Suppression of Antitumor Immunity. Cancer Disco. 2018;8:1026–43.
Mathieu M, Martin-Jaular L, Lavieu G, Thery C. Specificities of secretion and uptake of exosomes and other extracellular vesicles for cell-to-cell communication. Nat Cell Biol. 2019;21:9–17.
Tao SC, Yuan T, Zhang YL, Yin WJ, Guo SC, Zhang CQ. Exosomes derived from miR-140-5p-overexpressing human synovial mesenchymal stem cells enhance cartilage tissue regeneration and prevent osteoarthritis of the knee in a rat model. Theranostics. 2017;7:180–95.
Ma D, Gao X, Liu Z, Lu X, Ju H, Zhang N. Exosome-transferred long non-coding RNA ASMTL-AS1 contributes to malignant phenotypes in residual hepatocellular carcinoma after insufficient radiofrequency ablation. Cell Prolif. 2020;53:e12795.
Perez EA. Microtubule inhibitors: Differentiating tubulin-inhibiting agents based on mechanisms of action, clinical activity, and resistance. Mol Cancer Ther. 2009;8:2086–95.
Mohamed Z, Hassan MK, Okasha S, Mitamura T, Keshk S, Konno Y, et al. miR-363 confers taxane resistance in ovarian cancer by targeting the Hippo pathway member, LATS2. Oncotarget. 2018;9:30053–65.
Hisakane K, Seike M, Sugano T, Yoshikawa A, Matsuda K, Takano N, et al. Exosome-derived miR-210 involved in resistance to osimertinib and epithelial-mesenchymal transition in EGFR mutant non-small cell lung cancer cells. Thorac Cancer. 2021;12:1690–8.
Wu S, Luo M, To KKW, Zhang J, Su C, Zhang H, et al. Intercellular transfer of exosomal wild type EGFR triggers osimertinib resistance in non-small cell lung cancer. Mol Cancer. 2021;20:17.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82102792), Project of Clinical Research Supporting System, Clinical Medicine First-class Discipline, Scientific Research Project of Shanghai Municipal Health Commission (20194Y0207). Talent training plan of Shanghai Chest Hospital in 2020 (to LM, XZ, and WY). Excellent Talents Nurture Project of Shanghai Chest Hospital (2021YNZYY01, 2021YNZYY02).
Author information
Authors and Affiliations
Contributions
WY, CZ, and YW researched and analyzed data. XT constructed the plasmids. YM performed mice experiments. FM researched and analyzed data. LM, XZ, and JX designed the experiment, interpreted the data and analyzed data. XZ and JX wrote the paper. All authors read and approved the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This study was approved by the Ethics Review Committee of Shanghai Chest Hospital, Shanghai Jiao Tong University School of Medicine (permit number: KS [Y] 21057). All patients completed informed consent forms.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, W., Zhang, C., Wang, Y. et al. YAP 5-methylcytosine modification increases its mRNA stability and promotes the transcription of exosome secretion-related genes in lung adenocarcinoma. Cancer Gene Ther 30, 149–162 (2023). https://doi.org/10.1038/s41417-022-00533-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-022-00533-7
This article is cited by
-
Hippo pathway in cell–cell communication: emerging roles in development and regeneration
Inflammation and Regeneration (2024)
-
CircMAST1 inhibits cervical cancer progression by hindering the N4-acetylcytidine modification of YAP mRNA
Cellular & Molecular Biology Letters (2024)
-
Multi-omic analyses of m5C readers reveal their characteristics and immunotherapeutic proficiency
Scientific Reports (2024)
-
Identification of ALYREF in pan cancer as a novel cancer prognostic biomarker and potential regulatory mechanism in gastric cancer
Scientific Reports (2024)
-
LINC02159 promotes non-small cell lung cancer progression via ALYREF/YAP1 signaling
Molecular Cancer (2023)