Abstract
The abnormal regulation of alternative splicing is usually accompanied by the occurrence and development of tumors, which would produce multiple different isoforms and diversify protein expression. The aim of the present study was to conduct a systematic review in order to describe the regulatory mechanisms of alternative splicing, as well as its functions in tumor cells, from proliferation and apoptosis to invasion and metastasis, and from angiogenesis to metabolism. The abnormal splicing events contributed to tumor progression as oncogenic drivers and/or bystander factors. The alterations in splicing factors detected in tumors and other mis-splicing events (i.e., long non-coding and circular RNAs) in tumorigenesis were also included. The findings of recent therapeutic approaches targeting splicing catalysis and splicing regulatory proteins to modulate pathogenically spliced events (including tumor-specific neo-antigens for cancer immunotherapy) were introduced. The emerging RNA-based strategies for the treatment of cancer with abnormally alternative splicing isoforms were also discussed. However, further studies are still required to address the association between alternative splicing and cancer in more detail.
Similar content being viewed by others
Introduction
Since the first discovery of eukaryotic “split” genes in 1977 harboring the intervening sequences, it has been known that these intron sequences could be removed by the spliceosome complex.1,2 Increasing evidence has shown that ~94% of human genes have intronic regions during the pre-mRNA processes, and most eukaryotic genes undergo alternative splicing in a tempo-spatial-dependent manner,3 which is regulated by multiple RNA-binding proteins (RBPs) and depends on cis-acting elements and trans-acting factors. Different mature messenger RNA (mRNAs) with different functions could, therefore, be synthesized from a single gene through alternative splicing, which increases the complexity of mRNA for the diversity of proteins.4,5
Recently, it was reported that splicing may be closely associated with the occurrence of tumors, and that abnormal changes in alternative splicing could affect tumor progression.6 It could also disrupt the protein interaction pathways in tumor development.7,8
In the present review, the complexity of the splicing network, including cis-elements, spliceosome assembly and a plethora of trans-elements with antagonistic functions that define RNA maturation were first explained. Next, the function of alternative splicing as an oncogenic driver and/or passenger during tumor cell progression adaptive responses to metabolic stress and neo-angiogenesis was discussed, with emphasis on hematological malignancy. Finally, recent advances in therapeutics targeting splicing catalysis and splicing regulatory proteins were summarized, and emerging novel technologies were discussed using RNA-based therapies to modulate pathogenically spliced isoforms.
Regulatory splicing components and alternative splicing
The process of removing introns from pre-mRNAs and connecting the remaining exons to each other to produce mature mRNA is called splicing, while the different combination of exons in the mRNA producing diversified mature mRNA is called alternative splicing.9 RNA splicing can be accomplished inside the nucleus by an enzymatic machine termed spliceosome. Spliceosome mainly recognizes the junction of introns and exons, and generally follows the “GU-AG” rule to distinguish introns from exonic sequences by four canonical consensus sequences: (i) The 5′ splice-site (SS; characterized by a GU dinucleotide at the 5′ end of the intron), (ii) the 3′ SS (containing AG at the 3′ end of the intron), (iii) the branch point sequence (BPS; located upstream of the 3′ SS), and (iv) the polypyrimidine tract (located between the BPS and the 3′ SS).10 Alternative splicing depends on many trans-acting factors as well as certain cis-acting elements.11
Core spliceosome machinery
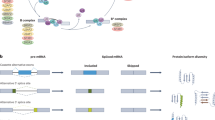
Alternative splicing is a regulated process, mainly executed by the spliceosome machinery and affected by the activity of splicing regulators, such as serine/arginine-rich (SR) proteins or heterogeneous nuclear ribonucleoproteins (hnRNPs), which are the most important mediators of SS recognition.12 The spliceosome is composed of 5 small nuclear RNPs (snRNPs; U1, U2, U4, U5, and U6 snRNPs), each of which contains its own snRNA complexed to a group of 300 associated proteins.13,14 Thanks to the recent structural studies using cryo-electron microscopy, an unprecedented high-resolution view of each step was obtained during spliceosome body assembly.15,16,17,18 These snRNPs bind to pre-mRNA, as shown in Fig. 1, U1 snRNP binds to 5′ SS, and U2 (including SF3A, SF3B) binds to 3′ SS and polypyrimidine sequences. Next, U5, U4/U6 were recruited, and a catalytically active complex was formed by rearrangement between snRNPs to complete intron excision and exon linkage.19,20
Spliceosome assembly. U1 snRNP recognizes 5′ SS and binds through base-pairing, SF1 binds to BPS, U2AF2 binds to polypyrimidine tract and U2AF1 binds to 3′ SS, forming Complex E. Next, U2 snRNP, with the assistance of U2AF, replaces SF1 with the BPS through base-pairing to form Complex A. Next, U5/U4/U6 is recruited and results in the rearrangement of Complex A. Among them, U4 and U6 snRNP are combined through complementary pairing of their RNA components, while U5 snRNP is loosely bound through protein interaction, at which time a Complex B is formed. Through a series of conformational changes, U1 snRNP leaves, U6 snRNP binds to 5′ SS, and at the same time, U4 snRNP leaves so that U6 snRNP and U2 snRNP pair through snRNA. After this rearrangement process, Pre-catalytic Spliceosome Complex B is formed, followed by two transesterification reactions. The first transesterification reaction generates Complex C. The rearrangements occur in Complex C, promoting second transesterification, resulting in a post-spliceosomal complex. As a result, exons are interconnected to form mature mRNA, introns are degraded and snRNPs are recycled. SF1 splicing factor 1, BPS branch point sequence, SS splice-site, snRNPs small nuclear RNPs
The SR proteins21 and hnRNP family12 are two essential auxiliary factors in enhancing or repressing splice-site usage by the recognition of specific cis-acting RNA elements. In general, SR proteins play a positive role in splicing regulation and preferentially bind to exonic splicing enhancer (ESE) and intronic splicing enhancer (ISE). Conversely, the binding of hnRNPs to exonic splicing silencer (ESS) and intronic splicing silencer (ISS) generally represses exon inclusion. Therefore, splicing factors could compete for the binding in the context of the cis-regulatory RNA elements and/or the recruitment of spliceosome component.22 In the present study, a simplified spliceosome assembly pathway and the core splicing factors required for exon/intron definition were summarized (Fig. 1).
Cis-acting elements and other RBPs
These cis-acting elements including the aforementioned 5′ SS, 3′ SS, BPS and splice enhancer (ESE, ISE) or silencer elements (ESS, ISS) in precursor mRNA are important for both constitutive and regulated splicing.23 The occurrence of alternative splicing is usually accompanied by changes in the ability of sequence-specific RBPs to bind to cis-acting sequences in their target pre-mRNA.24 Of note, both SR proteins and hnRNPs have been recently reported to be able to either promote or repress splicing when binding to different positions of pre-mRNAs.21,23,25 In addition, mutations in these cis-acting splice enhancer or silencer elements could largely affect alternative splicing.26
In addition to the above core components (snRNPs, SR proteins and hnRNPs), there are hundreds of RBPs genome-widely identified to bind to mRNA, including >300 previously not regarded as RBPs.27,28 This information will complicate the understanding of alternative splicing regulation based on competition or combination between different splicing factors and other RBPs [e.g., epithelial splicing regulatory protein 1 (ESRP1), RNA-binding motif proteins 4, 5, 6, 10]. The regulatory complexity of these groups of splicing factors could diversify the alternatively spliced RNA products.
Main alternative splicing patterns
In addition to removing intronic sequences, alternative splicing is also influenced by the RNA structure29 and other key regulatory pathways involved in gene regulation, such as transcription rate and the chromatin epigenetic signature.30,31 Alternative splicing events could affect one entire cassette exon or part of it through the usage of alternative splicing sites in a variant exon, and occur in genes bearing multiple transcription start sites or using multiple polyadenylation sites. The main alternative splicing patterns are divided into 5 types, as shown in Fig. 2: Exon skipping, intron retention, mutually exclusive exons, alternative 5′ SS and alternative 3′ SS. Growing evidence has revealed that mis-splicing events can change the function of protein and lead to human diseases. The study of its mechanism will provide important information for the treatment of diseases, including cancer initiation, maintenance and/or progression,32 which will be discussed in the next section.
The main alternative splicing patterns are divided into five types: exon skipping (also called cassette exon); intron retention; mutually exclusive exons (only some exons appear in mature mRNA); A5SS (the change of the splicing site causes the position of the 3′ end of the exon to change); A3SS (the change of the splicing site causes the position of the 5′ end of the exon to change). SS, splice site; A5SS, alternative SS
Functions of alternative splicing in cancer
Several studies have reported that alternative splicing is associated with tumors.33,34 During the last decades, recurrent somatic mutations in components of the human splicing machinery have occurred in human solid tumors, including bladder,35 brain,36 breast,37 cervix,38 colon,39 kidney,40 liver,41 lung,42 oral/HN,43 ovary,44 prostate,45 skin,46 stomach47,48 and thyroid49 tumors, as well as hematological malignancies, including acute myeloid leukemia (AML),50 myelodysplastic syndrome (MDS),51 chronic myelogenous leukemia,52 de novo AML,53 myelodysplastic syndrome without ringed sideroblasts (MDS w/o RS),54 myeloproliferative neoplasm MPN55 and refractory anemia with ringed sideroblasts and refractory cytopenia with multilineage dysplasia and ringed sideroblasts (RARS/RCMD).56 In addition to cancer, neurological diseases, such as Alzheimer’s disease (AD),57 Parkinson’s disease,58,59 Huntington’s disease (HD),60 schizophrenia,61,62 congenital myasthenic syndrome,63 Spinal muscular atrophy,64,65 and immunological and infectious diseases, such as celiac disease,66 psoriasis,67 systemic lupus erythematosus,68 asthma,69 inflammatory response,70 viral infections,71 cardiovascular disease72,73 and diabetes mellitus74,75 also have a connection with mis-splicing events (Table 1). Most of the diseases are due to either genetic mutation falling within the canonical RNA splicing sites, which directly influences mRNA maturation, or alterations in the expression level of spliceosomal/splicing regulatory factors that contribute to the splicing of pre-mRNA.
The transformation of normal cells into cancer cells mainly involves cellular proliferation, escape from cell death, growth inhibition, induction of angiogenesis, invasion and metastasis, energy metabolism and immune escape.76 This review focuses on four aspects: Proliferation and apoptosis, invasion and metastasis, and angiogenesis and metabolism under the light of alternative splicing (Fig. 3).
Roles of alternative splicing in tumorigenesis. The diagram illustrates different hallmarks of cancer along with examples of alternative splicing events that contribute to the transformation of normal cells into tumor cells mainly via proliferation and apoptosis, invasion and metastasis, angiogenesis and metabolism. Arrows up and down indicate the alternative isoforms contributing the most and the least to each process. ITGA6 integrin subunit α6, PKM pyruvate kinase, Bcl B-cell lymphoma, VEGF vascular endothelial growth factor, TAK1 TGF-β-activated kinase 1, CD44 cluster of differentiation 44
Balancing between proliferation and apoptosis
Tumor cells have the ability to divide more than they should, and to not die when they should. Therefore, there is a hallmark capability of cells to fine-tune and maintain a balance between proliferation and apoptosis. In general, some tumor suppressor genes, such as p53 and pRb, prevent normal cells from becoming carcinogenic by acting on the cell cycle and promoting genetic changes; it is therefore common for tumor cells to present an aberrant-splicing activity with an increased frequency of splicing isoforms that maintain the abnormal proliferative and apoptotic rhythm.77 Alternative splicing participates in the process of proliferation, differentiation and apoptosis via regulating the alternative expression of many oncogenic or tumor suppressor genes,78 as well as splicing factors. The integrin subunit α6 (ITGA6) pre-mRNA can be alternatively spliced into two splice isoforms: ITGA6A and ITGA6B. The pro-proliferative ITGA6A variant was enhanced in colon cancer cells, due to a process contributed to by Myc-mediated promoter activation, as well as the splicing factor epithelial splicing regulatory protein 2-mediated alternative splicing, thus promoting cell proliferation.79 Similarly, C-Myc can upregulate polypyrimidine tract-binding protein (PTB), hnRNPA1 and hnRNPA2 to change the splicing of pyruvate kinase (PKM) and make it develop into PKM2, which could promote tumor cell proliferation.80 Mutations in RNA-binding motif protein 10 (RBM10) had been identified in lung cancer cells, which disrupts the splicing of NUMB and inducing tumor proliferation.81 The complex apoptosis mechanisms involve two distinct regulatory pathways: The (extrinsic) death receptor-mediated pathway and the (intrinsic) mitochondrial pathway. Most SR splicing factors, such as RBMP, are associated with pro- and anti-apoptosis effects by influencing caspase 2, B-cell lymphoma (Bcl)-x, myeloid cell leukemia factor 1 (MCL-1), etc.82 For example, Bcl-x can obtain two different spliced isoforms, Bcl-xl and Bcl-xs, with Bcl-xl inhibiting apoptosis and Bcl-xs promoting it.83 It has been reported that PTB protein 1 (PTBP1)-overexpression can promote the 5′ SS end of the second exon of Bcl-x, which could produce Bcl-xs that promotes apoptosis; conversely, the downregulation of PTBP1 could produce Bcl-xl, which inhibits apoptosis.84 Induced MCL-1, as an apoptotic factor of the Bcl-2 family regulators, could cause apoptosis when its expression is decreased, and the alternative splicing of the Mcl-1 gene can be regulated by SR splicing factor (SRSF)5, as well as SRSF1.85 Therefore, the effect of alternative splicing on tumor cell proliferation and apoptosis may provide new therapeutic targets for cancer treatment.
Invasion and metastasis
Invasion and metastasis are two major obstacles in the treatment of cancer. Epithelial–mesenchymal transition (EMT) is abnormally activated during cancer metastasis and recurrence, which are dependent on interactions between tumor cells and the microenvironment.86 It may be important to define whether a tumor is invasive or metastatic, in order to determine its behavior. Transforming growth factor-β (TGF-β), as the main inducer of EMT, induces alternative splicing of TGF-β-activated kinase 1 to reject exon 12 during EMT.87 In EMT, the alternative splicing of cluster of differentiation 44 (CD44) has been changed through the regulation of the splicing factor ESRP1, which determines the interaction between CD44 and cell surface receptor tyrosine kinases.88 The generated CD44s could promote the occurrence and development of the EMT of breast cancer cells. ESRP1 and hnRNPM could compete for GU-rich binding sites in the pre-mRNA and modulate exon inclusion or skipping, defining either an epithelial or a mesenchymal state to determine a specific cell fate.89 It was recently revealed that hnRNPM and ESRP1 are key regulators in the EMT splicing program and are correlated with breast cancer.90
Angiogenesis
Angiogenesis is one of the critical features of tumor progression through the formation of new blood vessels. Many proteins act as vascularization activators, including basic fibroblast growth factor, tumor necrosis factor-α and vascular endothelial growth factor (VEGF). VEGF-A is composed of 8 exons, and exons 6, 7, and 8 alternately select 3′ and 5′ SSs to produce isoforms, which can control angiogenesis. VEGF-A could lead to both angiogenic and anti-angiogenic results, and might thus be utilized for anti-angiogenic therapeutics.91 It has been reported that inhibiting serine/arginine protein-specific splicing factor kinase 1 (SRPK1) by downregulating the tumor suppressor factor Wilms’ tumor suppressor 1 and indirectly suppressing SRSF1, could turn the splicing of VEGF into VEGF120 with an anti-angiogenic effect, thus inhibiting the growth of tumor endothelial cells.92 The evidence from a murine retinal model indicated that SRPK inhibitor 1 (SRPKIN-1), as an inhibitor of SRPK, can regulate the splicing of VEGF and then transform it into an anti-angiogenic VEGF-A165b isoform.93 Truncated glioma-associated oncogene homolog 1, as a splice variant of the GLI1 gene, could upregulate the expression of VEGF-A, thus enhancing the angiogenesis of glioblastomas.94 The alternative splicing of other factors in angiogenesis, in order to provide all the necessary nutrients for tumor development, need to be further demonstrated.
Alternative splicing in tumor metabolism
During tumor progression, cancer cells may experience nutrient deprivation and hypoxia, due to overgrowth. As compared with normal cells, tumor cells preferentially metabolize glucose through the aerobic glycolysis pathway, for which some genes encoding enzymes are susceptible to alternative splicing. The last step in glycolysis is the conversion of phosphoenolpyruvate to pyruvate, is catalyzed by PKM, which are encoded by two genes: PKLR and PKM, each producing two different variants.95,96 PKLR is mainly expressed in liver and hematopoietic cells, but the majority of tissues express PKM, coding PKM1 and PKM2 variants.97 Through alternative splicing of two mutually exclusive exons, exons 9 and 10, PKM1 is constitutively expressed in normal cells while PKM2 is expressed in tumor cells in a cell-signal-dependent manner.98 A study has shown that the alternative splicing of PKM is regulated by splicing factors: hnRNPA1/hnRNPA2, PTB/nPTB, and SRSF3,99 suggesting the role of splicing factors involved in alternative splicing for tumor metabolism. The upregulation of PKM2 is common in several types of cancer and contributes to a switch to glycolytic metabolism, reflecting the modulation of PKM1 and PKM2 ratio by alternative splicing to determine the choice between aerobic glycolysis and mitochondrial oxidative phosphorylation in response to metabolic stress.
Another aspect of tumor metabolism affected by alternative splicing events is the cellular response to hypoxia, which is a common characteristic of the inner bulk of many solid tumors. A recent RNA-seq study showed that breast cancer cells underwent extensive alternative splicing changes when cultured in hypoxic conditions, such as predominant intron retention events (for LDHA, TNFSF13 and ARHGAP4), as well as exon skipping (for MARCH7, PCBP2, and LRCH3).100
Alterations of splicing regulatory factors and related signal transduction pathways in tumor progression
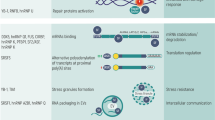
During alternative splicing under the regulation of certain splicing factors, specific splicing isoforms are produced.101 In tumors, especially in hematological malignancies, abnormal changes in splicing factors often occur.102 Hematological tumors are malignant clonal diseases of hematopoietic cells, with genetic mutations in ~85% of patients.103 This section contains a review of the detailed changes in core splicing factors SRSF2, SF3B1, U2AF1 and ZRSR2,104,105 since these are the most frequently mutated, as well as the contribution of other RBPs under the settings of related gene-regulatory network.
Aberrant expression of core splicing factors
SRSF2 (also as known as SC35) is rich in serine and arginine. As early as in 1990, SRSF2 was first discovered in HeLa cells and was identified as a non-snRNP protein-splicing component.106 The SRSF2 mutations often occur in the linking sequence of the RNA-recognition motifs (RRM) and RS regions, the mutations of which affect exons by recruiting U2AF to the upstream 3′ SS and U1 snRNP to the downstream 5′ SS.107,108 Studies have shown that mutations in SRSF2 have been found in hematological malignancies, with the highest frequency occurring in chronic myelomonocytic leukemia (CMML; 28–47%), and the lowest in MDS.109
Repeated mutations of the splicing factor SRSF2, such as P95 hotspot, often occur in hematological malignancies.110 Liang et al.111 confirmed the differential splicing of several hnRNP proteins by constructing an SRSF2P95H mutant cell line. The results showed that mutations in splicing factors could induce hematopoietic differentiation by affecting hnRNPA2B1 protein function, thereby promoting cancer development. Zhang et al.112 introduced common mutations in MDS into SRSF2. The results showed that the mutation of the splicing factor SRSF2 erroneously regulated the splicing of the pre-mRNA and produced abnormal proteins that could drive cancer.
SF3B1 (SF3B155) is a key component of U2 snRNP in the spliceosome. It binds to intron branch SSs and affects the 3′ SS. The most common mutation hotspots are K700, E622, R625, H662, and K666. SF3B1 has significant mutations not only in hematological malignancies, but also in solid tumors, which occur in 48–79% of RARS and RCMD-RS 6–26% of chronic lymphocytic leukemia (CLL).109
Mutations of SF3B1 were first found in patients with MDS.113 It was also found that mutations in SF3B1 were closely associated with the presence of ring sideroblasts in MDS, suggesting that mutations in SF3B1 are associated with refractory anemia with RARS.56 Wang et al.114 found that mutations in SF3B1 caused dysfunction in cells, including DNA damage, affecting Notch signaling, etc., thus indicating that it is associated with poor prognosis of CLL.
U2AF1 is also known as U2AF35. It can directly bind to the 3′ SS,115 with S34 and Q157 as the mutation hotspots. U2AF1 mutations mainly occur in hematological tumors, especially MDS (5–12%), CMML (8–17%)109 and lung cancer (3%).116 The transgenic mouse model showed that the mutations in U2AF1 altered the function of mouse hematopoietic cells, and simultaneously changed the splicing of precursor mRNA and the expression of downstream gene subtypes, U2AF1 (S34F) compared with U2AF1 (WT) transgenic mice, resulting in abnormal hematopoietic function in patients with MDS.117 The mutation of the S34 mutation hotspot in U2AF1 causes abnormal alternative splicing, affecting gene expression through canonical or non-canonical roles of translation regulation.118,119
ZRSR2 is also a member of the arginine and serine rich family and affects the 3′ SS through U12. ZRSR2 mutations are evenly distributed throughout the gene sequence, so there are no mutation hotspots. ZRSR2 mutations also mainly occur in hematological tumors. ZRSR2 have smaller mutations in MDS (1–11%) and CMML (0.8–8%), as compared to SRSF2, SF3B1 and U2AF1.109 By knocking down ZRSR2 in leukemia cells, which showed a slow growth trend, as compared with the control group, confirming the special role of ZRSR2 in RNA splicing and the U12 intron splicing disorder MDS.120 Of note, the three components of the major spliceosome (SF3B1, SRSF2, and U2AF1) generally occur in a mutually exclusive manner as heterozygous change-of-function mutations; ZRSR2 does not follow this pattern, suggesting an unknown association between the four spliceosome gene mutations and their potential roles in oncogenesis, which are unique to the different subtypes of hematological malignancies.121
According to reports, alterations in other splicing factors, such as PRPF8, SF3A1, LUCL7L2, SF1, U2AF2, PRPF8, PRPF40B, SRSF6, SRSF1, SRSF7, TRA2β, SRRM2, DDX1, DDX23, and CELF4108,122,123,124,125 have very low frequencies among hematological tumors.
Contribution of other RBPs to dysregulated alternative splicing
The expression of many RBPs can be autoregulated by the splicing of their own pre-mRNA, with the enhanced inclusion of premature termination codon-containing cassette exons, termed “poison exons”. These transcripts could be degraded by a nonsense-mediated decay (NMD) method, forming a negative feedback loop when SR-protein levels are upregulated.126,127 The coupling of splicing-factor-mediated alternative splicing with NMD in tumors needs further investigation. Furthermore, many other RBPs have been reported to be involved in the dysregulated alternative splicing of cancer cells.27,28 Herein hnRNPK and RNA-binding motif proteins were used as examples.
hnRNPK is an RBP containing three K homology domains (KH1, KH2 and KH3) and is closely associated with tumors.128 hnRNPK could play either a tumor suppressive or an oncogenic role in different types of cancer. hnRNPK has been shown to contribute to leukemogenesis in AML patients with deletions on the long arm of chromosome 9, which is correlated with a reduced expression of hnRNPK.129 However, another study found that hnRNPK is overexpressed in diffuse large B-cell lymphoma patients without Myc genomic alterations, representing a mechanism of C-Myc activation without Myc lesions, due to hnRNPK overexpression.130 In addition, the overexpression of hnRNPK can promote lung metastasis,131 and the increased hnRNPK expression is observed in multiple types of cancer, including breast, prostate, colorectal, gastric and pancreatic cancers.132 Therefore, considering the inhibition of tumor cell growth by hnRNPK as a haploinsufficient tumor repressor in myeloid malignancies,133 it is tempting to speculate that the dichotomous roles of hnRNPK in tumorigenesis may reflect the diverse hnRNPK functions in different cellular contexts. However, the exact molecular mechanisms of hnRNPK in tumorigenesis remain unclear and require further study.
RNA-binding motif proteins 4, 5, 6, and 10 are RBPs, which can regulate alternative splicing. In breast cancer cells, elevated SRPK1 induces cytoplasmic accumulation of phosphorylated RBM4, and eliminates its pro-apoptotic effect by inducing RBM4-regulated splicing transcripts of IR-B and MCL-1S.134 Downregulated RBM5 inhibits bladder cancer cell apoptosis.135 RBM6 could reduce the expression of EGFR, extracellular signal regulated kinase (ERK) and phosphorylated (p)-ERK in vitro and in vivo, thus repressing the growth and progression of laryngocarcinoma.136 Studies have shown that RBM10 inhibits the proliferation of endometrial cancer.137,138 Conversely, in lung adenocarcinoma, RBM10-overexpression reduced p53 expression in A549 cells and inhibited apoptosis;139 therefore, RBM10 could also promote lung adenocarcinoma cell proliferation through the RAP1/AKT/CREB signaling pathway.140 Many other RBPs, as splicing factors, need to be further characterized, especially in the setting of the complicated gene-regulatory network of alternative splicing.
Signal transduction driving alternative splicing during tumorigenesis
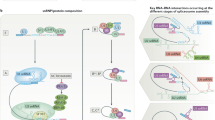
Although alterations in some splicing factors (snRNPs, SR proteins and hnRNAPs) have been described, a major driver of dysregulated splicing is oncogenic signal transduction pathways. Signaling could affect the expression of splicing factors at either the transcriptional or the post-translational modification levels, both of which modulate the subcellular localization to the nucleus or cytoplasm in a given cell. For example, in the process of spliceosome assembly and catalysis, the activity of SR proteins is usually regulated by phosphorylation.23 SRPKs in the cytoplasm and CDC-like kinases in the nucleus are two families of kinases involved in the regulation of SR-protein phosphorylation.141 These biochemical events are usually chained to transmit signals from receptors with upstream ligand stimulating downstream effectors inside the cell. Aberrant transduction of such signaling pathways is common during tumorigenesis and provides promising targets for drug development to control the growth and survival of malignant cells. These cancer cell intrinsic alterations were found to cause dysregulated alternative splicing, as will be briefly discussed in this section (Fig. 4):
Oncogenic signal transduction driving alternative splicing in cancer. The RAS/RAF/ERK, PI3K/AKT and Wnt signaling pathways play an important role in regulating splicing factor via transcriptional regulation and/or post-translational modification, which lead to aberrant-splicing events that promote oncogenesis. P phosphorylation, Wnt Wingless, ERK extracellular signal regulated kinase, PI3K phosphatidylinositol 3-kinase
The RAS/RAF/ERK pathway, which is characterized by the activation of the small GTPase RAS with a cascade stimulation of three mitogen-activated protein kinases (MAPK; namely RAF, MEK and ERK), is a hallmark event in many epithelia cell-derived tumors. Transcriptomic RNA-seq data from colon adenocarcinoma or lung squamous cell carcinoma showed that the oncogenic KRAS was correlated with an ETS transcription factor-mediated aberrant-splicing regulatory network. The network involved the induction of the expression of the alternative splicing factor PTBP1, which was associated with a shift in the alternative splicing of transcripts encoding the small GTPase RAC1, adaptor protein NUMB and PKM.142 The MEK/ERK signaling pathway was also reported to mediate the phosphorylation of splicing activator DAZAP1, which is required for its cytoplasm-to-nucleus translocation, as well as splicing regulatory activity.143 In addition, the activation of the MAPK/ERK downstream of RAS could cause the phosphorylation of splicing factors, such as signal transduction and activation of RNA metabolism 68 (SAM68). Phospho-SAM68 could then bind to and promote an intron retention event in the 3’UTR transcript of the SRSF1, thus leading to the subsequent switch in splicing profile, such as the RON gene transcripts.144,145 Of note, it was recently reported that the increased hnRNPA2 expression in hepatocellular carcinoma could induce an alternative splicing switch that decreases the dominant-negative isoform of A-RAF, leading to the activation of the RAF/MEK/ERK pathway and cellular transformation.146 This example highlights that the manipulation of certain alternative splicing variants can also shape cell signaling or even cell fates, which could be exploited to investigate the feedback regulatory mechanisms or therapeutic uses (see section 6).
The phosphatidylinositol 3-kinase (PI3K)/AKT pathway is another key pathway involved in regulating cell survival and apoptosis in a variety of tumors. For example, activated AKT has been shown to phosphorylate SRSF1 in lung cancer cells, thereby generating an anti-apoptotic caspase-9b isoform through the exclusion of an exon 3,4,5,6 cassette.147 EGF signaling was reported to regulate splicing through the phosphorylation of AKT/SRPK/SR protein148 and/or SPSB1-hnRNPA1 ubiquitination.149 The PI3K/AKT pathway is also known to activate the mammalian target of rapamycin complex 1 (mTORC1), a key regulator of cell metabolism and growth. The signaling of the mTORC1-S6K1 axis promotes the phosphorylation and cytoplasm-to-nucleus translocation of kinase SRPK2, which further activates SR proteins to participate in the splicing of lipogenesis-related transcripts to fuel tumor cell metabolism.150
Wingless (Wnt) signaling is well known for regulating development and stemness, and for its close association with many cancer types, especially colorectal cancer (CRC).151 Glycogen synthase 3β is part of the canonical Wnt pathway and has been reported to direct the phosphorylation of splicing factors, such as SRSF2152 or PTB-associated splicing factor (PSF).153 Activated Wnt/β-catenin signaling could directly enhance the transcript level of SRSF3.154 SRSF3 has also been reported to negatively affect one alternative exon inclusion variant of RAS-related C3 botulinum toxin substrate 1b.155
In addition to the three signaling pathways described above, there are many other tumor microenvironment-derived soluble factors, metabolic stress conditions and extracellular matrix-related signals, which also have a crosstalk with alternative splicing and are not discussed here.155 Therefore, more detailed studies are required to investigate the intricate interplay between oncogenic signaling pathways and dysregulated splicing in cancer.
Connection between mis-regulated splicing events of non-coding RNAs and cancer
During the past decades, thousands of non-coding RNAs (ncRNAs), including long ncRNAs (lncRNAs) and circular RNAs (circRNAs), have been characterized and found to play important biological roles in many cellular aspects, including cancers.156,157 Some mis-spliced RNA transcripts may also contribute to tumorigenesis.
lncRNAs and circRNAs
lncRNAs belong to an RNA subgroup and are usually longer than 200 nucleotides, have an important regulatory effect on cellular metabolism and are involved in the regulation of alternative splicing.158,159 At present, many studies have shown that the occurrence of diseases is associated with the mis-splicing of lncRNAs (Table 2). Serine/threonine-protein phosphatase 1 regulatory subunit 10 (lncRNA-PNUTS) is generated with the involvement of hnRNPE1, a non-coding isoform of PNUTS, and could functionally serve as a competitive sponge for miR-205 in breast tumor implantation, growth and metastasis.160 lncRNA HOXB-AS3 could interact with the ErbB3-binding protein 1 (EBP1) and regulate ribosomal RNA transcription and de novo protein synthesis in NPM1-mutated AML.161 The knockdown of metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) in ovarian cancer led to a reduced expression of RBP fox-1 homolog 2 (RBFOX2), thus inhibiting tumor cell proliferation.162 MALAT1 could also release SFPQ (also known as PSF (PTB-associated splicing factor)) from SFPQ/PTBP2 complex by binding to SFPQ to promote tumor growth and migration in CRC.163 In CRC, LINC01133 interacts with SRSF6 to inhibit EMT and metastasis.164 HOX transcript antisense RNA (HOTAIR) is an indicator of cell cycle dysregulation in lung cancer.165 An increasing number of mis-spliced lncRNAs are being reported in other diseases, such as adipogenesis,166 retinal diseases,167 Prader-Willi syndrome,168 and AD.169
circRNAs are widely expressed endogenous RNAs via a back-splicing process, which is the covalent joining of a downstream splice donor site with an upstream splice acceptor site.156 Since its biogenesis is performed by the canonical spliceosome machinery and regulated by the same cis-regulatory elements and trans-acting factors as that control linear mRNA splicing, circRNAs can be regarded as an additional form of alternative splicing,170 and in fact certain circRNAs are expressed in the context of cancer-specific alternative spicing.171
circRNAs may regulate multiple biological processes, and their aberrant expression has been reported in several types of cancer, even if the majority of circRNAs still lack functional annotations.172 Recent reports have identified their potential in gene expression regulation at the transcription and post-transcription levels through their interaction with the elongating RNA polymerase II complex173 or the U1 snRNP.174 Furthermore, circRNAs biogenesis may compete with pre-mRNA maturation, thus decreasing the levels of linear mRNAs that harbor circularized exons.175 In addition, circRNAs can function as sponges for microRNAs (miRNAs)176 to sequester normal functional miRNA. It may also act as a decoy for RBPs to regulate intracellular mRNA fate,177 or even produce abnormal small peptides if they contain internal ribosome entry site elements and AUG initiator sites in the open-reading frame, thus expanding the regulatory repertoire of circRNA.178,179 It is therefore tempting to speculate that circRNAs, as well as other RNA spliced variants, could be used as potential biomarkers for cancers, such as leukemia and lung cancer.180,181
Mis-regulated splicing events acting as oncogenic drivers and passenger factors in tumorigenesis
The above description indicates that mis-splicing events could be generally regarded as oncogenic drivers, which is indeed supported by recent multi-omic data from large-scale samples.182,183,184 However, aberrant spliced transcripts may also act as bystander or passenger factors in tumorigenesis. A recent study on multiple myeloma, using unbiased phosphoproteomics and transcriptomic/exomic sequencing following treatment with the proteasome inhibitor carfilzomib, found that the splicing-related proteins underwent the most prominent phosphorylation changes, and that the spliceosome might be a specific vulnerability in this hematological malignancy.185 Kahles et al.186 have provided a landscape of the splicing differences between the most common tumors and the healthy counterpart samples from 8,705 patients, suggesting that the increased splicing diversity in tumors might be an effect rather than a cause of oncogenesis. They also highlighted that certain cancer-specific alternative splice isoforms could produce neo-antigens, which can be used as a novel strategy for cancer immunotherapy (see next section).
Modulation of splicing as cancer therapeutics
As mentioned earlier, there is an important link between alternative splicing and cancer.187 Abnormal alternative splicing, which may produce abnormal proteins, could be explored as a marker for cancer diagnosis188 and a new target for cancer treatment.189 At present, small molecules and splice-switching antisense oligonucleotides (SSOs) are two major validated methods that target alternative splicing for the treatment of cancer (Fig. 5).
Modulation of splicing as potential cancer therapeutics. Several representative strategies are depicted in the simplified splicing regulation diagram, including small molecules targeting the core spliceosome SF3b-complex compound, SSOs, ASO, short hairpin RNA interference/small interference RNA, clustered regularly interspaced short palindromic repeats (CRISPR)-associated (Cas) system, single-BEs (CBEs or ABEs) as well as antibodies against tumor-specific neo-antigen due to alternative splicing. SR serine/arginine-rich, ESE exonic splicing enhancer, ESS exonic splicing silencer, ISE intronic splicing enhancer, ISS intronic splicing silencer, SSOs splice-switching antisense oligonucleotides, ASO antisense oligonucleotide, BEs base editors, CBEs cytosine-BEs, ABEs adenine-BEs, CRISPR-Cas clustered regularly interspaced short palindromic repeats-Cas
Small molecules modulating the levels of splicing regulators
Since some specific splice isoforms are used as potential tumor markers and therapeutic targets, it is reasonable to design inhibitors targeting splicing factors.190 In fact, SF3B1, having the highest mutation degree in various hematological malignancies, is often used as a target for cancer treatment. Some small molecules, such as spliceostatin A, pladienolide-B, GEX1A and E1707, can inhibit the splicing of SF3B1, and have a selective effect on tumor cells.191 For example, Amiloride is a small molecule first discovered in the 1960’s, which has been used clinically to treat edema and hypertension.192 It has been validated that amiloride can change the alternative splicing of Bcl-x, HIPK3 and RON/MISTR1 by affecting hypo-phosphorylation of splicing factor SF2/ASF and decreasing the expression of SRP20 and other SR proteins.193 H3B-8800 is an orally available small-molecule splicing modulator derived from pladienolide-B with a structural similarity to E7107 but with less potency, which interacts directly with the splicing factor SF3b complex to kill spliceosome-mutated epithelium and blood tumor cells by causing enhanced retention of GC-rich introns.194 H3B-8800 has recently been used in a phase 1 clinical trial (NCT02841540) targeting patients with relapsed/refractory myeloid neoplasms (MDS, CMML and AML) that carry splicing factor mutations (https://clinicaltrials.gov/ct2/show/NCT02841540).
Although the effects of small-molecule drugs are significant, they have an obvious disadvantage: They easily miss targets. Therefore, further clinical research on these small-molecule drugs is required to ensure that their mechanism of action is fully understood.121
SSOs
SSOs can affect the function of splicing factors through splicing modification, which has become an effective treatment method.181,195 The prototype for SSO technology is the antisense oligonucleotides (ASOs), which have been approved for the treatment of spinal muscular atrophy (SMA)196 and Duchenne muscular dystrophy.197 SMA is a congenital neuromuscular disease characterized by the loss of motor neurons, which leads to progressive muscle weakness and is difficult to cure.198 Many efficient ASOs are under development or have already been tested in clinical trials for the treatment of myotonic dystrophy (NCT023412011), HD (NCT02519036), amyotrophic lateral sclerosis (ALS; NCT02623699) and AD (NCT03186989).199
This base-pairing of oligonucleotides to target RNA can induce degradation or interfere with the splicing of pre-mRNA. In order to improve the stability of synthetic oligonucleotides, replacing the ribose ring of the oligonucleotide subunits with a morpholine ring, termed morpholino,200 seems especially suitable for targeting splicing, as termed morpholino are refractory to RNase H activity and thus not directly degrade the pre-mRNA. Studies have shown that Bcl-x SSOs could be combined with the downstream 5′ SS of the exon 2 in pre m-RNA of Bcl-x and modify Bcl-x pre-mRNA splicing. The pro-apoptotic effect on tumor cell lines demonstrates the anti-tumor activity of Bcl-x pre-mRNA spliced SSO.201,202 The decoy RNA oligonucleotides were designed and confirmed to inhibit the splicing and biological activity of RBFOX1/2, SRSF1 and PTBP1.203 Therefore, SSOs will be an effective way to treat tumors caused by the vital mis-spliced events during disease initiation and/or progression. Ongoing efforts to discover pathogenic isoform alterations are being made to exploit the full potential of this therapeutic approach.
Generation of alternative tumor-specific antigens for cancer immunotherapy
The advent of immunotherapy has revolutionized the treatment of many solid tumors, as well as blood cancers, and the research of suitable antigens specific to tumors for cancer vaccines and T cell therapies has accelerated but remains a challenge within the past decade. Based on certain different patterns of alternative splicing of mRNA between tumors and normal tissues, the development of neo-antigens that are produced by aberrant spliced mRNA and are not recognized by the immune system, including splice variants, gene fusions and other processes,204 would be a novel strategy. Over the past decades, there have been a few experimentally validated splicing-derived peptides with neo-epitopes that are recognized by T cells with evidence of immunogenicity. In a study on chronic myeloid leukemia, peptides derived from alternatively spliced out-of-frame BCR/ABL fusing transcripts were able to stimulate a peptide-specific cytotoxic T lymphocyte response, evidenced by the detection of out-of-frame peptide-specific IFNγ + CD8 + T cells in patients and the killing of peptide-pulsed target cells in vitro by these cytotoxic T lymphocytes.205 Another recent study on B-cell lineage marker CD20 showed that its alternative splicing isoform with a 168-nucleotide spliced out in exons 3–7 was only present in several patient-derived B lymphoma cell lines but not normal cells, and could generate a CD20-derived peptide with HLA-DR1 binding epitopes and vaccination, thus eliciting epitope-specific CD4+ and CD8+ responses in transgenic mice.206 Although there are many computational tools available and high-throughput studies to predict, screen and validate the presentation and immunogenicity of generated peptides, the field of investigating tumor-specific-splicing events as a promising anticancer treatment strategy remains in its nascence and presents many challenges, such as specific and cross-reactive immunogenicity, as well as tumor clonal heterogeneity (see recent more comprehensive reviews207,208).
Other RNA-based therapeutic approaches
Targeting mis-spliced RNA transcripts during tumorigenesis remains relatively nascent. Novel technologies such as short hairpin RNA interference, small interference RNA,209 clustered regularly interspaced short palindromic repeats (CRISPR)-Cas directed gene editing,210 and even single-base editors (BEs) cytosine-BEs (CBEs) or adenine-Bes (ABEs) represent powerful and exciting strategies.211,212 Recently, the CRISPR-Cas13a enzyme could target and edit RNA transcripts,213,214 such as endogenous post-transcriptional RNA editing and alternative polyadenylation, representing another approach to targeting RNA molecules from variant or aberrant-splicing processes with a potentially great clinical utility in the future, once its feasibility and safety have been established in humans.
Conclusion and future perspectives
In conclusion, alternative splicing can produce multiple isoforms with diverse functions for the same gene loci. The dysregulation of alternative splicing is closely associated with tumor progression, which is becoming a hot topic in the field of cancer research, providing new ideas for attractive methods of treating cancer. In-depth studies of the regulatory mechanism of alternative splicing have extended our understanding of tumorigenesis. In the light of high-throughput data of the genome, transcriptome, proteome and epigenome from large-scale samples and functional characterization, the aberrant-splicing events could be involved in tumorigenesis as oncogenic drivers and/or passengers. Targeted splicing has also been explored as a new treatment option for cancer and other diseases, including novel RNA-based CRISPR-Cas13a editing technology, SSOs, as well as small molecules that modulate splicing. By knocking down PTBP1 using ASOs or CRISPR-CasRx technologies, two recent papers successfully converted glial cells to neurons in mouse models of Parkinson’s disease.215,216 However, more recent studies have shown that one of the major challenges of targeting the splicing events and/or aberrant RNA species at the post-transcriptional level are the specificity and delivery efficiency.
Therefore, exploring the mutation site and studying the abnormality caused by mis-splicing in cancer is of great importance, as it may provide novel ideas for the follow-up research of splicing-targeting drugs and improve the promising RNA-based anti-tumor therapy.
References
Berget, S. M., Moore, C. & Sharp, P. A. Spliced segments at the 5′ terminus of adenovirus 2 late mRNA. Proc. Natl Acad. Sci. USA 74, 3171–3175 (1977).
Chow, L. T., Roberts, J. M., Lewis, J. B. & Broker, T. R. A map of cytoplasmic RNA transcripts from lytic adenovirus type 2, determined by electron microscopy of RNA:DNA hybrids. Cell 11, 819–836 (1977).
Pan, Q. et al. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Nilsen, T. W. & Graveley, B. R. Expansion of the eukaryotic proteome by alternative splicing. Nature 463, 457–463 (2010).
Biamonti, G. et al. The alternative splicing side of cancer. Semin. Cell Dev. Biol. 32, 30–36 (2014).
Agrawal, A. A., Yu, L., Smith, P. G. & Buonamici, S. Targeting splicing abnormalities in cancer. Curr. Opin. Genet. Dev. 48, 67–74 (2018).
Climente-Gonzalez, H., Porta-Pardo, E., Godzik, A. & Eyras, E. The functional impact of alternative splicing in cancer. Cell Rep. 20, 2215–2226 (2017).
Bonnal, S. C., Lopez-Oreja, I. & Valcarcel, J. Roles and mechanisms of alternative splicing in cancer-implications for care. Nat. Rev. Clin. Oncol. 17, 457–474 (2020).
Faustino, N. A. & Cooper, T. A. Pre-mRNA splicing and human disease. Genes Dev. 17, 419–437 (2003).
Shenasa, H., Movassat, M., Forouzmand, E. & Hertel, K. J. Allosteric regulation of U1 snRNP by splicing regulatory proteins controls spliceosomal assembly. RNA 26, 1389–1399 (2020).
Busch, A. & Hertel, K. J. Evolution of SR protein and hnRNP splicing regulatory factors. Wiley Interdiscip. Rev. RNA 3, 1–12 (2012).
Hoskins, A. A. et al. Ordered and dynamic assembly of single spliceosomes. Science 331, 1289–1295 (2011).
Jurica, M. S. & Moore, M. J. Pre-mRNA splicing: awash in a sea of proteins. Mol. Cell 12, 5–14 (2003).
Wan, R. et al. The 3.8 A structure of the U4/U6.U5 tri-snRNP: Insights into spliceosome assembly and catalysis. Science 351, 466–475 (2016).
Yan, C. et al. Structure of a yeast spliceosome at 3.6-angstrom resolution. Science 349, 1182–1191 (2015).
Wan, R. et al. Structure of a yeast catalytic step I spliceosome at 3.4 A resolution. Science 353, 895–904 (2016).
Yan, C. et al. Structure of a yeast activated spliceosome at 3.5 A resolution. Science 353, 904–911 (2016).
Dredge, B. K., Polydorides, A. D. & Darnell, R. B. The splice of life: alternative splicing and neurological disease. Nat. Rev. Neurosci. 2, 43–50 (2001).
Matlin, A. J., Clark, F. & Smith, C. W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Howard, J. M. & Sanford, J. R. The RNAissance family: SR proteins as multifaceted regulators of gene expression. Wiley Interdiscip. Rev. RNA 6, 93–110 (2015).
Martinez-Montiel, N. et al. Alternative Splicing as a target for cancer treatment. Int. J. Mol. Sci. 19, 545–572 (2018).
Fu, X. D. & Ares, M. Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet. 15, 689–701 (2014).
Goldammer, G. et al. Characterization of cis-acting elements that control oscillating alternative splicing. RNA Biol. 15, 1081–1092 (2018).
Geuens, T., Bouhy, D. & Timmerman, V. The hnRNP family: insights into their role in health and disease. Hum. Genet. 135, 851–867 (2016).
Yoshimi, A. & Abdel-Wahab, O. Molecular pathways: understanding and targeting mutant spliceosomal proteins. Clin. Cancer Res. 23, 336–341 (2017).
Barbosa-Morais, N. L., Carmo-Fonseca, M. & Aparicio, S. Systematic genome-wide annotation of spliceosomal proteins reveals differential gene family expansion. Genome Res. 16, 66–77 (2006).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Jacobs, E., Mills, J. D. & Janitz, M. The role of RNA structure in posttranscriptional regulation of gene expression. J. Genet. Genomics 39, 535–543 (2012).
Luco, R. F. et al. Regulation of alternative splicing by histone modifications. Science 327, 996–1000 (2010).
Naftelberg, S., Schor, I. E., Ast, G. & Kornblihtt, A. R. Regulation of alternative splicing through coupling with transcription and chromatin structure. Annu. Rev. Biochem. 84, 165–198 (2015).
Lee, Y. & Rio, D. C. Mechanisms and regulation of alternative Pre-mRNA splicing. Annu. Rev. Biochem. 84, 291–323 (2015).
Dvinge, H., Kim, E., Abdel-Wahab, O. & Bradley, R. K. RNA splicing factors as oncoproteins and tumour suppressors. Nat. Rev. Cancer 16, 413–430 (2016).
Singh, B. & Eyras, E. The role of alternative splicing in cancer. Transcription 8, 91–98 (2017).
Xie, R. et al. Polypyrimidine tract binding protein 1 promotes lymphatic metastasis and proliferation of bladder cancer via alternative splicing of MEIS2 and PKM. Cancer Lett. 449, 31–44 (2019).
Babic, I. et al. EGFR mutation-induced alternative splicing of Max contributes to growth of glycolytic tumors in brain cancer. Cell Metab. 17, 1000–1008 (2013).
Anczukow, O. et al. The splicing factor SRSF1 regulates apoptosis and proliferation to promote mammary epithelial cell transformation. Nat. Struct. Mol. Biol. 19, 220–228 (2012).
Liu, F. et al. SRSF10-mediated IL1RAP alternative splicing regulates cervical cancer oncogenesis via mIL1RAP-NF-kappaB-CD47 axis. Oncogene 37, 2394–2409 (2018).
Zhou, X. et al. BCLAF1 and its splicing regulator SRSF10 regulate the tumorigenic potential of colon cancer cells. Nat. Commun. 5, 4581 (2014).
Sokol, E. et al. microRNA-mediated regulation of splicing factors SRSF1, SRSF2 and hnRNP A1 in context of their alternatively spliced 3’UTRs. Exp. Cell Res 363, 208–217 (2018).
Duriez, M. et al. Alternative splicing of hepatitis B virus: a novel virus/host interaction altering liver immunity. J. Hepatol. 67, 687–699 (2017).
Sheng, J. et al. SRSF1 modulates PTPMT1 alternative splicing to regulate lung cancer cell radioresistance. EBioMedicine 38, 113–126 (2018).
Peiqi, L. et al. Expression of SRSF3 is correlated with carcinogenesis and progression of oral squamous cell carcinoma. Int. J. Med. Sci. 13, 533–539 (2016).
Iborra, S. et al. Alterations in expression pattern of splicing factors in epithelial ovarian cancer and its clinical impact. Int. J. Gynecol. Cancer 23, 990–996 (2013).
Fan, L. et al. Histone demethylase JMJD1A promotes alternative splicing of AR variant 7 (AR-V7) in prostate cancer cells. Proc. Natl Acad. Sci. USA 115, E4584–E4593 (2018).
Jensen, M. A., Wilkinson, J. E. & Krainer, A. R. Splicing factor SRSF6 promotes hyperplasia of sensitized skin. Nat. Struct. Mol. Biol. 21, 189–197 (2014).
Liang, X. et al. PTBP3 contributes to the metastasis of gastric cancer by mediating CAV1 alternative splicing. Cell Death Dis. 9, 569 (2018).
Ailiken, G. et al. Post-transcriptional regulation of BRG1 by FIRDeltaexon2 in gastric cancer. Oncogenesis 9, 26 (2020).
Lin, P. et al. Role of global aberrant alternative splicing events in papillary thyroid cancer prognosis. Aging (Albany NY) 11, 2082–2097 (2019).
Liu, J. et al. Aberrant expression of splicing factors in newly diagnosed acute myeloid leukemia. Onkologie 35, 335–340 (2012).
Kanagal-Shamanna, R. et al. Myeloid neoplasms with isolated isochromosome 17q demonstrate a high frequency of mutations in SETBP1, SRSF2, ASXL1 and NRAS. Oncotarget 7, 14251–14258 (2016).
Itzykson, R. et al. Prognostic score including gene mutations in chronic myelomonocytic leukemia. J. Clin. Oncol. 31, 2428–2436 (2013).
Wang, H. et al. Prognostic value of U2AF1 mutant in patients with de novo myelodysplastic syndromes: a meta-analysis. Ann. Hematol. 98, 2629–2639 (2019).
Mupo, A. et al. Hemopoietic-specific Sf3b1-K700E knock-in mice display the splicing defect seen in human MDS but develop anemia without ring sideroblasts. Leukemia 31, 720–727 (2017).
Schischlik, F. et al. Mutational landscape of the transcriptome offers putative targets for immunotherapy of myeloproliferative neoplasms. Blood 134, 199–210 (2019).
Malcovati, L. et al. SF3B1 mutation identifies a distinct subset of myelodysplastic syndrome with ring sideroblasts. Blood 126, 233–241 (2015).
Love, J. E., Hayden, E. J. & Rohn, T. T. Alternative splicing in Alzheimer’s disease. J Parkinsons Dis Alzheimers Dis. 2, 6–17 (2015).
Alieva, A. et al. Involvement of endocytosis and alternative splicing in the formation of the pathological process in the early stages of Parkinson’s disease. Biomed. Res. Int. 2014, 718732 (2014).
Shehadeh, L. A. et al. SRRM2, a potential blood biomarker revealing high alternative splicing in Parkinson’s disease. PLoS ONE 5, e9104 (2010).
Fernandez-Nogales, M. et al. Faulty splicing and cytoskeleton abnormalities in Huntington’s disease. Brain Pathol. 26, 772–778 (2016).
Hughes, T. et al. A loss-of-function variant in a minor isoform of ANK3 protects against bipolar disorder and schizophrenia. Biol. Psychiatry 80, 323–330 (2016).
Morikawa, T. et al. The expression of HMGA1a is increased in lymphoblastoid cell lines from schizophrenia patients. Neurochem. Int. 56, 736–739 (2010).
Ohno, K. et al. Splicing regulation and dysregulation of cholinergic genes expressed at the neuromuscular junction. J. Neurochem. 142, 64–72 (2017).
Singh, R. N., Seo, J. & Singh, N. N. RNA in spinal muscular atrophy: therapeutic implications of targeting. Expert Opin. Ther. Targets 24, 731–743 (2020).
Singh, R. N. & Singh, N. N. Mechanism of splicing regulation of spinal muscular atrophy genes. Adv. Neurobiol. 20, 31–61 (2018).
Serena, G. et al. Proinflammatory cytokine interferon-gamma and microbiome-derived metabolites dictate epigenetic switch between forkhead box protein 3 isoforms in coeliac disease. Clin. Exp. Immunol. 187, 490–506 (2017).
Zhou, H. et al. Alternative splicing directs two IL-20R2 isoforms and is responsible for the incomplete gene knockout via the exon I ablation. Genes Immun. 17, 220–227 (2016).
Dahal, L. N. et al. Immunoregulatory soluble CTLA-4 modifies effector T-cell responses in systemic lupus erythematosus. Arthritis Res. Ther. 18, 180 (2016).
Luzina, I. G. et al. Natural production and functional effects of alternatively spliced interleukin-4 protein in asthma. Cytokine 58, 20–26 (2012).
Gray, P. et al. Identification of a novel human MD-2 splice variant that negatively regulates Lipopolysaccharide-induced TLR4 signaling. J. Immunol. 184, 6359–6366 (2010).
Liu, S. et al. MITA/STING and Its alternative splicing isoform MRP restrict hepatitis B virus replication. PLoS ONE 12, e0169701 (2017).
Lara-Pezzi, E., Gomez-Salinero, J., Gatto, A. & Garcia-Pavia, P. The alternative heart: impact of alternative splicing in heart disease. J. Cardiovasc. Transl. Res. 6, 945–955 (2013).
Guo, W. et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat. Med. 18, 766–773 (2012).
Dlamini, Z., Mokoena, F. & Hull, R. Abnormalities in alternative splicing in diabetes: therapeutic targets. J. Mol. Endocrinol. 59, R93–R107 (2017).
Malakar, P. et al. Insulin receptor alternative splicing is regulated by insulin signaling and modulates beta cell survival. Sci. Rep. 6, 31222 (2016).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Oltean, S. & Bates, D. O. Hallmarks of alternative splicing in cancer. Oncogene 33, 5311–5318 (2014).
Belluti, S., Rigillo, G. & Imbriano, C. Transcription factors in cancer: when alternative splicing determines opposite cell fates. Cells 9, 760–787 (2020).
Groulx, J. F., Boudjadi, S. & Beaulieu, J. F. MYC regulates alpha6 integrin subunit expression and splicing under its pro-proliferative ITGA6A form in colorectal cancer cells. Cancers (Basel). 10, 42–56 (2018).
David, C. J. et al. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 463, 364–368 (2010).
Bechara, E. G. et al. RBM5, 6, and 10 differentially regulate NUMB alternative splicing to control cancer cell proliferation. Mol. Cell 52, 720–733 (2013).
Lin, J. C., Tsao, M. F. & Lin, Y. J. Differential impacts of alternative splicing networks on apoptosis. Int. J. Mol. Sci. 17, 2097–2115 (2016).
Kuan, C. Y., Roth, K. A., Flavell, R. A. & Rakic, P. Mechanisms of programmed cell death in the developing brain. Trends Neurosci. 23, 291–297 (2000).
Bielli, P., Bordi, M., Di Biasio, V. & Sette, C. Regulation of BCL-X splicing reveals a role for the polypyrimidine tract binding protein (PTBP1/hnRNP I) in alternative 5′ splice site selection. Nucleic Acids Res. 42, 12070–12081 (2014).
Gautrey, H. L. & Tyson-Capper, A. J. Regulation of Mcl-1 by SRSF1 and SRSF5 in cancer cells. PLoS ONE 7, e51497 (2012).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Tripathi, V., Shin, J. H., Stuelten, C. H. & Zhang, Y. E. TGF-beta-induced alternative splicing of TAK1 promotes EMT and drug resistance. Oncogene 38, 3185–3200 (2019).
Brown, R. L. et al. CD44 splice isoform switching in human and mouse epithelium is essential for epithelial-mesenchymal transition and breast cancer progression. J. Clin. Invest. 121, 1064–1074 (2011).
Xu, Y. et al. Cell type-restricted activity of hnRNPM promotes breast cancer metastasis via regulating alternative splicing. Genes Dev. 28, 1191–1203 (2014).
Harvey, S. E. et al. Coregulation of alternative splicing by hnRNPM and ESRP1 during EMT. RNA 24, 1326–1338 (2018).
Harper, S. J. & Bates, D. O. VEGF-A splicing: the key to anti-angiogenic therapeutics? Nat. Rev. Cancer 8, 880–887 (2008).
Wagner, K. D. et al. Altered VEGF splicing isoform balance in tumor endothelium involves activation of splicing factors Srpk1 and Srsf1 by the Wilms’ tumor suppressor Wt1. Cells 8, 41–58 (2019).
Hatcher, J. M. et al. SRPKIN-1: a covalent SRPK1/2 inhibitor that potently converts VEGF from pro-angiogenic to anti-angiogenic isoform. Cell Chem. Biol. 25, 460–470 e466 (2018).
Zhu, H., Carpenter, R. L., Han, W. & Lo, H. W. The GLI1 splice variant TGLI1 promotes glioblastoma angiogenesis and growth. Cancer Lett. 343, 51–61 (2014).
Jurica, M. S. et al. The allosteric regulation of pyruvate kinase by fructose-1,6-bisphosphate. Structure 6, 195–210 (1998).
Takenaka, M. et al. Isolation and characterization of the human pyruvate kinase M gene. Eur. J. Biochem. 198, 101–106 (1991).
Christofk, H. R. et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 452, 230–233 (2008).
Chen, M., Zhang, J. & Manley, J. L. Turning on a fuel switch of cancer: hnRNP proteins regulate alternative splicing of pyruvate kinase mRNA. Cancer Res. 70, 8977–8980 (2010).
Mazurek, S. Pyruvate kinase type M2: a key regulator of the metabolic budget system in tumor cells. Int. J. Biochem. Cell Biol. 43, 969–980 (2011).
Han, J. et al. Hypoxia is a key driver of alternative splicing in human breast cancer cells. Sci. Rep. 7, 4108 (2017).
Pajares, M. J. et al. Alternative splicing: an emerging topic in molecular and clinical oncology. Lancet Oncol. 8, 349–357 (2007).
Yoshida, K. & Ogawa, S. Splicing factor mutations and cancer. Wiley Interdiscip. Rev. RNA 5, 445–459 (2014).
Damm, F. et al. Mutations affecting mRNA splicing define distinct clinical phenotypes and correlate with patient outcome in myelodysplastic syndromes. Blood 119, 3211–3218 (2012).
Dvinge, H., Kim, E., Abdel-Wahab, O. & Bradley, R. K. RNA splicing factors as oncoproteins and tumor suppressors. Nat. Rev. Cancer 16, 413–430 (2016).
Visconte, V., M. O., Nakashima & H. J., Rogers. mutations in splicing factor genes in myeloid malignancies: significance and impact on clinical features. Cancers (Basel). 11, 1844–1857 (2019).
Fu, X. D. & Maniatis, T. Factor required for mammalian spliceosome assembly is localized to discrete regions in the nucleus. Nature 343, 437–441 (1990).
Chen, M. & Manley, J. L. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat. Rev. Mol. Cell Biol. 10, 741–754 (2009).
Makishima, H. et al. Mutations in the spliceosome machinery, a novel and ubiquitous pathway in leukemogenesis. Blood 119, 3203–3210 (2012).
Anczukow, O. & Krainer, A. R. Splicing-factor alterations in cancers. RNA 22, 1285–1301 (2016).
Arbab Jafari, P. et al. Prognostic significance of SRSF2 mutations in myelodysplastic syndromes and chronic myelomonocytic leukemia: a meta-analysis. Hematology 23, 778–784 (2018).
Liang, Y. et al. SRSF2 mutations drive oncogenesis by activating a global program of aberrant alternative splicing in hematopoietic cells. Leukemia 32, 2659–2671 (2018).
Zhang, J. et al. Disease-associated mutation in SRSF2 misregulates splicing by altering RNA-binding affinities. Proc. Natl Acad. Sci. USA 112, E4726–E4734 (2015).
Papaemmanuil, E. et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N. Engl. J. Med. 365, 1384–1395 (2011).
Wang, L. et al. Transcriptomic characterization of SF3B1 mutation reveals its pleiotropic effects in chronic lymphocytic leukemia. Cancer Cell 30, 750–763 (2016).
Wu, S., Romfo, C. M., Nilsen, T. W. & Green, M. R. Functional recognition of the 3′ splice site AG by the splicing factor U2AF35. Nature 402, 832–835 (1999).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
Shirai, C. L. et al. Mutant U2AF1 expression alters hematopoiesis and pre-mRNA splicing in vivo. Cancer Cell 27, 631–643 (2015).
Okeyo-Owuor, T. et al. U2AF1 mutations alter sequence specificity of pre-mRNA binding and splicing. Leukemia 29, 909–917 (2015).
Palangat, M. et al. The splicing factor U2AF1 contributes to cancer progression through a noncanonical role in translation regulation. Genes Dev. 33, 482–497 (2019).
Madan, V. et al. Aberrant splicing of U12-type introns is the hallmark of ZRSR2 mutant myelodysplastic syndrome. Nat. Commun. 6, 6042 (2015).
Taylor, J. & Lee, S. C. Mutations in spliceosome genes and therapeutic opportunities in myeloid malignancies. Genes Chromosomes Cancer 58, 889–902 (2019).
Quesada, V. et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat. Genet. 44, 47–52 (2011).
Cancer Genome Atlas Research, N. et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Walter, M. J. et al. Clonal diversity of recurrently mutated genes in myelodysplastic syndromes. Leukemia 27, 1275–1282 (2013).
Kurtovic-Kozaric, A. et al. PRPF8 defects cause missplicing in myeloid malignancies. Leukemia 29, 126–136 (2015).
Sureau, A. et al. SC35 autoregulates its expression by promoting splicing events that destabilize its mRNAs. EMBO J. 20, 1785–1796 (2001).
Sun, S. et al. SF2/ASF autoregulation involves multiple layers of post-transcriptional and translational control. Nat. Struct. Mol. Biol. 17, 306–312 (2010).
Xu, Y. et al. Post-translational modification control of RNA-binding protein hnRNPK function. Open Biol. 9, 180239 (2019).
Naarmann-de Vries, I. S. et al. Characterization of acute myeloid leukemia with del(9q) - Impact of the genes in the minimally deleted region. Leuk. Res. 76, 15–23 (2019).
Gallardo, M. et al. Uncovering the role of RNA-binding protein hnRNP K in B-cell lymphomas. J. Natl Cancer Inst. 112, 95–106 (2020).
Li, M. et al. Downregulation of HNRNPK in human cancer cells inhibits lung metastasis. Anim. Model Exp. Med. 2, 291–296 (2019).
Chen, Y. et al. Role of heterogeneous nuclear ribonucleoprotein K in tumor development. J. Cell Biochem. 120, 14296–14305 (2019).
Gallardo, M. et al. hnRNP K is a haploinsufficient tumor suppressor that regulates proliferation and differentiation programs in hematologic malignancies. Cancer Cell 28, 486–499 (2015).
Lin, J. C., Lin, C. Y., Tarn, W. Y. & Li, F. Y. Elevated SRPK1 lessens apoptosis in breast cancer cells through RBM4-regulated splicing events. RNA 20, 1621–1631 (2014).
Zhang, Y. P. et al. Down-regulated RBM5 inhibits bladder cancer cell apoptosis by initiating an miR-432-5p/beta-catenin feedback loop. FASEB J. 33, 10973–10985 (2019).
Wang, Q. et al. RNA-binding protein RBM6 as a tumor suppressor gene represses the growth and progression in laryngocarcinoma. Gene 697, 26–34 (2019).
Dou, X. Q. et al. miR-335 modulates Numb alternative splicing via targeting RBM10 in endometrial cancer. Kaohsiung J. Med. Sci. 36, 171–177 (2020).
Dou, X. Q. et al. Alternative splicing of VEGFA is regulated by RBM10 in endometrial cancer. Kaohsiung J. Med. Sci. 36, 13–19 (2020).
Sun, X. et al. Functional role of RBM10 in lung adenocarcinoma proliferation. Int. J. Oncol. 54, 467–478 (2019).
Jin, X. et al. RBM10 inhibits cell proliferation of lung adenocarcinoma via RAP1/AKT/CREB signalling pathway. J. Cell Mol. Med. 23, 3897–3904 (2019).
Zhou, Z. & Fu, X. D. Regulation of splicing by SR proteins and SR protein-specific kinases. Chromosoma 122, 191–207 (2013).
Hollander, D. et al. A network-based analysis of colon cancer splicing changes reveals a tumorigenesis-favoring regulatory pathway emanating from ELK1. Genome Res. 26, 541–553 (2016).
Choudhury, R. et al. The splicing activator DAZAP1 integrates splicing control into MEK/Erk-regulated cell proliferation and migration. Nat. Commun. 5, 3078 (2014).
Weg-Remers, S., Ponta, H., Herrlich, P. & Konig, H. Regulation of alternative pre-mRNA splicing by the ERK MAP-kinase pathway. EMBO J. 20, 4194–4203 (2001).
Valacca, C. et al. Sam68 regulates EMT through alternative splicing-activated nonsense-mediated mRNA decay of the SF2/ASF proto-oncogene. J. Cell Biol. 191, 87–99 (2010).
Shilo, A. et al. Splicing factor hnRNP A2 activates the Ras-MAPK-ERK pathway by controlling A-Raf splicing in hepatocellular carcinoma development. RNA 20, 505–515 (2014).
Shultz, J. C. et al. Alternative splicing of caspase 9 is modulated by the phosphoinositide 3-kinase/Akt pathway via phosphorylation of SRp30a. Cancer Res. 70, 9185–9196 (2010).
Zhou, Z. et al. The Akt-SRPK-SR axis constitutes a major pathway in transducing EGF signaling to regulate alternative splicing in the nucleus. Mol. Cell 47, 422–433 (2012).
Wang, F. et al. SPSB1-mediated HnRNP A1 ubiquitylation regulates alternative splicing and cell migration in EGF signaling. Cell Res. 27, 540–558 (2017).
Lee, G. et al. Post-transcriptional regulation of de novo lipogenesis by mTORC1-S6K1-SRPK2 signaling. Cell 171, 1545–1558 e1518 (2017).
Zhan, T., Rindtorff, N. & Boutros, M. Wnt signaling in cancer. Oncogene 36, 1461–1473 (2017).
Hernandez, F. et al. Glycogen synthase kinase-3 plays a crucial role in tau exon 10 splicing and intranuclear distribution of SC35. Implications for Alzheimer’s disease. J. Biol. Chem. 279, 3801–3806 (2004).
Heyd, F. & Lynch, K. W. Phosphorylation-dependent regulation of PSF by GSK3 controls CD45 alternative splicing. Mol. Cell 40, 126–137 (2010).
Goncalves, V., Matos, P. & Jordan, P. The beta-catenin/TCF4 pathway modifies alternative splicing through modulation of SRp20 expression. RNA 14, 2538–2549 (2008).
Goncalves, V., Matos, P., Jordan, P. & Antagonistic, S. R. proteins regulate alternative splicing of tumor-related Rac1b downstream of the PI3-kinase and Wnt pathways. Hum. Mol. Genet. 18, 3696–3707 (2009).
Chen, L. L. The biogenesis and emerging roles of circular RNAs. Nat. Rev. Mol. Cell Biol. 17, 205–211 (2016).
Ransohoff, J. D., Wei, Y. & Khavari, P. A. The functions and unique features of long intergenic non-coding RNA. Nat. Rev. Mol. Cell Biol. 19, 143–157 (2018).
Romero-Barrios, N. et al. Splicing regulation by long noncoding RNAs. Nucleic Acids Res. 46, 2169–2184 (2018).
Gawronski, A. R. et al. MechRNA: prediction of lncRNA mechanisms from RNA-RNA and RNA-protein interactions. Bioinformatics 34, 3101–3110 (2018).
Grelet, S. et al. A regulated PNUTS mRNA to lncRNA splice switch mediates EMT and tumour progression. Nat. Cell Biol. 19, 1105–1115 (2017).
Papaioannou, D. et al. The long non-coding RNA HOXB-AS3 regulates ribosomal RNA transcription in NPM1-mutated acute myeloid leukemia. Nat. Commun. 10, 5351 (2019).
Gordon, M. A. et al. The long non-coding RNA MALAT1 promotes ovarian cancer progression by regulating RBFOX2-mediated alternative splicing. Mol. Carcinog. 58, 196–205 (2019).
Ji, Q. et al. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br. J. Cancer 111, 736–748 (2014).
Kong, J. et al. Long non-coding RNA LINC01133 inhibits epithelial-mesenchymal transition and metastasis in colorectal cancer by interacting with SRSF6. Cancer Lett. 380, 476–484 (2016).
Liu, M. et al. HOTAIR, a long noncoding RNA, is a marker of abnormal cell cycle regulation in lung cancer. Cancer Sci. 109, 2717–2733 (2018).
Cooper, D. R. et al. Long non-coding RNA NEAT1 associates with SRp40 to temporally regulate PPARgamma2 splicing during adipogenesis in 3T3-L1 cells. Genes (Basel) 5, 1050–1063 (2014).
Rapicavoli, N. A., Poth, E. M. & Blackshaw, S. The long noncoding RNA RNCR2 directs mouse retinal cell specification. BMC Dev. Biol. 10, 49 (2010).
Wu, H. et al. Unusual processing generates SPA LncRNAs that sequester multiple RNA binding proteins. Mol. Cell 64, 534–548 (2016).
Massone, S. et al. 17A, a novel non-coding RNA, regulates GABA B alternative splicing and signaling in response to inflammatory stimuli and in Alzheimer disease. Neurobiol. Dis. 41, 308–317 (2011).
Zhang, X. O. et al. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 26, 1277–1287 (2016).
Feng, J. et al. Genome-wide identification of cancer-specific alternative splicing in circRNA. Mol. Cancer 18, 35 (2019).
Kristensen, L. S., Hansen, T. B., Veno, M. T. & Kjems, J. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene 37, 555–565 (2018).
Zhang, Y. et al. Circular intronic long noncoding RNAs. Mol. Cell 51, 792–806 (2013).
Li, Z. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 22, 256–264 (2015).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 56, 55–66 (2014).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Du, W. W. et al. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 44, 2846–2858 (2016).
Yang, Y. et al. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 27, 626–641 (2017).
Wang, Y. & Wang, Z. Efficient backsplicing produces translatable circular mRNAs. RNA 21, 172–179 (2015).
Meng, S. et al. CircRNA: functions and properties of a novel potential biomarker for cancer. Mol. Cancer 16, 94 (2017).
de Fraipont, F., Gazzeri, S., Cho, W. C. & Eymin, B. Circular RNAs and RNA splice variants as biomarkers for prognosis and therapeutic response in the liquid biopsies of lung cancer patients. Front. Genet. 10, 390 (2019).
Shuai, S. et al. The U1 spliceosomal RNA is recurrently mutated in multiple cancers. Nature 574, 712–716 (2019).
Suzuki, H. et al. Recurrent noncoding U1 snRNA mutations drive cryptic splicing in SHH medulloblastoma. Nature 574, 707–711 (2019).
Inoue, D. et al. Spliceosomal disruption of the non-canonical BAF complex in cancer. Nature 574, 432–436 (2019).
Huang, H. H. et al. Proteasome inhibitor-induced modulation reveals the spliceosome as a specific therapeutic vulnerability in multiple myeloma. Nat. Commun. 11, 1931 (2020).
Kahles, A. et al. Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell 34, 211–224 e216 (2018).
da Silva, M. R. et al. Splicing regulators and their roles in cancer biology and therapy. Biomed. Res. Int. 2015, 150514 (2015).
Kim, Y. J. & Kim, H. S. Alternative splicing and its impact as a cancer diagnostic marker. Genomics Inf. 10, 74–80 (2012).
Bonnal, S., Vigevani, L. & Valcarcel, J. The spliceosome as a target of novel antitumour drugs. Nat. Rev. Drug Discov. 11, 847–859 (2012).
Effenberger, K. A., Urabe, V. K. & Jurica, M. S. Modulating splicing with small molecular inhibitors of the spliceosome. Wiley Interdiscip. Rev. RNA. 8, 1381–1399 (2017).
Ohe, K. & Hagiwara, M. Modulation of alternative splicing with chemical compounds in new therapeutics for human diseases. ACS Chem. Biol. 10, 914–924 (2015).
Swift, P. A. & MacGregor, G. A. The epithelial sodium channel in hypertension. Am. J. Pharmacogenomics 4, 161–168 (2004).
Chang, J. G. et al. Small molecule amiloride modulates oncogenic RNA alternative splicing to devitalize human cancer cells. PLoS ONE 6, e18643 (2011).
Seiler, M. et al. H3B-8800, an orally available small-molecule splicing modulator, induces lethality in spliceosome-mutant cancers. Nat. Med. 24, 497–504 (2018).
Bauman, J., Jearawiriyapaisarn, N. & Kole, R. Therapeutic potential of splice-switching oligonucleotides. Oligonucleotides 19, 1–13 (2009).
Touznik, A. et al. LNA/DNA mixmer-based antisense oligonucleotides correct alternative splicing of the SMN2 gene and restore SMN protein expression in type 1 SMA fibroblasts. Sci. Rep. 7, 3672 (2017).
Cirak, S. et al. Exon skipping and dystrophin restoration in patients with Duchenne muscular dystrophy after systemic phosphorodiamidate morpholino oligomer treatment: an open-label, phase 2, dose-escalation study. Lancet 378, 595–605 (2011).
Chen, T. H. New and developing therapies in spinal muscular atrophy: from genotype to phenotype to treatment and where do we stand? Int. J. Mol. Sci. 21, 3297–3316 (2020).
Wurster, C. D. & Ludolph, A. C. Antisense oligonucleotides in neurological disorders. Ther. Adv. Neurol. Disord. 11, 1756286418776932 (2018).
Osman, E. Y. et al. Morpholino antisense oligonucleotides targeting intronic repressor Element1 improve phenotype in SMA mouse models. Hum. Mol. Genet. 23, 4832–4845 (2014).
Li, Z. et al. Pro-apoptotic effects of splice-switching oligonucleotides targeting Bcl-x pre-mRNA in human glioma cell lines. Oncol. Rep. 35, 1013–1019 (2016).
Bauman, J. A. et al. Anti-tumor activity of splice-switching oligonucleotides. Nucleic Acids Res. 38, 8348–8356 (2010).
Denichenko, P. et al. Specific inhibition of splicing factor activity by decoy RNA oligonucleotides. Nat. Commun. 10, 1590 (2019).
Slansky, J. E. & Spellman, P. T. Alternative splicing in tumors-a path to immunogenicity? N. Engl. J. Med. 380, 877–880 (2019).
Volpe, G. et al. Alternative BCR/ABL splice variants in Philadelphia chromosome-positive leukemias result in novel tumor-specific fusion proteins that may represent potential targets for immunotherapy approaches. Cancer Res. 67, 5300–5307 (2007).
Vauchy, C. et al. CD20 alternative splicing isoform generates immunogenic CD4 helper T epitopes. Int. J. Cancer 137, 116–126 (2015).
Frankiw, L., Baltimore, D. & Li, G. Alternative mRNA splicing in cancer immunotherapy. Nat. Rev. Immunol. 19, 675–687 (2019).
Smith, C. C. et al. Alternative tumour-specific antigens. Nat. Rev. Cancer 19, 465–478 (2019).
Coutinho, M. F., Matos, L., Santos, J. I. & Alves, S. RNA therapeutics: how far have we gone? Adv. Exp. Med. Biol. 1157, 133–177 (2019).
Wang, H., La Russa, M. & Qi, L. S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Setten, R. L., Rossi, J. J. & Han, S. P. The current state and future directions of RNAi-based therapeutics. Nat. Rev. Drug Discov. 18, 421–446 (2019).
Komor, A. C. et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Abudayyeh, O. O. et al. RNA targeting with CRISPR-Cas13. Nature 550, 280–284 (2017).
Granados-Riveron, J. T. & Aquino-Jarquin, G. CRISPR-Cas13 precision transcriptome engineering in cancer. Cancer Res. 78, 4107–4113 (2018).
Qian, H. et al. Reversing a model of Parkinson’s disease with in situ converted nigral neurons. Nature 582, 550–556 (2020).
Zhou, H. et al. Glia-to-neuron conversion by CRISPR-CasRx alleviates symptoms of neurological disease in mice. Cell 181, 590–603 e516 (2020).
Acknowledgements
This work was supported by National Key Research and Development Program: Precision Medicine sub-program 2016YFC0905900 (to YY); National Natural Science Foundation of China 81970196 and 81670200 (to C.G. & Y.Y.); A Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (Integration of Chinese and Western Medicine); Innovation Team of Six Talent Peaks Project in Jiangsu Province (TD-SWYY-015); Natural Science Foundation of Jiangsu Province for Excellent Young Scholars.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Qian, J., Gu, C. et al. Alternative splicing and cancer: a systematic review. Sig Transduct Target Ther 6, 78 (2021). https://doi.org/10.1038/s41392-021-00486-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-021-00486-7
This article is cited by
-
MAGOH promotes gastric cancer progression via hnRNPA1 expression inhibition-mediated RONΔ160/PI3K/AKT signaling pathway activation
Journal of Experimental & Clinical Cancer Research (2024)
-
SIRT7: a novel molecular target for personalized cancer treatment?
Oncogene (2024)
-
Alternative splicing and related RNA binding proteins in human health and disease
Signal Transduction and Targeted Therapy (2024)
-
Modification of BCLX pre-mRNA splicing has antitumor efficacy alone or in combination with radiotherapy in human glioblastoma cells
Cell Death & Disease (2024)
-
The pleiotropic nature of NONO, a master regulator of essential biological pathways in cancers
Cancer Gene Therapy (2024)