Abstract
Mucosal melanomas are rare, and less is known about the biomarkers of this subtype in comparison to cutaneous or uveal melanomas. Preferentially expressed antigen in melanoma (PRAME) has been studied as a tool for prognostication of uveal melanomas, and immunotherapy against PRAME-expressing tumor cells has already shown promise. Our goal was to retrospectively analyze 29 cases of mucosal melanomas at our institution to determine if any molecular and histopathologic prognosticators could be identified, as well as to study PRAME expression and its association with prognosis. We found that the majority of mucosal melanomas expressed PRAME and a high PRAME expression score predicted a poor prognosis. There was no association between prognosis and the histomorphologic features analyzed, such as presence of spindle cell or epithelioid predominance. BRAF mutations were absent in 16 of 16 cases tested. Pathogenic NRAS mutations were detected in 3 of 11 cases tested and were associated with shorter overall survival compared to those without NRAS alterations, but the presence of NRAS mutations did not correlate with PRAME expression. In conclusion, an increase in PRAME expression and the presence of a pathogenic NRAS were both associated with a worse prognosis in mucosal melanomas.
Similar content being viewed by others
Introduction
Mucosal melanoma is a rare cancer that arises from the melanocytes present within any mucosal lining. The most common primary site is the sinonasal tract; however, other possible primary sites include the gastrointestinal, oropharyngeal, and genitourinary tracts. Owing to a delayed clinical presentation especially in areas not easily visualized, this subtype of melanoma is typically already invasive when discovered, thus conferring a poor prognosis. Compared to cutaneous melanomas, research on mucosal melanomas is not as comprehensive primarily due to the sheer rarity of the diagnosis, and there is still much to learn about the risk factors, pathogenesis, prognosis, and treatment of this entity. Although immunohistochemical staining patterns are similar to that of cutaneous melanoma, recent studies have shown that the molecular profiles are notably different, making them eligible for different targeted molecular therapies [1]. Whereas BRAF mutations have been implicated in the majority of cutaneous melanomas, they are rare in mucosal melanomas, and instead the molecular characteristics of mucosal melanomas are more similar to uveal melanomas and lack the molecular signature of ultraviolet damage [1].
Recently, PRAME (preferentially expressed antigen in melanoma), which is a tumor-associated antigen in the family of cancer testis antigens (CTA), has been of interest in melanomas [2,3,4]. In cutaneous melanomas, Lezcano et al. [2] found PRAME immunohistochemical staining to have utility in supporting a diagnosis of melanoma, as well as when there are challenges in the assessment of melanoma margins. In uveal melanomas, PRAME mRNA expression has been found to be an important and independent prognostic indicator for metastatic risk [3]. Furthermore, a study has already shown promising results with PRAME-specific T cells recognizing uveal melanoma cells positive for PRAME expression [4]. Immunotherapeutic treatment using anti-PRAME antibodies against metastatic cutaneous melanoma has already undergone Phase I study, which demonstrated an acceptable safety profile and thus was cleared for Phase II testing [5].
These promising findings led us to explore the expression of PRAME in mucosal melanomas, as well as its association with prognosis. Additionally, we retrospectively analyzed the histopathologic and molecular characteristics of cases at our institution in hopes of elucidating any other prognostic indicators to better understand the pathology of mucosal melanomas.
Materials and methods
Following IRB approval, our institution’s database was searched for cases diagnosed as primary mucosal melanoma from 2003 to 2018. Twenty-nine mucosal melanoma cases of various sites from 29 different patients were identified, and we reviewed all available medical records as well as diagnostic and subsequent pathology reports for each case. Physical slides were available to us for 24 of these cases, and all were retrospectively reviewed by a resident (AT) with a staff head and neck pathologist (FL) to assess their histopathologic features.
Clinical data such as age at diagnosis, months survived, and primary site of malignancy were analyzed with the histopathologic features, immunohistochemical staining, and molecular profile of the tumor. Sixteen total cases had clinical molecular testing for review (Supplemental Table 1). Four cases were tested only for BRAF V600E using a CLIA-validated allele-specific real-time PCR assay. Twelve cases had CLIA-validated clinical next-generation sequencing data for focused gene panels [6, 7]. Two cases had Sanger sequencing results available performed at an outside hospital.
For the same 24 cases for which slides were available for review, blocks were available for staining. Formalin-fixed, paraffin-embedded tissues were de-paraffinized and rehydrated using standard methods. Antigens were retrieved (Reveal Decloaking reagent; Biocare Medical, Concord, CA), and subsequent steps were automated using an immunohistochemical staining platform (IntelliPath; Biocare). After quenching the endogenous peroxidase activity (Peroxidazed; Biocare), a serum-free blocking solution (Sniper; Biocare Medical, Concord, CA) was used, then removed, and the slides were incubated with rabbit monoclonal anti-PRAME (clone EPR20330; Abcam, Cambridge, MA, 1:1000) for 60 min at room temperature followed by TBST rinse and detection with Novocastra Novolink Polymer Kit (Leica Microsystems Inc., Buffalo Grove, IL) using the manufacturer’s specifications [2]. Slides were then rinsed with TBST and detected with diaminobenzidine (DAB) (Biolegent, Dedham, MA), followed by counterstain with CAT hematoxylin (Biocare, Concord, CA), dehydration, and then coverslipping. Testicular tissue served as both positive and negative controls, with high positivity in spermatogonia and negative staining in Sertoli cells of the same tissue. This tissue was also used to determine the best titer, using serial dilutions of the primary and secondary antibodies. All stained slides were reviewed by a resident (AT) and a fellow (MN), both of whom independently and qualitatively scored PRAME expression using percentages of 3+ (strong), 2+ (moderate), 1+ (weak), and 0 (negative) staining of malignant melanocytes for each case. The overall PRAME expression score was derived from the sum of each of the score values multiplied by the percentage of positive tumor cells (3 × x% + 2 × x% + 1 × x% = total score) to equal a range of 0–300, similar to the scoring system utilized for estrogen and progesterone receptor evaluation in breast carcinomas [8, 9]. The mean of the resident and fellow’s scores were calculated and used for analysis. For any cases with discrepancies of a score >50 between the two scores, the slides were instead reviewed and scored together with a staff head and neck pathologist (FL). PRAME scores were then compared against morphologic, molecular, and clinical characteristics of the corresponding tumor using an online unpaired t-test calculator [10]. To assess for any differences of PRAME scores among the primary tumor sites as well as between sinonasal vs. non-sinonasal sites, ANOVA tests were performed using R v.3.4.1 [11].
Three out of the 24 patients in our study had incomplete clinical histories and months of survival were unknown, and thus these patients were excluded in our analyses of survival and prognosis. Cox proportional hazards models were used to assess the association between PRAME score and survival in three ways: univariately, adjusted for age, and adjusted for the “stage” of disease. In an effort to include data from the gastrointestinal, genitourinary, and the oropharyngeal sites, as these sites do not have specific AJCC staging systems for mucosal melanoma, we adjusted for the extent of disease instead of stage, grouped into either local disease, metastatic to lymph node, or metastatic to distant site (Supplemental Table 2). These models were fit using R v.3.4.1, and scaled Schoenfeld residuals were used to check for clear violations of the proportional hazards assumption [11,12,13]. We generated Kaplan–Meier curves comparing survival between cases with PRAME scores above and below the median using Excel, as well as comparing cases positive and negative for NRAS. A log-rank test was used to compare the two survival curves for both variables. A separate log-rank test was used to determine any differences in survival among females and males. In all analyses, a two-sided p value of <0.05 was considered to be statistically significant.
Results
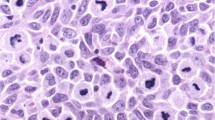
Clinicopathologic characteristics of our cases of mucosal melanoma are highlighted in Table 1. The mean age at diagnosis among all 29 cases was 67.1 years. At 12 months (1 year) after diagnosis, 72.0% of patients were alive, and this number decreased to 38.5% at 24 months (2 years). The sinonasal tract was the most common primary site in our population, at 65.5% of cases, followed by the gastrointestinal tract at 24.1%, the genitourinary tract at 13.8%, and the oropharyngeal tract at 3.4%. The most common cell morphology seen was epithelioid (75%), followed by spindle cells (33.3%) (Fig. 1). The immunohistochemical stains that were most commonly performed by staff pathologists to support the diagnosis and to exclude other diagnoses were S100, HMB45, tyrosinase, Melan-A, AE1/AE3, CD45, and chromogranin. AE1/AE3 (21 cases), CD45 (11 cases), and chromogranin (8 cases) were negative for all cases tested. All 28 cases (100%) stained for S100 were positive, with 78.6% (22/28 cases) showing strong, diffusely positive staining and 21.4% (6/28 cases) showing focal, scattered, weak, or patchy staining. HMB45, tyrosinase, and Melan-A were not as sensitive, with 88%, 82.8%, and 87.5% of cases staining positive, respectively.
H&E. a Low power view of epithelioid morphology, ×10 magnification. b High-power view of epithelioid morphology with pleomorphic cells, ×40 magnification. c Rhabdoid morphology, ×40 magnification. d Plasmacytoid morphology, ×40 magnification. e Low-power view of spindle cell morphology, ×10 magnification. f High-power view of spindle cell morphology, ×40 magnification
The most common pathogenic mutation found among our cases was within the NRAS gene. Three of 11 cases tested (27.3%) had pathogenic NRAS mutations, (p.Q61K, p.G12C, and p.G12D). All 16 cases tested (100%) were negative for activating BRAF alterations. Two variants of uncertain significance were identified in KIT (p.K492R) and HRAS (p.Q70*) as passenger mutations in a case where an NRAS mutation was also present.
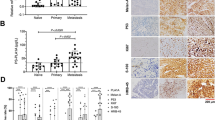
PRAME immunohistochemical staining revealed only four cases (4/24, 16.7%) that had negative or weak staining (Fig. 2). These cases had 70% or more tumor cells stain negative for PRAME, corresponding to an overall score of ≤30. The remaining 20 cases (83.3%) all had a higher overall score, with 45% as the highest percentage of PRAME-negative cells in the tumor among this group. PRAME scores were analyzed against morphologic features, molecular studies, and survival at 1 and 2 years (Table 2). Although the mean PRAME score was lower in cases with predominant spindle cells, this difference in PRAME scores across different cell types did not reach statistical significance (p = 0.054). Mean PRAME scores did not differ significantly between NRAS mutated and non-mutated cases (p = 0.237). The mean PRAME staining score was significantly lower in patients who survived 2 years after diagnosis compared to patients who did not survive 2 years (p = 0.012), but the same was not observed for 1-year survival (p = 0.061). Using a Cox proportional hazards model, we found that a 100-point increase in PRAME score was associated with a 170% increase in the hazard of death (HR: 2.70, 95% CI: [1.11, 6.59], p = 0.029). Kaplan–Meier curves for patients with PRAME scores above and below the median (202.5) are presented in Fig. 3a. There was little change in the estimated hazard ratio for PRAME when adding age as a covariate in the model. When the model was then adjusted for extent of disease, the estimated hazard ratio decreased only slightly (HR: 2.53, 95% CI: [1.04, 6.15], p = 0.041).
PRAME staining patterns, ×20 magnification. Left column demonstrates epithelioid predominant tumors exhibiting strong, mostly 3+ PRAME staining (a); moderate, mostly 1–2+ staining (c); and negative staining (e). Right column demonstrates spindle cell-predominant tumors with strong, mostly 3+ PRAME staining (b); moderate, mostly 1–2+ staining (d); and negative staining (f)
There was no significant difference between the PRAME scores among sinonasal, gastrointestinal, genitourinary, and oropharyngeal sites (p = 0.365). Furthermore, we found no significant differences when comparing PRAME scores among sinonasal vs. non-sinonasal sites (p = 0.347).
Kaplan–Meier curves and the log-rank test demonstrated that activating, clinically significant NRAS mutations conferred a lower survivability (p = 0.004) when compared to those that were negative for NRAS mutations (Fig. 3b). Due to our small sample size, we were unable to include both variables as covariates in a single Cox proportional hazards model, which would have allowed us to directly test whether PRAME expression and NRAS status have independent prognostic value.
Discussion
Little is still known on the topic of mucosal melanomas due to the sheer sparsity of cases as compared to other types of primary melanomas. Pathologic staging for these tumors is currently established by the American Joint Committee on Cancer (AJCC) for primary mucosal melanomas of the head and neck; however, none has been established for other primary sites [14]. Surgery and subsequent radiation therapy for a subset of these cases is the mainstay for treatment; however, the prognosis of this cancer continues to remain dismal, with an estimated 25% 5-year survival as compared to 80.8% for cutaneous melanomas [15,16,17]. For this reason, we sought to help shed light on mucosal melanomas by analyzing clinicopathologic and molecular data as well as PRAME immunohistochemical staining patterns for cases diagnosed in our institution to determine any associations with prognosis.
PRAME immunohistochemical staining patterns nor mRNA expression have yet to be studied in mucosal melanomas. We hypothesized that due to mucosal melanomas perhaps being more molecularly similar to uveal melanomas than cutaneous melanomas, PRAME expression may act as a prognostic indicator for mucosal melanomas as was seen in the uveal type [1, 3].
We observed a trend toward higher PRAME scores for epithelioid-cell-predominant tumors as compared to spindle cell-predominant tumors, as well as for the presence of NRAS mutations relative to the absence of NRAS mutations (Table 2). However, these differences were not statistically significant. No significant difference was seen in PRAME scores among the different primary sites nor between primary sinonasal and non-sinonasal cases. Notably, we found that patients who were alive at least 2 years after diagnosis have a lower average PRAME score than those who died prior to 2 years, which first suggested to us that PRAME may indeed be a prognostic indicator for mucosal melanomas.
There was a significant association between PRAME score and hazard of death, based on the Cox proportional hazards model, which suggests that a high PRAME-positive tumor cell burden portends a worse prognosis in mucosal melanomas. This association persisted even after taking into account the patients’ age and extent of disease at diagnosis. Analyzing PRAME as a continuous variable allowed us greater power to detect an association with survival while being limited by a small sample size. We also believe that a difference in power may explain why a significant difference in PRAME score was observed between 2-year survivors and non-survivors, but not when analyzing survival at the 1-year mark. Our finding is similar to the aforementioned discovery of PRAME expression as a prognostic indicator in uveal melanomas, and our study offers additional evidence that suggests more of a commonality between mucosal and uveal melanomas as compared to cutaneous melanomas [1, 3].
The average age among our cases was similar to that reported in the literature, with a majority of patients being over 60 years old [16, 18]. Also similar to the literature, the sinonasal tract was the most common primary site [16, 18]. Histopathologically, epithelioid-cell-predominant tumors were more common than cases with spindle cell predominance. A prior study with 61 cases of sinonasal mucosal melanoma showed no correlation of histologic subtypes with prognosis and survival [19]. We were also unable to identify any statistically significant correlations of histological subtypes with prognosis.
Among the immunohistochemical stains performed, S100 was the most sensitive, with positivity seen in 100% of our cases. The high sensitivity of S100 in the detection of melanocytic tumors ranging from 93% to 100% sensitivity have been well established [20]. There does not appear to be any studies showing an increase in negative S100 staining in mucosal melanomas as is the case for uveal melanomas [21].
It is well known that BRAF mutations are frequently seen in cutaneous melanomas; however, these are infrequent in mucosal melanomas [1, 22, 23]. This was observed in our cases as well, as all were negative for BRAF mutations. A whole-genome sequencing study recently revealed KIT mutations to be more frequent in mucosal melanomas, at 25% of cases (2 of 8) as compared to the 4.3% of cases (6 of 140) of cutaneous melanomas [1]. Among our cases, however, no pathogenic KIT mutations were identified in 14 cases assessed by either clinical NGS or Sanger sequencing. The same study also identified the SF3B1 gene, which was known to be more commonly mutated in uveal melanomas, to be significantly mutated in mucosal melanomas and thus perhaps a subject of future study, but unfortunately the SF3B1 gene was not a part of the clinical gene panel at our institution during the study period [1, 24].
The prognostic significance of an NRAS mutation was recently studied in cutaneous melanomas by Heppt et al. [25], who observed tumors with NRAS mutations exhibit more aggressive behavior. Similarly, we observed that cases with pathogenic NRAS mutations had shorter overall survival as compared to cases where no mutation was present, suggesting that the presence of a pathogenic NRAS mutation also portends a poor prognosis in mucosal melanomas. However, our data are in contrast to the study by Amit et al. [23] who observed that NRAS mutations did not significantly affect survival in sinonasal mucosal melanomas, although the prevalence of NRAS mutations in their cases (30%, 22/66 cases) was similar to our present study. The review of the current literature and the molecular findings in our study led us to suggest that although BRAF mutation status may be important to determine eligibility for possible targeted therapy options, broadening the search to include pathogenic mutations in other genes such as NRAS is more appropriate in this clinical setting.
Despite being limited by the small number of mucosal melanoma cases at our institution, we were able to provide insight on a novel marker for assessment of prognosis in mucosal melanomas. We hope that our findings will motivate larger studies and will be useful for future meta-analyses. With the majority of mucosal melanomas in our study staining for PRAME, specifically targeting tumor cells with PRAME expression may be a strategy for future therapy of this devastating cancer, just as it has shown potential in cutaneous and uveal melanomas [4, 5].
References
Hayward NK, Wilmott JS, Waddell N, Johansson PA, Field MA, Nones K, et al. Whole-genome landscapes of major melanoma subtypes. Nature. 2017;545:175–80.
Lezcano C, Jungbluth AA, Nehal KS, Hollmann TJ, Busam KJ. PRAME expression in melanocytic tumors. Am J Surg Pathol. 2018;42:1456–65.
Field MG, Decatur CL, Kurtenbach S, Gezgin G, van der Velden PA, Jager MJ, et al. PRAME as an independent biomarker for metastasis in uveal melanoma. Clin Cancer Res. 2016;22:1234–42.
Gezgin G, Luk SJ, Cao J, Dogrusöz M, van der Steen DM, Hagedoorn RS, et al. PRAME as a potential target for immunotherapy in metastatic uveal melanoma. JAMA Ophthalmol. 2017;135:541–9.
Gutzmer R, Rivoltini L, Levchenko E, Testori A, Utikal J, Ascierto PA, et al. Safety and immunogenicity of the PRAME cancer immunotherapeutic in metastatic melanoma: results of a Phase I dose escalation study. ESMO Open. 2016;1:e000068.
Yang R, Nelson AC, Henzler C, Thyagarajan B, Silverstein KAT. ScanIndel: a hybrid framework for indel detection via gapped alignment, split reads and de novo assembly. Genome Med. 2015;7:127.
Henzler C, Schomaker M, Yang R, Lambert AP, LaRue R, Kincaid R, et al. Optimization of a microfluidics-based next generation sequencing assay for clinical oncology diagnostics. Ann Transl Med. 2018;6:162.
McCarty KS Jr, Miller LS, Cox EB, Konrath J, McCarty KS Sr. Estrogen receptor analyses: correlation of biochemical and immunohistochemical methods using monoclonal antireceptor antibodies. Arch Pathol Lab Med. 1985;109:706–21.
Fitzgibbons PL, Dillon DA, Asalbeh R, Berman MA, Hayes DF, Hicks DG, et al. Template for reporting results of biomarker testing of specimens from patients with carcinoma of the breast. Arch Pathol Lab Med. 2014;138:595–601.
GraphPad QuickCalcs: t test calculator. https://www.graphpad.com/quickcalcs/ttest1.cfm. Accessed 26 February 2019.
R core team. R: A language and environment for statistical computing. Vol. 1. Vienna, Austria: R foundation for Statistical Computing; 2017. https://www.R-project.org/.
Therneau T. A package for survival analysis in S version 2.38, 2015. https://CRAN.R-project.org/package=survival.
Therneau TM, Grambsch PM. Modeling survival data: extending the Cox model. New York: Springer; 2000.
Amin MBES, Greene F, Byrd DR, Brookland RK, Washington MK, Gershenwald JE, et al. (eds.). AJCC cancer staging manual. 8th ed. Switzerland: Springer International Publishing; 2017.
Pfister DG, Ang KK, Brizel DM, Burtness B, Cmelak AJ, Colevas AD, et al. Mucosal melanoma of the head and neck. J Natl Compr Canc Netw. 2012;10:320–38.
Mihajlovic M, Vlajkovic S, Jovanovic P, Stefanovic V. Primary mucosal melanomas: a comprehensive review. Int J Clin Exp Pathol. 2012;5:739–53.
Chang AE, Karnell LH, Menck HR. The National Cancer Data Base report on cutaneous and noncutaneous melanoma: a summary of 84,836 cases from the past decade. The American College of Surgeons Commission on Cancer and the American Cancer Society. Cancer. 1998;83:1664–78.
Heppt MV, Roesch A, Weide B, et al. Prognostic factors and treatment outcomes in 444 patients with mucosal melanoma. Eur J Cancer. 2017;81:36–44.
Dauer EH, Lewis JE, Rohlinger AL, Weaver AL, Olsen KD. Sinonasal melanoma: a clinicopathologic review of 61 cases. Otolaryngol Head Neck Surg. 2008;138:347–52.
Ordóñez NG. Value of melanocytic-associated immunohistochemical markers in the diagnosis of malignant melanoma: a review and update. Hum Pathol. 2014;45:191–205.
Iwamoto S, Burrows RC, Kalina RE, George D, Boehm M, Bothwell MA, et al. Immunophenotypic differences between uveal and cutaneous melanomas. Arch Ophthalmol. 2002;120:466–70.
Zebary A, Jangard M, Omholt K, Ragnarsson-Olding B, Hansson J. KIT, NRAS and BRAF mutations in sinonasal mucosal melanoma: a study of 56 cases. Br J Cancer. 2013;109:559–69.
Amit M, Tam S, Abdelmeguid AS, Roberts DB, Takahashi Y, Raza SM, et al. Mutation status among patients with sinonasal mucosal melanoma and its impact on survival. Br J Cancer. 2017;116:1564–71.
Harbour JW, Roberson ED, Anbunathan H, Onken MD, Worley LA, Bowcock AM. Recurrent mutations at codon 625 of the splicing factor SF3B1 in uveal melanoma. Nat Genet. 2013;45:133–5.
Heppt MV, Siepmann T, Engel J, Schubert-Fritschle G, Eckel R, Mirlach L, et al. Prognostic significance of BRAF and NRAS mutations in melanoma: a German study from routine care. BMC Cancer. 2017;17:536.
Acknowledgements
This research received histology assistance from the University of Minnesota’s Biorepository and Laboratory Services program and was supported by the National Institutes of Health’s National Center for Advancing Translational Sciences, grant UL1TR002494. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health’s National Center for Advancing Translational Sciences. ACN is supported by the American Cancer Society, Clinical Scientist Development Grant, CSDG-18-139-01-CSM.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41379_2019_335_MOESM1_ESM.pdf
1. Molecular variants tested and methodologies used. 2. Primary site of mucosal melanoma as compared to PRAME scores, months survived, and extent of disease
Rights and permissions
About this article
Cite this article
Toyama, A., Siegel, L., Nelson, A.C. et al. Analyses of molecular and histopathologic features and expression of PRAME by immunohistochemistry in mucosal melanomas. Mod Pathol 32, 1727–1733 (2019). https://doi.org/10.1038/s41379-019-0335-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41379-019-0335-4
This article is cited by
-
Refining the application of PRAME—a useful marker in high CSD and acral melanoma subtypes
Virchows Archiv (2023)
-
PRAME Staining in Sinonasal Mucosal Melanoma: A Single-Center Experience
Head and Neck Pathology (2022)
-
Diagnostic utility of immunohistochemistry in concordance with mRNA analysis of PRAME in the stratification of high-risk uveal melanoma patients
Human Cell (2022)