Abstract
AlphaFold2 (AF2) is an artificial intelligence (AI) system developed by DeepMind that can predict three-dimensional (3D) structures of proteins from amino acid sequences with atomic-level accuracy. Protein structure prediction is one of the most challenging problems in computational biology and chemistry, and has puzzled scientists for 50 years. The advent of AF2 presents an unprecedented progress in protein structure prediction and has attracted much attention. Subsequent release of structures of more than 200 million proteins predicted by AF2 further aroused great enthusiasm in the science community, especially in the fields of biology and medicine. AF2 is thought to have a significant impact on structural biology and research areas that need protein structure information, such as drug discovery, protein design, prediction of protein function, et al. Though the time is not long since AF2 was developed, there are already quite a few application studies of AF2 in the fields of biology and medicine, with many of them having preliminarily proved the potential of AF2. To better understand AF2 and promote its applications, we will in this article summarize the principle and system architecture of AF2 as well as the recipe of its success, and particularly focus on reviewing its applications in the fields of biology and medicine. Limitations of current AF2 prediction will also be discussed.
Similar content being viewed by others
Introduction
In December of 2020, AlphaFold2 (AF2),1 a machine-learning based model to predict protein structures developed by DeepMind, won the championship in the 14th Critical Assessment of Structure prediction (CASP14).2 One and a half years later, DeepMind and the EMBL’s European Bioinformatics Institute (EMBL-EBI) released structures of more than 200 million proteins predicted by AF2,3 which cover almost all the known proteins on the planet (protein universe). These two events have drawn great attention to AF2 in the science community. AF2 represents a milestone advance in protein structure prediction. It is considered as the greatest contribution of artificial intelligence (AI) to the scientific field and one of the most important scientific breakthroughs made by mankind in the 21st century. This is a very remarkable historical achievement in the human understanding of nature. The high appraisal to AF2 is not excessive because understanding the three-dimensional (3D) structures of proteins is one of the most challenging issues in the field of biology, which has puzzled scientists for 50 years.4 Although multiple technologies including nuclear magnetic resonance (NMR),5 X-ray crystallography,6 and cryo-electron microscopy (cryo-EM)7 have been adopted to solve the protein structures, only about 200,000 proteins’ structures have been determined (https://www.rcsb.org/), covering less than 0.1% of the protein universe.
AF2 is expected to have a significant influence on the fields of biology and medicine, and may change the way we do related researches such as structural biology, drug discovery, protein design, etc. Despite that the time is short since AF2 was developed, there are already many studies related to AF2 reported. To better understand AF2 and promote its applications, we will in this review paper summarize the algorithm and working principle of AF2 and recipe of its success, particularly focus on reviewing its applications in the fields of biology and medicine. Limitations of current AF2 prediction will also be discussed. The remaining part of this paper is organized as follows. We will firstly give a brief introduction to the protein structure prediction, followed by analyzing the principle and architecture of AF2 and the secret of its success. Then we will summarize the applications of AF2 in the fields of biology and medicine, and discuss limitations of current AF2 prediction. It will end in concluding remarks.
A brief introduction to the protein structure prediction
In 1961, Anfinsen8 raised the famous thermodynamic hypothesis of protein folding (“Anfinsen’s dogma”) that a protein’s native structure stands for a free energy minimum determined by its amino acid sequence, or in other words, the 3D structure of a protein is only determined by its amino acid sequence. This hypothesis is the theoretical foundation of protein structure prediction. Since then, people began to look for algorithms to directly predict 3D structures of proteins from amino acid sequences. In the field of protein structure prediction, CASP, founded in 1994, is a milestone event.9,10,11 This competition is held every two years. The CASP committee publishes the “target sequences” globally, for which the experimental structures are known but not yet released. Each participant team that registers for the competition will predict and submit structures of proteins corresponding to the “target sequences” by using their own algorithm within a specified period. Finally, the CASP committee will assess their predicted structures by comparing with those experimentally solved. The competition is double blinded: participants have no access to the experimental structures and referees do not know who make the submissions. Because of the objectivity and fairness, the CASP competition has a very high reputation in structural biology and computational biology communities.
Until now, many algorithms for protein structure prediction have been reported and readers can refer to several recent review papers.12,13,14,15,16,17 Despite vastly different, they can be roughly grouped into three major classes: homology modeling, de novo modeling, and machine learning (ML) -based modeling.
(1) Homology modeling
Homology modeling, also known as comparative modeling or template-based modeling, is based on the hypothesis that proteins’ 3D structures are more conserved than their amino acid sequences, and that therefore similar amino acid sequences should have similar 3D structures.18,19 The homology modeling method mainly uses two techniques: sequence alignment and molecular modeling. The basic workflow of homology modeling is as follows: Given a target amino acid sequence, the first step is to look for its homologous sequences from structure-known protein databases, followed by sequence alignment. Then, coordinates of amino acids of the structure-known homologous proteins are taken as the coordinates of the corresponding amino acids of the target protein. Subsequently, molecular modeling is performed to relax the unfavorable interactions between amino acid pairs. Finally, the generated 3D structure is evaluated.
The homology modeling method is the most popular approach decades ago.19,20,21,22,23,24 Advantages of the homology modeling include simple algorithm, fast prediction speed, and high accuracy for proteins that have structure-known homologs. The defect is that it strongly depends on the template structures, which means that it cannot predict structures of proteins whose homologs’ structures have not been determined.25
(2) De novo modeling
De novo modeling is a protein structure prediction method based on the “first principles”.26 Unlike the homology modeling, the de novo modeling does not depend on the known protein structures, but generating the 3D structure of a target protein only based on the established laws of physics (quantum mechanics). In brief, a de novo modeling method conducts conformation search guided by a designed energy function with the atomic coordinates of amino acids as variables. Many possible conformations are produced in this process and that with the lowest energy is picked. Obviously, the de novo modeling method depends on two factors: (1) an energy function that represents the free energy of target protein with respect to the atomic coordinates of amino acids; (2) an effective conformational search algorithm that can quickly identify low energy states.
There are many investigations regarding protein structure prediction based on de novo modeling.27,28,29,30,31,32,33 The advantages of de novo modeling include: (1) it does not rely on the known protein structures, which means that it is able to predict protein structures where no any prior structural knowledge exists; (2) it has the possibility of finding new protein structural types. Nevertheless, this method faces two major obstacles. The first one is the free energy function. Theoretically, accurate calculation of free energy needs to solve the Schrödinger’s equation, which requires huge amount of calculation that we cannot afford even now. Therefore, empirical formulae have to be used. Currently, a majority of empirical formulae are based on molecular mechanics or Newtonian mechanics. The second one is the conformational space of protein, which is an astronomical number. The possible conformational number of a protein with several hundred amino acids is estimated to be about 10300.34 Although great progresses have been made in conformational search algorithms, as well as computing power and storage space, de novo modeling is still only applicable to small proteins with the number of amino acid residues ranging from 10 to 80.
(3) ML-based modeling
ML-based modeling is a strategy that utilizes ML algorithms and known protein structures to predict the structures of target proteins. Despite many ML algorithms, the most noteworthy is deep learning (DL). DL has achieved rapid development in recent years, which was driven by the fast growing of data volume (“big data”), a large increase in computing power (e.g., GPU, TPU, etc.) and the continuous optimization of DL algorithms (e.g., Recurrent Neural Networks,35 Convolutional Neural Networks,36 Generative Adversarial Networks,37 Transformer,38 etc.). DL has demonstrated its great power in computer vision,39 natural language processing,40 auto-driving,41 and other fields.42,43,44,45,46 Recently, DL has also been applied to the protein structure prediction.12 At present, there are many modeling methods based on DL, among which AlphaFold,1,47 RoseTTAFold,48 ESMFold49 (ESMFold also offers an extensive database of protein structural predictions, which include 617 million metagenomic protein structures) and the recent language model by Chowdhury et al.50 are the most famous ones.
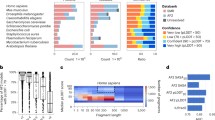
Compared with homology modeling and de novo modeling, the DL-based method is a data-driving approach and is the latest emerging one. Due to the great success of DL in other fields, the DL-based protein prediction approach is expected to have a better performance. Indeed, the DL-based method lived up to people’s expectation and won the champion of CASP1351 and 14.2 Particularly, AF2 in CASP14 could predict the structures of proteins with atomic-level accuracy. Figure 1a summarizes the trend of performance denoted as the backbone accuracy for the best models obtained in each CASP.52 Here the backbone accuracy is measured by the Global Distance Test (GDT_TS)53 value, which is a multi-scale metric to indicate the proximity of the Cα atoms in a model to those in the corresponding structure determined by experiments. The GDT_TS values were calculated respectively according to proteins with different target difficulties: “easy”, “medium”, and “difficult”; an “easy” target implies a protein whose structure is easy to be predicted, for example, a well-folded protein with no loop and structures of its highly homologous proteins being available, while a “difficult” target implies a protein whose structure is difficult to be predicted, for example, a protein with some un-folded domains or many loops, and structures of its homologous proteins being not available. As shown in Fig. 1a, the “easy” proteins can be predicted accurately in CASP1 - CASP14 with GDT_TS values around or larger than 80%. However, for the “medium” and “difficult” proteins, the prediction accuracies were significantly improved only in CASP13 and CASP14. Especially, the GDT_TS values for the “medium” and “difficult” proteins have reached more than 85% in CASP14, largely due to the contribution of AF2; Fig. 1b shows a comparison of the GDT_TS values with or without AF2 prediction included in CASP14. We will elaborate the principle and architecture of AF2 as well as the secret of AF2’ success in the next section.
Performances of protein structure prediction indicated as backbone agreement with that of structures determined by experiments for the best models in CASPs. a The trend of performance (denotated by GDT_TS values) with regard to the backbone accuracy for best models obtained in each CASP. b A comparison of the GDT_TS values with or without AF2 prediction included in CASP14. Prediction accuracies for proteins with different target difficulty (“easy”, “medium” and “difficult”) are presented in indicated colors (blue, gold and green, respectively)
Principle and architecture of AF2 and secrets of AF2’ success
AF2 is the most advanced protein structure prediction method of DeepMind. Its principle is based on the state-of-the-art DL algorithms as well as the conservation of protein structures in evolution. It uses a new end-to-end deep neural network which is trained to generate protein structures from amino acid sequences, by utilizing information of homologous proteins and multiple sequence alignments.
In AF2, some new DL algorithms developed recently are used, of which attention mechanism-based transformer38 plays a critical role in improving AF2’s performance. Transformer is a newly emerging deep neural network, which applies the self-attention mechanism to obtain intrinsic features and displays great potential of broad applications in AI. Transformer38 was first applied in the area of natural language processing (NLP). It is composed of an encoder module and a decoder module with several transformer blocks of the same architecture. Each transformer block is composed of a multi-head attention layer, a feed-forward neural network, shortcut connection and layer normalization.
The conservation of protein structure in evolution is the biological principle behind AF2. A protein is often conservative in evolution, and the evolution is mostly neutral, which means that most of the mutations don’t affect the protein function. More importantly, protein structure is more conservative than its amino acid sequence. Typically, for example, for a sequence that change by 80% between distant species, the 3D structure may remain almost the same. Conservation of a position in alignment usually implies its importance for protein folding or function. Co-evolution of two amino acid residues of a protein often implies interaction between those amino acids. This information has been used as the basis for 3D structure prediction in AF2.
AF2 adopts an architecture that is completely different from that of previous DL models, including AlphaFold1.47 A detailed description for the architecture of AF2 can be found in reference.1 Here we present an overview to the architecture and work principle of AF2. As shown in Fig. 2, the pipeline of AF2 includes three modules.
-
1.
The first one is the input module. Given an amino acid sequence, AF2 finds its homologs in sequence databases and conducts MSA by aligning the input sequence and its homolog sequences. AF2 also checks whether any of the homologs has a 3D structure available in protein structure databases, and constructs a pairwise distance matrix between amino acids. Then AF2 generates MSA representation and pair representation. It should be noted that, although both AF2 and homology modeling use MSA, AF2 extracts and utilizes the co-evolution information from the MSA, but homology modeling does not. Intuitively, when two residues (A and B) are spatially near to each other in the folded structure, the mutations of residue A may provoke a selective pressure for residue B to mutate. Such co-evolutionary information54 detected in MSAs has been utilized to assist the protein structure prediction in AF2.
It is necessary to mention that AF2 uses many high-quality protein sequence databases, including Uniref90,55 Uniclust30,56 MGnify57 and BFD (Big Fantastic Database);1 BFD is a database constructed by the team themselves. Pertaining to the structure databases used for training and as templates, it adopts PDB and PDB7058 respectively. AF2 also utilizes several efficient search algorithms, including JackHMMER59 and HHBlits60 for genetic searching, and HHSearch61 for template searching.
-
2.
The second one is the Evoformer module, which is likely an encoder. In this module, AF2 takes the inputs (MSA representation and pair representation) from the first module and passes them through a deep learning module (called Evoformer). Evoformer produces processed MSA representation and pair representation. The key benefit of using Evoformer blocks is that they are able to switch information between MSA representation and pair representation: the MSA information can be reinterpreted as the pairwise information is improved, and in the similar way, the pairwise information can be further improved as the MSA information is reinterpreted.
The Evoformer contains 48 blocks with weights not shared. Each block has two inputs: an MSA representation and a pair representation. The outputs from each Evoformer block are an updated MSA representation and an updated pair representation (Fig. 2b). The MSA representation and pair representation are processed with several layers. The Dropout approach is also used, which is commonly used for alleviating the problem of overfitting.
Each Evoformer block (Fig. 2b) contains two pathways of transformer-based layers and two “communication channels” between the two pathways. The first pathway of transformer-based layers acts on the MSA. It computes attention over a large matrix of protein symbols. To reduce computational cost, the MSA attention is factorized in row-wise gated self-attention and column-wise gated self-attention components. The row-wise gated self-attention mechanism, allowing the network to identify which pairs of amino acids are more related, constructs attention weights for amino acid pairs. It also combines the information from the input pair representation, and this information can be considered as an extra term. The column-wise gated self-attention, allowing the network to determine which sequences are more informative, enables the components which belong to the same target amino acid to process information exchange. After the row-wise gated self-attention and column-wise gated self-attention steps, the MSA pathway has an MSA transition layer which includes a 2-layer MLP. This trick enhances the attention mechanism and allows it to pinpoint interacting pairs of amino acids.
The second pathway of transformer-based layers acts on the pair representation. The key feature of this network is that attention is arranged in terms of triangles of residues, which is based on the straightforward principle that in a triangle, any two edges can affect the third edge. The intuition here is to enforce the triangle equivariance. As shown in Fig. 2b, the first two rounds of update are triangular multiplicative updates, which are based on non-attention method. Each of the “outgoing” and “incoming” edges obtains an update from another two edges of all the triangles where the edge is included. The second two rounds of update are triangular self-attention. They update the pair representation in the Evoformer block. Two versions are also involved: “starting node” version and “ending node” version. The “starting node” version renews the edge based on all the edges which has the same starting node. The “ending node” version operates the similar way, but it works on the edges which share the same ending node instead. Pairwise representation pathway also contains a transition layer after the triangular self-attention layers, which works the same way as the transition layer introduced above.
-
3.
The third one is the structure module, likely a decoder. The structure module also uses a transformer neural network. It achieves the transition from abstract representation of protein structure to 3D atom coordinates of target proteins. The structure module takes each residue as a separate object and predicts the rotations and translations required to place it.
Schematic work principle and architecture of AF2. a The overall architecture of AF2. The pipeline of AF2 contains three modules. The first one is the input module, which takes an amino acid sequence as input, and generates the MSA representation and the pair representation. The second one is the Evoformer module, which takes the MSA representation and the pair representation from the first module and passes them through the deep learning module, Evoformer. The third one is the structure module, which achieves the transition from abstract representation of protein structure to 3D atom coordinates of target protein. b Components of a block in Evoformer. Evoformer contains 48 blocks with weights not shared. The MSA representation and the pair representation are renewed through each block. c Components of a block in the structure module. Structure module contains 8 blocks with shared weights. Single representation and backbone frames are updated through each block of the structure module
The structure module has three input elements. The first one is the single representation containing the abstract information of the target sequence. It is a linear projection of the MSA representation’s first row. The second one is the pair representation output from the Evoformer module. Backbone frames serve as an additional input. Each residue is represented as a triangle, where its vertex next to obtuse angle is Cα atom, and two other vertices are N atom of amino group and C atom of carbonic acid group. The backbone frames, which are the main part of the system prediction, are formed with triangles of the whole amino acid sequence. At the beginning of the structure module, all the backbone frames are placed at the same point in the same orientation. The structural module’s output is the 3D coordinates of all the protein atoms.
The structure module has 8 blocks with shared weight. Each block (Fig. 2c) updates the single representation and the backbone frames. The critical component of each block is the Invariant Point Attention (IPA), which is a geometry-aware attention mechanism used for updating the single representation. The final attention values of the IPA operation are 3D equivariant, which means that they are invariant to global rigid motion including rotations and translations. After the processing of the IPA operation, the module block predicts relative rotations and translations of each backbone frame. The utilization of these operations enables the overall attention and process equivariantly on the backbone frames. In the next step, the structure module block predicts the side-chain χ angles and computes all atom positions using the updated single representation from IPA and the renewed backbone frames. However, the final output might not meet all the stereochemical constraints. For this reason, AF2 applies Amber relaxation to resolve the violations and clashes without harmfully impact the prediction accuracy. OpenMM62 with Amber99sb force field63 is used for the process.
Finally, AF2 adopts recycling mechanism for three times to process iterative refinement of training and testing; the recycling mechanism has been broadly utilized in computer vision, which allows the network to be deeper and to process multiple versions of the input features without significantly increasing the quantity of parameters or training time. In each recycling, the model incorporates the previous outputs as additional inputs. AF2 recycles the predicted backbone atom coordinates from the structure module, the output pair representations and the first row of MSA representations from the Evoformer.
AF2 has achieved the best performance compared to previous models. Although we have presented the principle and architecture of AF2, the secret of AF2’ success is not explicitly indicated. Here, we present our analysis on the most critical points leading to the success of AF2. From the technological point of view, it is indisputable that the delicate algorithms used are the major causes. Of the most importance is the use of attention mechanism-based transformer. In AF2, several types of attention mechanisms are used, with each one focusing on a specific aspect for the model to learn. In the encoder part, AF2 uses two groups of transformers which are intertwined with each other: one mainly operates on the raw MSA and the other one mainly operates on pairwise information, which update each other through specific information channels between them. The MSA row-wise gated self-attention allows the model to capture long-range dependencies in amino acid sequences and protein structures. The MSA column-wise gated self-attention is a kind of ‘conservation-aware’ attention mechanism, which lets the elements exchange information among species. The triangular self-attention module in the decoder enables the model to learn geometric restrictions within the protein molecules. In the decoder part, AF2 also employs a transformer to geometrically encode residues as a cloud of oriented reference frames in 3D space.
The training method is also a factor which makes AF2 success. The designers utilized the idea of self-distillation.64 They used a combination of PDB and a new self-distillation unlabeled data set of predicted protein structures as the training data to train AF2, among which, 25% of the training example comes from the known structures in PDB while 75% of data was from the new self-distillation data set. The aim is to make AF2 recap the protein structures predicted previously challenging by using different training data augmentation methods. This integration data set approach makes use of the data predicted by AF2 and largely improves the performance of the model.
Other algorithms or tricks that may contribute to the success of AF2 include the use of recycling approach, end-to-end framework for learning from protein data, and so on. Moreover, big data of amino acid sequences and structures also contribute a lot to AF2’s success. The complete sequence library and sufficient number of single domain protein structures allow deep learning neural networks to explore various dependencies in protein sequence and structure, which could be another important intrinsic cause for the success of AF2.
Applications of AF2 in the fields of biology and medicine
The excellent performance of protein structure prediction by AF2 and the release of structures of more than 200 million proteins are reshaping structural biology, and hence will profoundly impact the fields of biology and medicine that require protein structural information. AF2 and its predicted protein structures will enable researchers to have more opportunity to solve problems that are previously thought to be highly challenging. We will in the follows review the progress of applications of AF2 in the fields of biology and medicine. These applications are classified into eight categories: structural biology, drug discovery, protein design, target prediction, protein function prediction, protein-protein interaction, biological mechanism of action, and others (Fig. 3).
Application areas of AF2 in the fields of biology and medicine. AF2 can be applied in many areas of biology and medicine, including structural biology, drug discovery, protein design, protein-protein interaction, target prediction, protein function prediction, biological mechanism of action, and others (such as protein evolution, rare disease treatment studies, effects of mutation on treatment, vaccine design and so on)
Structural biology
Undoubtedly, structural biology is the most impacted area by AF2.65 Rather than saying that AF2 may make structural biologists unemployed, we prefer to the viewpoint that the AF2 and its predicted structures will change the way we do structural biology, including X-ray crystallography, cryo-EM, and NMR spectroscopy. Firstly, predicted structures could be utilized as templates for molecular replacement in solving X-ray crystal structures, implying that traditional selenomethionine phasing is almost not necessary.66,67 Secondly, these predicted structures may also be helpful for structure determination of large protein assemblies by cryo-EM, which usually needs structures of the component proteins or their domains as a starting point for fitting to the cryo-EM densities. Thirdly, one could also benefit from the predicted structures in using NMR to solve protein structures.68,69 Typically, the de novo structure determination of domains or proteins using NMR, which is time-consuming, may be replaced by the AF2 structures. Therefore, the application of AF2 prediction allows to make full use of the advantages of NMR in studying protein folding and dynamics.
Currently there are already many successful applications in this respect. For example, Hu et al.70 utilized X-ray crystallography and AF2 prediction to determine the structure of the VP8* domain (VP8*B) of VP4, which is a spike protein, in group B rotaviruses. In this study, the authors expressed and purified the VP8*B protein. Then they obtained the crystals of this protein and received diffraction data of X-ray. In the process of solving the 3D structure of this protein, instead of using the traditional selenomethionine phasing method, they used AF2 to generate a suitable search model for molecular replacement. The results showed that the AF2 predicted structure almost perfectly matched the diffraction density. Besides the overall fold, AF2 also successfully predicted the orientation of sidechain with high accuracy, which is very close to that determined by experiments. Of note is that they found a novel fold mode by AF2, which has never been reported in homology proteins.
Hutin et al.71 recently revealed a structure of the vaccinia virus DNA helicase, the helicase-primase D5, by utilizing combined cryo-EM and AF2 prediction. The obtained structure of D5 shows an AAA + helicase core, which is flanked by N- as well as C-terminal domains. The structure of D5 predicted by AF2 largely helped the construction of the model. The N-terminal domain, which has a 3.9 Å resolution, forms a well-defined tight ring, while the resolution decreases towards the C-terminus, which still allows the fit of the predicted structure. This structure validates AF2 calculations of a large number of structures of viral helicase associated with D5. Jin et al.72 solved the structure of interleukin (IL) −27 signal complex by using cryo-EM with the aid of AF2 prediction, through which they revealed a new mechanism for the assembly and activation of IL-27 receptor recognition complex. Skalidis et al.73 utilized cryo-EM and AF2 to characterize metabolon-embedded architectures of a 60S pre-ribosome, fatty acid synthase, and pyruvate/oxoglutarate dehydrogenase complex E2 cores. Though cryo-EM 3D reconstructions were resolved at resolution ranging from 3.84 to 4.52 Å by collecting less than 3,000 micrographs of a single cellular fraction, AF2 enabled polypeptide hydrogen bonding patterns discernible at this resolution range. These results proposed an integrated approach, powered by ML, which enables the cryo-EM characterization of native cell extracts.
Fowler and Williamson68 recently evaluated the accuracy of NMR structures and AF2 prediction. They used the program Accuracy of NMR Structures Using RCI and Rigidity (ANSURR), which calculates a protein structure’s local rigidity.74 They compared AF2 predicted structures and those determined by NMR and found that AF2 tends to be more accurate than NMR ensembles. They also found that the NMR ensembles are more accurate in some cases, which tend to be dynamic structures, where AF2 had low confidence. They finally suggested that AF2 could be utilized as the model for refining NMR-structure.
There are also some other similar studies in which AF2 is applied to help structural determination, and some of which combines AF2 and experimental methods to verify if the protein structure is solved correctly, for example, combining AF2 with X-ray crystallography,75,76 cryo-EM,77,78,79 NMR,80 and multiple methods.81,82,83,84,85
Besides structure determination, AF2 prediction can even be applied to the design of expression constructs. They enable researchers to better determine where the starting and ending points of a domain locate in the sequence, and avoid less ordered regions;86 Neglecting less ordered regions from protein sequences is often conducive to designing recombinant proteins for investigations pertaining to structures.
Drug discovery
Drug discovery is one of the major application areas that require protein structure information. Although the confidence level of prediction varies, the AF2 predicted structures still could considerably promote the structure-based drug discovery, especially against protein targets with limited or no structural information. At present, protein structures used in structure-based drug discovery mainly come from the RCSB Protein Data Bank (PDB). However, the number of protein structures in the PDB database are quite limited, which is far from meeting the current vigorous demand for drug discovery. The structures’ release of the entire protein universe is expected to accelerate existing and new drug discovery projects.
Zhang et al.87 recently used Glide,88 a molecular docking program, to benchmark the performance of virtual screening towards 28 common drug targets, each with a known experimental structure and an AF2 structure. The AF2 structures show comparable performance with experimental structures in terms of the enrichment factor, especially when flexible docking was used. The results clearly show that AF2 structures can completely replace the experimental structures in virtual screening.
Ren et al.89 applied AF2 in their end-to-end AI-powered drug discovery engines, which include a biocomputational platform named PandaOmics and a generative chemistry platform named Chemistry42.90 PandaOmics provides the targets of interest and Chemistry42 is responsible for generating molecules based on the AF2 predicted structures, and the selected molecules are then synthesized and tested in biological assays. Through this approach, they discovered a small molecule hit compound for CDK20 (Cyclin-dependent Kinase 20)91 with a Kd value of 8.9 ± 1.6 μM within 30 days from target selection and after only synthesizing 7 compounds. This compound was the first small molecule targeting CDK20 at that time, and this work is the first demonstration of AF2’s successful application in the early drug discovery process.
Weng et al.92 applied AF2 to predict the 3D structure of WSB1 (SOCS-box-containing WD-40 protein), a new potential anticancer target93,94,95 with 3D structural information not available. The predicted structure was then optimized by molecular dynamics simulations. The optimized 3D structure of WSB1 was taken as the receptor structure to perform molecular docking to screen for WSB1 inhibitors. Finally, they obtained a number of potential active compounds. Among these compounds, G490-0341 displayed the best stable structure and deserved further research and development.
Liang et al.96 identified JMJD897,98,99 as a novel oncogene correlated with immunosuppression and DNA repair by bioinformatics analysis. Then they used AF2 to predict the 3D structure of JMJD8 and performed virtual screening to retrieve JMJD8 inhibitors. Liu et al.100 proposed a multi-target drug discovery method and applied this method to drug discovery of therapeutic hypothermia.101 In this study, they first predicted the structure for all related protein targets by using AF2 and RoseTTAFold. After that, they applied molecular docking to estimate the interaction between proteins and drugs, and determined optimal single drugs or drug combinations. Considering the differences in the weights of different protein targets, the approach could refrain from inhibiting beneficial proteins effectively while inhibiting harmful proteins.
Except for the above examples, some researches also showed that the side chain quality modeled by AF2 is not good enough for drug discovery, and some recent studies also found that the docking test based on AF2-predicted structures showed weak enrichment performance.102,103
Other researches with AF2 applied in drug discovery include the literature.104,105,106
Protein design
Design of proteins means creating novel proteins with desired structures and functions. De novo protein design is a longstanding fundamental goal of synthetic biology.107,108,109 It is a complex and challenging task, which is mainly hindered by the difficulty in reliable prediction of protein 3D structures from amino acid sequences. AF2 as well as other machine learning algorithms (such as RoseTTAFold and recent language models49,50) likely removes this obstacle. It is no exaggeration to say that with AF2 prediction, we will step into a new era of protein design. Some typical protein design examples by using AF2 are given as follows.
Jendrusch et al110 developed a computational framework for de novo protein design that embeds AF2 as an oracle within an optimizable design process. This is an adaptable framework for protein design through sequence optimization utilizing evolutionary algorithms. It extends previous studies towards protein design by leveraging structure predictors.111,112 The integrity of the structures predicted is validated and confirmed by standard ab initio folding, protein structure analysis methods, and rigorous all-atom molecular dynamics simulations. They also showed a potential application of their method in designing de novo protein monomers, dimers and oligomers, as well as protein binders for target proteins and proteins which change conformation upon complex formation.
Goverde et al.113 designed a pipeline for de novo protein design based on AF2. In the beginning, they just inverted the AF2 model, utilizing a loss function and the prediction weight set to bias the generated sequences for the objective of adopting a target fold. However, as observed in the protein surface’s hydrophilic versus hydrophobic patterning, the approach does not seem to fully capture basic principles of de novo protein design. Then they made modifications to their pipeline system with minimal post-design intervention, and conducted in vitro validation, which demonstrated that some designs were folded and stable in solution in the condition of high melting temperatures. Overall, the revised pipeline generated viable sequences as assessed experimental characterization, showing the possibility of contributing to solving outstanding challenges in the field of de novo protein design.
Other interesting studies with AF2 assisting protein design include the literature.114,115,116,117 It is also necessary to mention that, relative to AF2, RoseTTAFold has more applications in protein design, largely due to Baker’s groundbreaking work.118,119,120
Target prediction
Target prediction, including on-target and off-target identifications, are important not only for understanding physiological and pathological processes, but also for identifying novel drug targets and evaluating selectivity of drugs. Experimental approaches to target identification, such as various activity-based protein profiling (ABPP)-based methods,121,122,123,124 are often expensive and time consuming. Computer-aided target prediction may help narrow the scope of target identification, which is often based on protein-ligand docking, usually called inverse docking. Previously, the inverse docking faces a challenge of lacking 3D structures of all possible protein targets. The AF2 structures provide an unprecedented opportunity to develop feasible target prediction methods.
Wang et al.125 utilized the AF2 structures to construct the first pocket library for all the proteins in the human proteome, called the CavitySpace database. CavitySpace can be applied to identify novel targets for known drugs in drug repurposing or side effect researches. This database can be easily used to the target prediction by inverse docking. The building workflow of database is as follows: they collected 23,391 human proteins from AlphaFold protein structure database and 6956 human reference proteins from PDB. Then, they applied CAVITY, a tool developed by the same research group to detect all the possible cavities on protein surfaces,126 to identify all the potential cavities on protein surfaces. The CavitySpace database is freely available at http://www.pkumdl.cn:8000/cavityspace/.
There are also other related studies applied AF2 for target prediction.127,128,129
Protein function prediction
Currently, there are still many proteins whose functions are not known or poorly understood. Since 3D structures of proteins completely determine their functionality, this characteristic can be utilized to establish data-driven prediction models of protein function. Nevertheless, the insufficient number of available protein structures severely limits the performance of these models. The structures predicted by AF2 have provided a promising solution towards this problem, and are expected to improve the performance of these models via increasing the amount of training samples.
Ma et al.130 recently conducted a comprehensive study to investigate whether AF2-predicted structures could enhance the protein function prediction performance. In this study, they proposed a state-of-the-art structure-based protein function prediction approach and constructed a new benchmark data set. After that, they evaluated whether the performance of the protein function prediction model could be improved by putting additional protein structures predicted by AF2 into the training data set. They further compared the performance differences between two models separately trained with structures predicted by AF2 only and with real protein structures only. Their results demonstrated that protein function prediction models based on structures could benefit from virtual training data composed of structures predicted by AF2. Even, the model trained only using the structures predicted by AF2 achieved comparable performances to the model based on real protein structures, which are solved through experiments. This indicates that the structures predicted by AF2 were almost equally effective in protein function prediction.
Hu et al.131 explored the utility of the Protein Language Models (PLMs) module in AF2, Evoformer, in protein function prediction, and particularly compared the performance of evolution-based & evolution-free protein language models as protein function predictors. They showed that evolution-based PLMs performed better than evolution-free models only in the structure prediction tasks, but in general, were worse than evolution-free models in most function prediction tasks. Consistent with structure prediction, evolution-based PLMs are also sensitive to the amount of MSAs when predicting protein function.
Interpretable and compact structural feature representations are important for accurate prediction of protein properties and function. In a recent study, Rappoport and Jinich132 constructed and evaluated 3D feature representations of protein structures using space-filling curves, in which AF2 predicted protein structures were used. In this study, two enzyme substrate predictions were used as case studies: the S-adenosylmethionine dependent methyltransferases (SAM-MTases) and the short-chain dehydrogenase/reductases (SDRs). As their results demonstrate, enzymatic function could be predicted from feature representations on the basis of the 3D structures of SAM-MTAses and SDRs with good accuracy.
By searching proteins that contain Zα domain (experimentally validated Z-DNA/Z-RNA133 binding protein domain) from AF2 predicted structure database, Bartas et al.134 identified 185 proteins with a putative Zα domain, which may bind to Z-DNA/Z-RNA and play an important role in a variety of cellular processes.
There are also other interesting researches related to protein function prediction involving AF2.135,136
Protein–protein interaction
Protein-protein interaction (PPI) refers to the process in which two or more protein molecules form a protein complex through non-covalent bonds.137,138 A majority of proteins need to recruit other proteins through PPI to form protein complexes to perform their functions. Understanding the structure of interacting proteins is a fundamental step towards revealing the protein function and mechanism. However, there is lack of computational tools that can produce accurate structures of protein complexes. The emergence of AF2 can be greatly conducive to this area.
Evans et al.139 extended AF2 to the prediction of multiple-chain complex, and the system was named as AlphaFold-Multimer. On a benchmark data set of 17 heterodimer proteins without templates, they achieved at least medium accuracy on 14 targets and high accuracy on 6 targets. They also predicted structures for a large data set containing more than 4,000 recent protein complexes, from which they scored all the non-redundant interfaces with low template identity from these protein complexes. For heteromeric interfaces, they successfully predicted the interface in 67% of the cases, and 23% of the cases were predicted with high accuracy. For homomeric interfaces they effectively predicted the interface in 69% of cases, and produced high accuracy predictions in 34% of cases. All these results demonstrated superior performance compared to existing approaches. The AlphaFold2-multimers has now been used to predict protein-protein complex structures. For example, Gómez-Marín et al.140 applied AlphaFold-multimer for the prediction of PHF14-HMG20A complex models. Ivanov et al.141 also applied AlphaFold-multimer to predict the homodimers structure of CYP102A1.
Recently, Bryant et al.142 applied AF2 to predict heterodimeric protein complexes. In this work, they explored the docking effect by using the AF2 pipeline combined with different input MSAs, which is for studying the relationship between the output model quality and these inputs. Through scoring multiple PPI models with a predicted DockQ score (pDockQ), they could distinguish from incorrect models with high confidence acceptable (pDockQ > = 0.23). They concluded that AF2-based docking outperformed another docking method.143
Yin et al.144 examined the performance of AF2 in predicting structures of protein complexes from amino acid sequence. They used 152 diverse heterodimeric protein complexes to form a benchmark test data set. In this test, 43% of cases that had near-native models were produced as top ranking prediction results by AF2, substantially outperforming the performance of unbound protein–protein docking method (9%). To examine the effect of AlphaFold_Multimer in predicting antibody–antigen interaction, the authors used a set of antibody-antigen structures, which were released recently. The testing results confirmed a low success rate for the modeling of antibody–antigen complexes. They further observed that, via the algorithm, T cell receptor–antigen complexes are similarly not accurately modeled. These findings demonstrate that AF2 faces challenges in handling the adaptive immune recognition. Gao et al.145 developed an AF2-based system called AF2Complex, which can predict direct physical interactions in multimeric proteins. Contrary to normal approaches, paired MSAs are not necessary for AF2Complex. It improves significantly over AlpahFold-Multimer and reaches higher accuracy compared to some complex protein-protein docking approaches. Moreover, the authors introduced metrics used for direct protein-protein interactions prediction between arbitrary pairs of proteins and validate AF2Complex on the E. coli proteome as well as some challenging benchmark sets.
In addition to PPI, AF2 can also been used in the prediction of peptide-protein interactions. For example, Tsaban et al.146 suggested an AF2-based strategy to model peptide-protein complex which did not require MSA information for the peptide partner. In this way, binding-induced conformational changes of the receptor could be handled. The outcomes demonstrate that AF2 could be expected to provide structural insight into a broad range of peptide-protein complexes.
For some similar investigations related to protein-protein interaction or protein-peptide interaction by using AF2, readers can see the literature.147,148,149,150,151,152,153
Biological mechanism of action
Exploring biological mechanism of action is often complicated and remains a challenge. Studies of biological mechanism of action include many aspects, such as drug-target interaction mode, mechanism of biological enzyme catalysis, and so on.
In silico molecular docking methods have been broadly applied to the prediction of drug-target interaction. Nevertheless, this kind of methods strongly rely on existing protein structures. AF2 provides alternative approach to retrieval of accurate protein structures. Wong et al.102 combined molecular docking simulations with AF2 for protein-ligand interactions prediction. They successfully predicted the interactions between 296 proteins spanning the essential proteome of Escherichia coli, and 218 active antibacterial compounds and 100 inactive compounds, respectively. They measured the enzymatic activity of 12 essential proteins which were treated with each antibacterial compound to benchmark the performance of the model. This research suggests that advanced approaches in modeling protein–ligand interactions, especially utilizing methods based on machine learning, are needed to better leverage AF2 for mechanism of action studies as well as drug discovery.
Lactate oxidation with NAD+ as electron acceptor154 is a highly endergonic reaction. Some anaerobic bacteria conquer the energetic barrier through electron bifurcation/confurcation (FBEB/FBEC) based on flavin utilizing a lactate dehydrogenase (Ldh) combining with EtfA and EtfB, which are the electron-transferring proteins. However, the mechanism of action is poor understood. In a recent study, Kayastha et al.155 utilized AF2 calculations and obtained a plausible new B (bifurcation-connected) state, which allows electron to transmit between the shuttle FADs and EtfAB base. Based on the findings, they put forward an integrated catalytic mechanism of the FBEC process.
Post-transcriptional RNA editing regulates the expression of gene in a condition-dependent manner, which mechanism remains unclear. In a recent study, Kimura et al.156 characterized the C-to-Ψ editing mechanisms. They showed that TrcP mediates the stepwise editing of C-to-U followed by the conversion of U to Ψ. The structure modeling based on AF2 revealed a distinct long helical domain within TrcP which possibly binds and orients the substrate tRNA during both reactions, The findings suggest that TrcP mediated C-to-Ψ editing depends on a substrate channeling mechanism. These discoveries offer mechanistic views into an RNA editing process which could possibly stimulate environmental adaptation.
Liang et al.157 explored the substrate determinants and recognition mechanism of separase, which is a giant cysteine protease. In budding yeast, they identified a conserved motif downstream of the cleavage site. Using AF2 and molecular dynamics simulations, they discovered that in a conserved cleft near the binding groove of separase’s inhibitor securin, the motif is recognized by separase. The binding is mutually exclusive and requires separase’s conformation changes. Their research could let scientists get a deeper understanding of mechanism of substrate recognition and activation of separase.
Lorenz et al.158 applied AF2 to predict the structure of selected KRAB159 domains. They discovered an evolutionary conserved L-shaped body of two α-helices in all the domains of KRAB. It is changed into a typical spatial arrangement especially for mKRAB-AB after the replacement of amino acid and together with a third helix provided by mKRAB-B. This provides basic insights of how KRAB form complex with TRIM28. McMullen et al.160 found from yeast that EKP-GCSF and GCSF exhibits similar binding to its receptor GCSF-R. Similarly, to study the structural effects of EKP161,162 on GCSF, they applied computational modeling using AF2 in conjunction with molecular dynamics simulations. Computational modeling shows that EKP does not change the structural behavior of GCSF, which demonstrates that EKP does not hinder receptor binding. Furthermore, the initial conformation of EKP-GCSF from AF2 shows that the EKP which is around GCSF might provide evidence of the thermal-protectivity of EKP on GCSF.
There are also other studies pertaining to biological mechanism of action.163,164,165,166,167,168,169,170,171,172,173,174,175,176
Other applications
In addition to the application areas mentioned above, AF2 prediction could also be applied to some other fields, such as protein evolution,177,178,179,180 rare disease treatment studies,181 effects of mutation on treatment,182,183,184,185,186,187 vaccine design,188,189,190 and so on. For example, Tang et al.180 investigated the relationship between organism evolution and protein evolution based on the structures of proteomes from 48 organisms predicted by AF2. They found some interesting phenomena, including: (1) constituent proteins of organisms with higher complexity would have larger gyration radii, higher coil fractions as well as slower vibrations, and (2) higher degree of functional specialization of proteins is associated with higher degree of organismal complexity. This research brings new views about how the proteins’ functionality diversity increases, and how the dimensionality of the manifold of protein dynamics decreases in the process of evolution. Sebastiano et al.181 found that protein structures predicted by AF2 have a potential to assist rare disease treatment studies. In this investigation, the authors focused on Alsin, a protein responsible for rare motor neuron diseases. With the AF2 predicted protein structures, they evaluated the flexibility profile of Alsin and its mutants, and models of dimeric/tetrameric Alsin responsible for its physiological action. They concluded that efforts of drug discovery targeting Alsin-involving diseases should be pursued. Yang et al.184 applied AF2 to predict the S, N, and M proteins’ structures of the Omicron variant of SARS-CoV-2. They analyzed how the S protein and its parts, S1 RBD and NTD, have been affected by the mutations in detail, and also how the current SARS-CoV-2 vaccines and treatments would be affected by these mutations. Zeng et al.188 utilized AF2 to design a hemagglutinin stem vaccine “B60-Stem-8070”. This vaccine showed better performance compared with the original hemagglutinin stem antigen.
Limitations of current AF2 prediction
The invention of AF2 is a game-changer event in structural biology. It has reformed the field of protein structure prediction by utilizing sequence information to model protein folds quickly with atomic-level accuracy. However, current AF2 was trained on protein structures from the Protein Data Bank in which X-ray crystallographic structures dominate. Therefore, it is best considered as a predictor of the structured state under experimental conditions where a protein is likely to crystallize, other than a predictor of the lowest free-energy state under physiological conditions. This together with some inherent limitations in methods and techniques limits applications of AF2 predictions in many aspects, which are summarized as follows.
The protein dynamics
Protein dynamics is a very important research area.191,192,193,194,195,196,197 The protein structure predicted by AF2 is a static state. However, proteins are very dynamic with multiple states. Many important physiological and pathological proteins (such as ion channel proteins) have very subtle conformational changes under different active states, and will also show ever-changing spatial configurations due to their combination with various other proteins inside and outside the cell. At this point, AF2 often gives a single optimal solution, which is difficult to cover the conformational diversity of proteins. However, this does not mean that it is not possible to understand protein dynamics with AF2. According to several recent studies,198,199,200,201 AF2 can still be used for some analysis of protein dynamics. For example, Del Alamo et al.199 recently presented a method to drive AF2 for sampling alternative conformations of topologically diverse transporters as well as the G-protein-coupled receptors that do not exist in the training dat set of AF2. Nevertheless, the exploration of the conformational space is in part a by-product of low sequence information which is provided for inference.
There are also other studies pointing out that AF2’s weak performance in identifying conformational ambiguity.202 Additionally, AF2 could hardly be used for the structure prediction of a protein with multiple domains, such as a transmembrane receptor with a large extracellular domain.203 There is still a demand of new deep learning methods to be designed for the prediction of ensembles of biophysically correlated states.
Structures for disordered regions of proteins
The AF2 database consists of highly accurate predictions for the folded part of a large number of proteins. Nevertheless, AF2 does not well in predicting structures of proteins, in cases fewer sequences are available for alignment, and regions that are natively unfolded or disordered regions, for example, loops. The loop structures are relatively stable in crystals, but are very flexible in solutions. Although many approaches have been tried,204,205,206,207 it is difficult for existing methods to predict the morphology, dynamics and interactions of disordered regions of proteins in solutions.
Structures of proteins in complex with small molecules or other proteins
It has been well known that small molecule ligands or proteins may induce a protein to undergo conformational changes. The most representative example is allosteric modulators,208 which refers to small molecules or peptides that bind to a site of an enzyme protein different from its endogenous ligand binding site to cause conformational changes, thereby changing the activity of the enzyme. Besides allosteric modulators, plenty of orthosteric ligands, which bind to the identical site as the endogenous ligand, can also induce conformational changes. Nevertheless, AF2 is not designed to determine how proteins change their shape in the presence of other interacting ligands or proteins.
Structures of proteins with point mutations
Point mutations are frequently encountered in proteins, particularly pathological state. Understanding the effect of missense mutations on protein structure may help unveil their biological or pathological mechanism. Even though AF2 could predict wild-type (WT) structures, it likely performs poorly in predicting the effect of missense mutations on the proteins’ 3D structures. Although there are researches showing that AF2 could predict the phenotypic effect of missense mutations,209,210 it was also observed that the performance of missense mutations prediction is not good in other studies,211 and there were only weak or no correlations between the output metrics of AF2 and changes in protein stability or functionality.212
Structures of proteins with post-translational modifications
Post-translational modifications,213,214,215 such as phosphorylation,216 methylation,217 acetylation,218 and glycosylation,219 are common in proteins. These post-translational modifications may lead to conformational changes in protein structures.220 For example, phosphorylation of inactive kinases in their active loop often results in a large conformational change, and eventually activate the kinases. However, AF2 can predict protein structures only based on their amino acid sequence, and post-translational modifications of residues cannot be recognized. Therefore, the conformational changes due to post-translational modifications cannot be predicted with current AF2.
Prediction of orphan proteins and artificially designed proteins
In addition to the above-mentioned limitations, AF2 and other computational systems that use DL and the information of co-evolutionary relationships encoded in MSAs also face challenges in prediction of orphan proteins and artificially designed proteins because an MSA cannot be generated. Recently Chowdhury et al.51 developed an end-to-end deep neural network model, namely differentiable recurrent geometric network (RGN) model, in which a protein language model (AminoBERT) based on the Bidirectional Encoder Representations from Transformers (BERT)221 was used to learn latent structural information from unaligned proteins. Language models were firstly introduced to extract semantic information from words. The RGN model showed better performance on orphan proteins than AF2, while significantly reducing the computing time by up to 106-folds. These results demonstrate the theoretical and practical strengths of protein language models in structure prediction compared with MSAs.
Limitations in methods and techniques
We finally have to mention that AF2 itself has some limitations in methods and techniques. For example, (i) deep learning models have low interpretability currently; (ii) the AF2 structural prediction is based on the data of MSA, i.e., a large number of evolutionarily related sequences is needed for the structure predictions, which might cause side effects such as comparably slower prediction speed. As a comparison, language models (such as ESMfold49 and RGN50) enable end-to-end protein structure prediction directly from amino acid sequences with high speed and accuracy.
Concluding remarks
The excellent performance of AF2 in predicting protein structure together with the release of structures of more than 200 million proteins predicted by AF2 is reshaping structural biology. AF2 will certainly have a significant impact on researches that need protein structure information, and could be applied in many fields such as drug discovery, protein design, target prediction, protein function prediction, PPI, biological mechanism of action, and others, in addition to experimental structural biology. Despite just a very short time since AF2 was developed, we have already witnessed a number of successful applications. We believe that, as time goes on, more applications or new application fields will be developed, for example, design of protein machines with complex or specific functions, design of new organisms, and disease diagnosis. Even so, AF2 prediction is not a panacea and there are many issues still needing to be solved, including protein dynamics, structures of disordered regions of proteins, structures of mutants, structures of protein-ligand complexes, structures of proteins with post-translational modifications, and so on. With the further development of AI algorithm, ever-increasing data, and computing power, it is expected that more surprises will surely come to us in future.
Finally, it is necessary to mention that, during the revision of this article, the new CASP competition, CASP15, is over. Different from previous CASPs, which took protein structure prediction as the main track, CASP 15 also paid attention to predicting protein complex and RNA structures. This is consistent with the CASP’s style or philosophy that keeping pace with the times, which means that protein complex structures and RNA structures might be the new focuses in the “post AlphaFold era”. Another noteworthy point is that DeepMind did not take part in CASP15, which reasons are unclear. Nevertheless, all teams that have achieved better results have more or less used AF2 algorithms or AF2 predicted structures, which implies that AF2 invisibly won CASP15 again, further highlighting the great influence of AF2 in structural biology. Overall, we look forward to new breakthroughs of AI in structural biology, and more application achievements by using AF2 in the future.
References
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Kryshtafovych, A. et al. Critical assessment of methods of protein structure prediction (CASP) - Round XIV. Proteins 89, 1607–1617 (2021).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
Dill, K. A. & MacCallum, J. L. The protein-folding problem, 50 years on. Science 338, 1042–1046 (2012).
Wüthrich, K. Protein structure determination in solution by NMR spectroscopy. J. Biol. Chem. 265, 22059–22062 (1990).
Shi, Y. A glimpse of structural biology through X-ray crystallography. Cell 159, 995–1014 (2014).
Earl, L. A., Falconieri, V. & Milne, J. L. Subramaniam, S. Cryo-EM: beyond the microscope. Curr. Opin. Struct. Biol. 46, 71–78 (2017).
Anfinsen, C. B. et al. The kinetics of formation of native ribonuclease during oxidation of the reduced polypeptide chain. Proc. Natl Acad. Sci. USA 47, 1309–1314 (1961).
Cozzetto, D., Di, Matteo, A. & Tramontano, A. Ten years of predictions… and counting. FEBS J. 272, 881–882 (2005).
Moult, J. A decade of CASP: progress, bottlenecks and prognosis in protein structure prediction. Curr. Opin. Struct. Biol. 15, 285–289 (2005).
Lupas, A. N. et al. The breakthrough in protein structure prediction. Biochem. J. 478, 1885–1890 (2021).
Torrisi, M., Pollastri, G. & Le, Q. Deep learning methods in protein structure prediction. Comput. Struct. Biotechnol. J. 18, 1301–1310 (2020).
Kuhlman, B. & Bradley, P. Advances in protein structure prediction and design. Nat. Rev. Mol. Cell Biol. 20, 681–697 (2019).
AlQuraishi, M. Machine learning in protein structure prediction. Curr. Opin. Chem. Biol. 65, 1–8 (2021).
Jisna, V. A. & Jayaraj, P. B. Protein structure prediction: conventional and deep Learning perspectives. Protein J. 40, 522–544 (2021).
Pearce, R. & Zhang, Y. Toward the solution of the protein structure prediction problem. J. Biol. Chem. 297, 100870 (2021).
Gao, W. et al. Deep learning in protein structural modeling and design. Patterns 1, 100142 (2020).
Al-Lazikani, B. et al. Protein structure prediction. Curr. Opin. Chem. Biol. 5, 51–56 (2001).
Xiang, Z. Advances in homology protein structure modeling. Curr. Protein Pept. Sci. 7, 217–227 (2006).
Tramontano, A. et al. Assessment of homology‐based predictions in CASP5. Proteins 53, 352–368 (2003).
Bordoli, L. et al. Protein structure homology modeling using SWISS-MODEL workspace. Nat. Protoc. 4, 1–13 (2009).
Cardozo, T., Totrov, M. & Abagyan, R. Homology modeling by the ICM method. Proteins 23, 403–414 (1995).
Bower, M. J., Cohen, F. E. & Dunbrack, R. L. Jr. Prediction of protein side-chain rotamers from a backbone-dependent rotamer library: a new homology modeling tool. J. Mol. Biol. 267, 1268–1282 (1997).
Aszódi, A. & Taylor, W. R. Homology modelling by distance geometry. Fold. Des. 1, 325–334 (1996).
Muhammed, M. T. & Aki-Yalcin, E. Homology modeling in drug discovery: overview, current applications, and future perspectives. Chem. Biol. Drug Des. 93, 12–20 (2019).
Abriata, L. A. et al. State-of-the-art web services for de novo protein structure prediction. Brief. Bioinform. 22, bbaa139 (2021).
Bradley, P., Misura, K. M. S. & Baker, D. Toward high-resolution de novo structure prediction for small proteins. Science 309, 1868–1871 (2005).
Wu, R. et al. High-resolution de novo structure prediction from primary sequence. Preprint at https://www.biorxiv.org/content/10.1101/2022.07.21.500999v1 (2022).
Liu, J. et al. CGLFold: a contact-assisted de novo protein structure prediction using global exploration and loop perturbation sampling algorithm. Bioinformatics 36, 2443–2450 (2020).
Bhattacharya, D., Cao, R. & Cheng, J. UniCon3D: de novo protein structure prediction using united-residue conformational search via stepwise, probabilistic sampling. Bioinformatics 32, 2791–2799 (2016).
Lee, J. et al. De novo protein structure prediction by dynamic fragment assembly and conformational space annealing. Proteins 79, 2403–2417 (2011).
Zhao, K. L. et al. MMpred: a distance-assisted multimodal conformation sampling for de novo protein structure prediction. Bioinformatics 37, 4350–4356 (2021).
Peng, C. X., Zhou, X. G. & Zhang, G. J. De novo protein structure prediction by coupling contact with distance profile. IEEE/ACM Trans. Comput. Biol. Bioinform. 19, 395–406 (2020).
Levinthal, C. How to fold graciously. Mössbaun Spectr. Biol. Sys. Proc. 67, 22–24 (1969).
Salehinejad, H. et al. Recent advances in recurrent neural networks. Preprint at https://arxiv.org/abs/1801.01078 (2017).
Gu, J. et al. Recent advances in convolutional neural networks. Pattern Recogn. 77, 354–377 (2018).
Goodfellow, I. et al. Generative adversarial networks. Commun. ACM 63, 139–144 (2020).
Vaswani, A. et al. Attention is all you need. Proc. Adv. Neural Inf. Process. Syst. 30, 5998–6008 (2017).
Voulodimos, A. et al. Deep learning for computer vision: a brief review. Comput. Intell. Neurosci. 2018, 7068349 (2018).
Otter, D. W. et al. A survey of the usages of deep learning for natural language processing. IEEE Trans. Neural Netw. Learn. Syst. 32, 604–624 (2020).
Grigorescu, S. et al. A survey of deep learning techniques for autonomous driving. J. Field Rob. 37, 362–386 (2020).
Purwins, H. et al. Deep learning for audio signal processing. IEEE J. Sel. Top. Signal Process. 13, 206–219 (2019).
Khan, S. & Yairi, T. A review on the application of deep learning in system health management. Mech. Syst. Signal Process. 107, 241–265 (2018).
Zhou, L. et al. Application of deep learning in food: a review. Compr. Rev. Food Sci. Food Saf. 18, 1793–1811 (2019).
Christin, S., Hervet, É. & Lecomte, N. Applications for deep learning in ecology. Methods Ecol. Evol. 10, 1632–1644 (2019).
Mater, A. C. & Coote, M. L. Deep learning in chemistry. J. Chem. Inf. Model. 59, 2545–2559 (2019).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Lin, Z. et al. Evolutionary-scale prediction of atomic level protein structure with a language model. Preprint at https://www.biorxiv.org/content/10.1101/2022.07.20.500902v3 (2022).
Chowdhury, R. et al. Single-sequence protein structure prediction using a language model and deep learning. Nat. Biotechnol. 40, 1617–1623 (2022).
Kryshtafovych, A. et al. Critical assessment of methods of protein structure prediction (CASP)—Round XIII. Proteins 87, 1011–1020 (2019).
Kryshtafovych, A., Schwede, T., Topf, M., Fidelis, K. & Moult, J. Critical assessment of methods of protein structure prediction (CASP)-Round XIV. Proteins 89, 1607–1617 (2021).
Zemla, A. LGA: A method for finding 3D similarities in protein structures. Nucleic Acids Res. 31, 3370–3374 (2003).
De, Juan, D., Pazos, F. & Valencia, A. Emerging methods in protein co-evolution. Nat. Rev. Genet. 14, 249–261 (2013).
Suzek, B. E., Wang, Y., Huang, H., McGarvey, P. B. & Wu, C. H. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015).
Mirdita, M. et al. Uniclust databases of clustered and deeply annotated protein sequences and alignments. Nucleic Acids Res. 45, D170–D176 (2017).
Mitchell, A. L. et al. MGnify: the microbiome analysis resource in 2020. Nucleic Acids Res. 48, D570–D578 (2020).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinform. 20, 473 (2019).
Johnson, L. S., Eddy, S. R. & Portugaly, E. Hidden markov model speed heuristic and iterative HMM search procedure. BMC Bioinform. 11, 1–8 (2010).
Remmert, M., Biegert, A., Hauser, A. & Söding, J. Hhblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 9, 173–175 (2012).
Steinegger, M. et al. HH-suite3 for fast remote homology detection and deep protein annotation. BMC Bioinform. 20, 1–15 (2019).
Eastman, P. et al. OpenMM 7: rapid development of high performance algorithms for molecular dynamics. PLoS Comput. Biol. 13, e1005659 (2017).
Hornak, V. et al. Comparison of multiple Amber force fields and development of improved protein backbone parameters. Proteins 65, 712–725 (2006).
Xie, Q., Luong, M.-T., Hovy, E. & Le, Q. V. Self-training with noisy student improves imagenet classification. In Proc. IEEE Conf. Comput. Vis. Pattern Recognit. 10687–10698 (2020).
Akdel, M. et al. A structural biology community assessment of AlphaFold2 applications. Nat. Struct. Mol. Biol. 29, 1056–1067 (2022).
Cramer, P. AlphaFold2 and the future of structural biology. Nat. Struct. Mol. Biol. 28, 704–705 (2021).
Hendrickson, W. A., Horton, J. R. & LeMaster, D. M. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. Embo. J. 9, 1665–1672 (1990).
Fowler, N. J. & Williamson, M. P. The accuracy of protein structures in solution determined by AlphaFold and NMR. Structure 30, 925–933 (2022).
Tejero, R., Huang, Y. J., Ramelot, T. A. & Montelione, G. T. AlphaFold models of small proteins rival the accuracy of solution NMR structures. Front. Mol. Biosci. 9, 877000 (2022).
Hu, L. et al. Novel fold of rotavirus glycan-binding domain predicted by AlphaFold2 and determined by X-ray crystallography. Commun. Biol. 5, 419 (2022).
Hutin, S. et al. The Vaccinia virus DNA helicase structure from combined single-particle cryo-electron microscopy and AlphaFold2 prediction. Viruses 14, 2206 (2022).
Jin, Y. et al. Structural insights into the assembly and activation of the IL-27 signaling complex. Embo. Rep. 23, e55450 (2022).
Skalidis, I. et al. Cryo-EM and artificial intelligence visualize endogenous protein community members. Structure 30, 575–589.e6 (2022).
Jacobs, D. J. et al. Protein flexibility predictions using graph theory. Proteins 44, 150–165 (2001).
Nagaratnam, N. et al. Structural and biophysical properties of FopA, a major outer membrane protein of Francisella tularensis. PLoS One 17, e0267370 (2022).
Paul, B. et al. Structural predictions of the SNX-RGS proteins suggest they belong to a new class of lipid transfer proteins. Front. Cell Dev. Biol. 10, 826688 (2022).
Liu, H. et al. Cryo-EM structures of human hepatitis B and woodchuck hepatitis virus small spherical subviral particles. Sci. Adv. 8, eabo4184 (2022).
Tai, L. et al. 8 Å structure of the outer rings of the Xenopus laevis nuclear pore complex obtained by cryo-EM and AI. Protein Cell 13, 760–777 (2022).
Chang, L. et al. DeepTracer-ID: De novo protein identification from cryo-EM maps. Biophys. J. 121, 2840–2848 (2022).
Arantes, P. R. et al. Assessing structure and dynamics of AlphaFold2 prediction of GeoCas9. Biophys. J. 121, 45 (2022).
Stsiapanava, A. et al. Structure of the decoy module of human glycoprotein 2 and uromodulin and its interaction with bacterial adhesin FimH. Nat. Struct. Mol. Biol. 29, 190–193 (2022).
Overduin, M. et al. Transmembrane membrane readers form a novel class of proteins that include peripheral phosphoinositide recognition domains and viral spikes. Membranes (Basel) 12, 1161 (2022).
Burnim, A. A. et al. Comprehensive phylogenetic analysis of the ribonucleotide reductase family reveals an ancestral clade. Elife 11, e79790 (2022).
Allison, T. M. et al. Complementing machine learning-based structure predictions with native mass spectrometry. Protein Sci. 31, e4333 (2022).
Murphy, R. D. et al. The Toxoplasma glucan phosphatase TgLaforin utilizes a distinct functional mechanism that can be exploited by therapeutic inhibitors. J. Biol. Chem. 298, 102089 (2022).
Edich, M. et al. The impact of AlphaFold2 on experimental structure solution. Faraday Discuss 240, 184–195 (2022).
Zhang, Y. et al. Benchmarking Refined and Unrefined AlphaFold2 Structures for Hit Discovery. Preprint at https://chemrxiv.org/engage/chemrxiv/article-details/62b41f0c0bbbc117477285a4 (2022).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Ren, F. et al. AlphaFold accelerates artificial intelligence powered drug discovery: efficient discovery of a novel CDK20 small molecule inhibitor. Chem. Sci. 14, 1443–1452 (2023).
Ivanenkov, Y. A. et al. Chemistry42: An AI-Driven Platform for Molecular Design and Optimization. J. Chem. Inf. Model. 63, 695–701 (2023).
Mok, M. T. et al. CCRK is a novel signalling hub exploitable in cancer immunotherapy. Pharmacol. Ther. 186, 138–151 (2018).
Weng, Y. et al. Identification of potential WSB1 inhibitors by AlphaFold modeling, virtual screening, and molecular dynamics simulation studies. Evid.-Based Complement. Alternat. Med. 2022, 4629392 (2022).
Cao, J. et al. Hypoxia-induced WSB1 promotes the metastatic potential of osteosarcoma cells. Cancer Res. 75, 4839–4851 (2015).
Zhang, Y. et al. WD repeat and SOCS box containing protein 2 in the proliferation, cycle progression, and migration of melanoma cells. Biomed. Pharmacother. 116, 108974 (2019).
Che, J. et al. Discovery of 5, 6-Bis (4-methoxy-3-methylphenyl) 21yridine-2-amine as a WSB1 degrader to inhibit cancer cell metastasis. J. Med. Chem. 64, 8621–8643 (2021).
Liang, X. et al. JMJD8 is an M2 macrophage biomarker, and it associates with DNA damage repair to facilitate stemness maintenance, chemoresistance, and immunosuppression in Pan-cancer. Front. Immunol. 13, 875786 (2022).
Su, Y. & Wang, J. JmjC domain-containing protein 8 (JMJD8) represses Ku70/Ku80 expression via attenuating AKT/NF-κB/COX-2 signaling. Biochim. Biophys. Acta - Mol. Cell Res. 1866, 118541 (2019).
Zhang, B. et al. JMJD8 promotes malignant progression of lung cancer by maintaining EGFR stability and EGFR/PI3K/AKT pathway activation. J. Cancer 12, 976 (2021).
Wang, L. et al. MiR-873-5p suppresses cell proliferation and epithelial–mesenchymal transition via directly targeting Jumonji domain-containing protein 8 through the NF-κB pathway in colorectal cancer. J. Cell Commun. Signal. 13, 549–560 (2019).
Liu, F. et al. A chronotherapeutics-applicable multi-target therapeutics based on AI: Example of therapeutic hypothermia. Brief. Bioinform. 23, bbac365 (2022).
Song, S. S. & Lyden, P. D. Overview of therapeutic hypothermia. Curr. Treat. Options Neurol. 14, 541–548 (2012).
Wong, F. et al. Benchmarking AlphaFold-enabled molecular docking predictions for antibiotic discovery. Mol. Syst. Biol. 18, e11081 (2022).
Xu, G. et al. OPUS-Rota4: a gradient-based protein side-chain modeling framework assisted by deep learning-based predictors. Brief. Bioinform. 23, bbab529 (2022).
Yang, Q. et al. Structural comparison and drug screening of spike Proteins of Ten SARS-CoV-2 Variants. Research 2022, 9781758 (2022).
Park, H. M. et al. Rethinking Protein Drug Design with Highly Accurate Structure Prediction of Anti-CRISPR Proteins. Pharm. (Basel) 15, 310 (2022).
Yang, Q. et al. Highly accurate protein structure prediction and drug screen of monkeypox virus proteome. J. Infect. 86, 66–117 (2023).
Korendovych, I. V. & DeGrado, W. F. De novo protein design, a retrospective. Q. Rev. Biophys. 53, e3 (2020).
Huang, P. S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320–327 (2016).
Pan, X. & Kortemme, T. Recent advances in de novo protein design: Principles, methods, and applications. J. Biol. Chem. 296, 100558 (2021).
Jendrusch, M. et al. AlphaDesign: A de novo protein design framework based on AlphaFold. Preprint at https://www.biorxiv.org/content/10.1101/2021.10.11.463937v1 (2021).
Anishchenko, I. et al. De novo protein design by deep network hallucination. Nature 600, 547–552 (2021).
Norn, C. et al. Protein sequence design by conformational landscape optimization. Proc. Natl Acad. Sci. USA 118, e2017228118 (2021).
Goverde, C. et al. De novo protein design by inversion of the AlphaFold structure prediction network. Preprint at https://www.biorxiv.org/content/10.1101/2022.12.13.520346v1 (2022).
Arenas, N. E. et al. Design of a specific peptide against phenolic glycolipid-1 from Mycobacterium leprae and its implications in leprosy bacilli entry. Mem. Inst. Oswaldo. Cruz. 117, e220025 (2022).
Peñas-Utrilla, D. & Marcos, E. Identifying well-folded de novo proteins in the new era of accurate structure prediction. Front. Mol. Biosci. 9, 991380 (2022).
Listov, D. et al. Assessing and enhancing foldability in designed proteins. Protein Sci. 31, e4400 (2022).
Casadevall, G. et al. Estimating conformational heterogeneity of tryptophan synthase with a template-based Alphafold2 approach. Protein Sci. 31, e4426 (2022).
Dauparas, J. et al. Robust deep learning-based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Humphreys, I. R. et al. Computed structures of core eukaryotic protein complexes. Science 374, eabm4805 (2021).
Wang, J. et al. Scaffolding protein functional sites using deep learning. Science 377, 387–394 (2022).
Chen, X. et al. Target identification with quantitative activity based protein profiling (ABPP). Proteomics 17, https://doi.org/10.1002/pmic.201600212 (2017).
Fuerst, R. & Breinbauer, R. Activity-based protein profiling (ABPP) of oxidoreductases. Chembiochem 22, 630–638 (2021).
Xu, J. et al. Applications of activity-based protein profiling (ABPP) and bioimaging in drug discovery. Chem. Asian J. 15, 34–41 (2020).
Yang, W. et al. Non-classical ferroptosis inhibition by a small molecule targeting PHB2. Nat. Commun. 13, 7473 (2022).
Wang, S. et al. CavitySpace: a database of potential ligand binding sites in the human proteome. Biomolecules 12, 967 (2022).
Yuan, Y., Pei, J. & Lai, L. Binding site detection and druggability prediction of protein targets for structure-based drug design. Curr. Pharm. Des. 19, 2326–2333 (2013).
Wu, M. & Zhang, Y. Integrated bioinformatics, network pharmacology, and artificial intelligence to predict the mechanism of celastrol against muscle atrophy caused by colorectal cancer. Front. Genet. 13, 1012932 (2022).
Hegedűs, T., Geisler, M., Lukács, G. L. & Farkas, B. Ins and outs of AlphaFold2 transmembrane protein structure predictions. Cell Mol. Life Sci. 79, 73 (2022).
Wu, M. & Zhang, Y. Combining bioinformatics, network pharmacology and artificial intelligence to predict the mechanism of celastrol in the treatment of type 2 diabetes. Front. Endocrinol. 13, 1030278 (2022).
Ma, W. et al. Enhancing protein function prediction performance by utilizing AlphaFold-predicted protein structures. J. Chem. Inf. Model. 62, 4008–4017 (2022).
Hu, M. et al. Exploring evolution-based &-free protein language models as protein function predictors. Preprint at https://arxiv.org/abs/2206.06583 (2022).
Rappoport, D. & Jinich, A. Enzyme Substrate Prediction from Three-Dimensional Feature Representations Using Space-Filling Curves. J. Chem. Inf. Model. https://doi.org/10.1021/acs.jcim.3c00005 (2023).
Herbert, A. Z-DNA and Z-RNA in human disease. Commun. Biol. 2, 1–10 (2019).
Bartas, M. et al. Searching for new Z-DNA/Z-RNA binding proteins based on structural similarity to experimentally validated Zα domain. Int. J. Mol. Sci. 23, 768 (2022).
Dawson, J. E. et al. Shape shifting: the multiple conformational substates of the PTEN N-terminal PIP2 -binding domain. Protein Sci. 31, e4308 (2022).
Feng, Y. et al. Naturally occurring I81N mutation in human cytochrome c regulates both inherent peroxidase activity and interactions with neuroglobin. ACS Omega 7, 11510–11518 (2022).
Athanasios, A. et al. Protein-protein interaction (PPI) network: recent advances in drug discovery. Curr. Drug Metab. 18, 5–10 (2017).
Rabbani, G. et al. Protein-protein interactions and their role in various diseases and their Prediction Techniques. Curr. Protein Pept. Sci. 19, 948–957 (2018).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. Preprint at https://www.biorxiv.org/content/10.1101/2021.10.04.463034v2 (2022).
Gómez-Marín, E. et al. The high mobility group protein HMG20A cooperates with the histone reader PHF14 to modulate TGFβ and Hippo pathways. Nucleic Acids Res. 50, 9838–9857 (2022).
Ivanov, Y. D. et al. Prediction of monomeric and dimeric structures of CYP102A1 using AlphaFold2 and AlphaFold multimer and assessment of point mutation effect on the efficiency of intra- and interprotein electron transfer. Molecules 27, 1386 (2022).
Bryant, P. et al. Improved prediction of protein-protein interactions using AlphaFold2. Nat. Commun. 13, 1265 (2022).
Green, A. G. et al. Large-scale discovery of protein interactions at residue resolution using co-evolution calculated from genomic sequences. Nat. Commun. 12, 1396 (2021).
Yin, R., Feng, B. Y., Varshney, A. & Pierce, B. G. Benchmarking AlphaFold for protein complex modeling reveals accuracy determinants. Protein Sci. 31, e4379 (2022).
Gao, M., Nakajima, An. D., Parks, J. M. & Skolnick, J. AF2Complex predicts direct physical interactions in multimeric proteins with deep learning. Nat. Commun. 13, 1744 (2022).
Tsaban, T. et al. Harnessing protein folding neural networks for peptide-protein docking. Nat. Commun. 13, 176 (2022).
McCafferty, C. L. et al. Integrative modeling reveals the molecular architecture of the intraflagellar transport A (IFT-A) complex. Elife 11, e81977 (2022).
Maeda, Y. et al. Biomolecular fluorescence complementation profiling and artificial intelligence structure prediction of the Kaposi’s sarcoma-associated herpesvirus ORF18 and ORF30 interaction. Int. J. Mol. Sci. 23, 9647 (2022).
Liu, Y. et al. Cargo recognition mechanisms of yeast Myo2 revealed by AlphaFold2-powered protein complex prediction. Biomolecules 12, 1032 (2022).
Van Breugel, M., Rosa E Silva, I. & Andreeva, A. Structural validation and assessment of AlphaFold2 predictions for centrosomal and centriolar proteins and their complexes. Commun. Biol. 5, 312 (2022).
Österlund, N. et al. Mass spectrometry and machine learning reveal determinants of client recognition by antiamyloid chaperones. Mol. Cell Proteom. 21, 100413 (2022).
Jovine, L. Using machine learning to study protein-protein interactions: from the uromodulin polymer to egg zona pellucida filaments. Mol. Reprod. Dev. 88, 686–693 (2021).
Burke, D. F. et al. Towards a structurally resolved human protein interaction network. Nat. Struct. Mol. Biol. 30, 216–225 (2023).
Weghoff, M. C., Bertsch, J. & Müller, V. A novel mode of lactate metabolism in strictly anaerobic bacteria. Environ. Microbiol. 17, 670–677 (2015).
Kayastha, K. et al. Structure-based electron-confurcation mechanism of the Ldh-EtfAB complex. Elife 11, e77095 (2022).
Kimura, S. et al. Sequential action of a tRNA base editor in conversion of cytidine to pseudouridine. Nat. Commun. 13, 5994 (2022).
Liang, M. et al. Identification of a novel substrate motif of yeast separase and deciphering the recognition specificity using AlphaFold2 and molecular dynamics simulation. Biochem. Biophys. Res. Commun. 620, 173–179 (2022).
Lorenz, P. et al. The KRAB domain of ZNF10 guides the identification of specific amino acids that transform the ancestral KRAB-A-related domain present in human PRDM9 into a canonical modern KRAB-A domain. Int. J. Mol. Sci. 23, 1072 (2022).
Ecco, G., Imbeault, M. & Trono, D. KRAB zinc finger proteins. Development 144, 2719–2729 (2017).
McMullen, P. et al. Impacts of a Zwitterionic peptide on its fusion protein. Bioconjug. Chem. 33, 1485–1493 (2022).
Nowinski, A. K. et al. Sequence, structure, and function of peptide self-assembled monolayers. J. Am. Chem. Soc. 134, 6000–6005 (2012).
Keefe, A. J. et al. Screening nonspecific interactions of peptides without background interference. Biomaterials 34, 1871–1877 (2013).
Darai, N. et al. Theoretical studies on RNA recognition by Musashi 1 RNA-binding protein. Sci. Rep. 12, 12137 (2022).
Zhorov, B. S. & Dong, K. Pyrethroids in an AlphaFold2 Model of the Insect Sodium Channel. Insects 13, 745 (2022).
Ding, Y. W. et al. Directed evolution of the fusion enzyme for improving astaxanthin biosynthesis in Saccharomyces cerevisiae. Synth. Syst. Biotechnol. 8, 46–53 (2022).
Taka, J. R. H., Sun, Y. & Goldstone, D. C. Mapping the interaction between Trim28 and the KRAB domain at the center of Trim28 silencing of endogenous retroviruses. Protein Sci. 31, e4436 (2022).
Bentaleb, C. et al. The endocytic recycling compartment serves as a viral factory for hepatitis E virus. Cell Mol. Life Sci. 79, 615 (2022).
Mehrtash, A. B. & Hochstrasser, M. Elements of the ERAD ubiquitin ligase Doa10 regulating sequential poly-ubiquitylation of its targets. iScience 25, 105351 (2022).
Patel, O. et al. Crystal structure of the putative cell-wall lipoglycan biosynthesis protein LmcA from Mycobacterium smegmatis. Acta Crystallogr. D. Struct. Biol. 78, 494–508 (2022).
Zheng, L. et al. Distinct structural bases for sequence-specific DNA binding by mammalian BEN domain proteins. Genes Dev. 36, 225–240 (2022).
Tao, H. et al. Discovery of non-squalene triterpenes. Nature 606, 414–419 (2022).
Pasquadibisceglie, A., Leccese, A. & Polticelli, F. A computational study of the structure and function of human Zrt and Irt-like proteins metal transporters: An elevator-type transport mechanism predicted by AlphaFold2. Front. Chem. 10, 1004815 (2022).
Goulet, A., Mahony, J., Cambillau, C. & van, Sinderen, D. Exploring structural diversity among adhesion devices encoded by Lactococcal P335 phages with AlphaFold2. Microorganisms 10, 2278 (2022).
Goulet, A. et al. A structural discovery journey of streptococcal phages adhesion devices by AlphaFold2. Front. Mol. Biosci. 9, 960325 (2022).
Ries, J. I. et al. CipA mediates complement resistance of Acinetobacter baumannii by formation of a factor I-dependent quadripartite assemblage. Front. Immunol. 13, 942482 (2022).
Pinheiro, F. et al. AlphaFold and the amyloid landscape. J. Mol. Biol. 433, 167059 (2021).
Burnim, A. A. et al. Analysis of insertions and extensions in the functional evolution of the ribonucleotide reductase family. Protein Sci. 31, e4483 (2022).
Kolesnik, M. V. et al. Type III CRISPR-Cas systems: deciphering the most complex prokaryotic immune system. Biochemistry 86, 1301–1314 (2021).
Alvarez-Carreño, C., Penev, P. I., Petrov, A. S. & Williams, L. D. Fold evolution before LUCA: common ancestry of SH3 domains and OB domains. Mol. Biol. Evol. 38, 5134–5143 (2021).
Tang, Q. Y., Ren, W., Wang, J. & Kaneko, K. The statistical trends of protein evolution: a lesson from AlphaFold database. Mol. Biol. Evol. 39, msac197 (2022).
Sebastiano, M. R. et al. AI-based protein structure databases have the potential to accelerate rare diseases research: AlphaFoldDB and the case of IAHSP/Alsin. Drug Discov. Today 27, 1652–1660 (2022).
Iqbal, S. et al. PROST: AlphaFold2-aware sequence-based predictor to estimate protein stability changes upon missense mutations. J. Chem. Inf. Model. 62, 4270–4282 (2022).
Zhu, Y. et al. Deep whole-genome resequencing sheds light on the distribution and effect of amphioxus SNPs. BMC Genom. Data 23, 26 (2022).
Yang, Q. et al. Structural analysis of the SARS-CoV-2 Omicron variant proteins. Research 2021, 9769586 (2021).
Ivanov, Y. D. et al. Prediction of monomeric and dimeric structures of CYP102A1 using AlphaFold2 and AlphaFold multimer and assessment of point mutation effect on the efficiency of intra-and interprotein electron transfer. Molecules 27, 1386 (2022).
Pan, Q., Nguyen, T. B., Ascher, D. B. & Pires, D. E. V. Systematic evaluation of computational tools to predict the effects of mutations on protein stability in the absence of experimental structures. Brief. Bioinform. 23, bbac025 (2022).
Guan, W. et al. A lysine residue from an extracellular turret switches the ion preference in a Cav3 T-Type channel from calcium to sodium ions. J. Biol. Chem. 298, 102621 (2022).
Zeng, D. et al. A hemagglutinin stem vaccine designed rationally by AlphaFold2 confers broad protection against influenza B infection. Viruses 14, 1305 (2022).
Molini, B. et al. B-cell epitope mapping of TprC and TprD variants of treponema pallidum subspecies informs vaccine development for human treponematoses. Front. Immunol. 13, 862491 (2022).
Li, V. et al. In silico SARS-CoV-2 vaccine development for Omicron strain using reverse vaccinology. Genes Genomics 44, 937–944 (2022).
Dobson, C. M. Protein folding and misfolding. Nature 426, 884–890 (2003).
Daggett, V. & Fersht, A. R. Is there a unifying mechanism for protein folding? Trends Biochem. Sci. 28, 18–25 (2003).
Glazer, D. S., Radmer, R. J. & Altman, R. B. Improving structure-based function prediction using molecular dynamics. Structure 17, 919–929 (2009).
Hummer, G. & Köfinger, J. Bayesian ensemble refinement by replica simulations and reweighting. J. Chem. Phys. 143, 243150 (2015).
Childers, M. C. & Daggett, V. Validating molecular dynamics simulations against experimental observables in light of underlying conformational ensembles. J. Phys. Chem. B. 122, 6673–6689 (2018).
Yang, Y. I. et al. Enhanced sampling in molecular dynamics. J. Chem. Phys. 151, 070902 (2019).
Hanson, J., Paliwal, K. K., Litfin, T. & Zhou, Y. SPOT-Disorder2: improved protein intrinsic disorder prediction by ensembled deep learning. Genom. Proteom. Bioinform. 17, 645–656 (2019).
Guo, H. B. et al. AlphaFold2 models indicate that protein sequence determines both structure and dynamics. Sci. Rep. 12, 10696 (2022).
Del Alamo, D., Sala, D., Mchaourab, H. S. & Meiler, J. Sampling alternative conformational states of transporters and receptors with AlphaFold2. Elife 11, e75751 (2022).
Chang, L. & Perez, A. Deciphering the folding mechanism of proteins G and L and their mutants. J. Am. Chem. Soc. 144, 14668–14677 (2022).
Nussinov, R., Zhang, M., Liu, Y. & Jang, H. AlphaFold, artificial intelligence (AI), and allostery. J. Phys. Chem. B. 126, 6372–6383 (2022).
Chakravarty, D. & Porter, L. L. AlphaFold2 fails to predict protein fold switching. Protein Sci. 31, e4353 (2022).
He, X. et al. AlphaFold2 versus experimental structures: evaluation on G protein-coupled receptors. Acta Pharmacol. Sin. 44, 1–7 (2023).
Ward, J. et al. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J. Mol. Biol. 337, 635–645 (2004).
Peng, Z., Mizianty, M. J. & Kurgan, L. Genome-scale prediction of proteins with long intrinsically disordered regions. Proteins 82, 145–158 (2013).
Liu, Y., Wang, X. & Liu, B. A comprehensive review and comparison of existing computational methods for intrinsically disordered protein and region prediction. Brief. Bioinform. 20, 330–346 (2017).
Miskei, M., Horvath, A., Vendruscolo, M. & Fuxreiter, M. Sequence-based prediction of Fuzzy protein interactions. J. Mol. Biol. 432, 2289–2303 (2020).
Yang, X. et al. Molecular mechanism of allosteric modulation for the cannabinoid receptor CB1. Nat. Chem. Biol. 18, 831–840 (2022).
Reynisdottir, T., Anderson, K. J., Boukas, L. & Bjornsson, H. T. Missense variants causing Wiedemann-Steiner syndrome preferentially occur in the KMT2A-CXXC domain and are accurately classified using AlphaFold2. PLoS Genet. 18, e1010278 (2022).
McBride, J. M., Polev, K., Reinharz, V., Grzybowski, B. A., & Tlusty, T. AlphaFold2 can predict structural and phenotypic effects of single mutations. Preprint at https://www.biorxiv.org/content/10.1101/2022.04.14.488301v2 (2023).
Buel, G. R. & Walters, K. J. Can AlphaFold2 predict the impact of missense mutations on structure? Nat. Struct. Mol. Biol. 29, 1–2 (2022).
Pak, M. A. et al. Using AlphaFold to predict the impact of single mutations on protein stability and function. Preprint at https://www.biorxiv.org/content/10.1101/2021.09.19.460937v1 (2021).
Tolsma, T. O. & Hansen, J. C. Post-translational modifications and chromatin dynamics. Essays Biochem. 63, 89–96 (2019).
Samaržija, I. Post-translational modifications that drive prostate cancer progression. Biomolecules 11, 247 (2021).
Salas-Lloret, D. & González-Prieto, R. Insights in post-translational modifications: ubiquitin and SUMO. Int. J. Mol. Sci. 23, 3281 (2022).
Singh, V. et al. Phosphorylation: implications in cancer. Protein J. 36, 1–6 (2017).
Dai, X., Ren, T., Zhang, Y. & Nan, N. Methylation multiplicity and its clinical values in cancer. Expert Rev. Mol. Med. 23, e2 (2021).
Gil, J., Ramírez-Torres, A. & Encarnación-Guevara, S. Lysine acetylation and cancer: a proteomics perspective. J. Proteom. 150, 297–309 (2017).
Eichler, J. Protein glycosylation. Curr. Biol. 29, R229–R231 (2019).
Tikhonov, D. et al. Changes in protein structural motifs upon post-translational modification in kidney cancer. Diagnostics 11, 1836 (2021).
Devlin, J. et al. Bert: pre-training of deep bidirectional transformers for language understanding. Preprint at https://arxiv.org/abs/1810.04805 (2019).
Acknowledgements
The authors would like to thank Dr. Jun Zou (West China Hospital, Sichuan University) for useful discussion. This work was supported by grants from the National Key R&D Program of China (2021YFC2500203), Beijing Natural Science Foundation Haidian Origination and Innovation Joint Fund (L222007), the National Natural Science Foundation of China (32070670), and Innovation Project for Institute of Computing Technology, CAS. (E161080).
Author information
Authors and Affiliations
Contributions
R.C., Y.Z., and X.Z. conceived and supervised the literature review. Z.Y. performed the literature summary, data analysis and draft writing. Z.Y., R.C., Y.Z., and X.Z. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, Z., Zeng, X., Zhao, Y. et al. AlphaFold2 and its applications in the fields of biology and medicine. Sig Transduct Target Ther 8, 115 (2023). https://doi.org/10.1038/s41392-023-01381-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-023-01381-z
This article is cited by
-
Evolutionary divergence of TLR9 through ancestral sequence reconstruction
Immunogenetics (2024)
-
Weighted gene coexpression network analysis and machine learning reveal oncogenome associated microbiome plays an important role in tumor immunity and prognosis in pan-cancer
Journal of Translational Medicine (2023)
-
Deep learning algorithms applied to computational chemistry
Molecular Diversity (2023)
-
SARS-CoV-2 proteins structural studies using synchrotron radiation
Biophysical Reviews (2023)