Abstract
Non-alcoholic fatty liver disease (NAFLD) is the most prevalent chronic liver disease, characterized by excessive hepatic lipid accumulation. Recently, we demonstrated that Smad ubiquitination regulatory factor 1 (Smurf1) deficiency significantly alleviates mouse hepatic steatosis. However, the mechanism of Smurf1-regulating hepatic lipid accumulation requires further exploration and clarification. Hence, this study explores the potential mechanism of Smurf1 in hepatic steatosis. In this study, hepatic Smurf1 proteins in NAFLD patients and healthy individuals were determined using immunohistochemical staining. Control and NAFLD mouse models were established by feeding Smurf1-knockout (KO) and wild-type mice with either a high-fat diet (HFD) or a chow diet (CD) for eight weeks. Oleic acid (OA)-induced steatotic hepatocytes were used as the NAFLD mode cells. Lipid content in liver tissues was analyzed. Smurf1-MDM2 interaction, MDM2 and p53 ubiquitination, and p53 target genes expression in liver tissues and hepatocytes were analyzed. We found that hepatic Smurf1 is highly expressed in NAFLD patients and HFD-induced NAFLD mice. Its deletion attenuates hepatocyte steatosis. Mechanistically, Smurf1 interacts with and stabilizes mouse double minute 2 (MDM2), promoting p53 degradation. In Smurf1-deficient hepatocytes, an increase in p53 suppresses SREBP-1c expression and elevates the expression of both malonyl-CoA decarboxylase (MCD) and lipin1 (Lpin1), two essential proteins in lipid catabolism. Contrarily, the activities of these three proteins and hepatocyte steatosis are reversed by p53 knockdown in Smurf1-deficient hepatocytes. This study shows that Smurf1 is involved in the pathogenesis of NAFLD by balancing de novo lipid synthesis and lipolysis.
Similar content being viewed by others
Introduction
Non-alcoholic fatty liver disease (NAFLD), the most prevalent chronic liver disease affecting a quarter of the global population worldwide1,2, is the second primary indicator for liver transplantation and the third leading cause of hepatocellular carcinoma (HCC)3. Additionally, it is also associated with type II diabetes, chronic kidney disease, and cardiovascular disease4. Currently, there are no effective treatments for NAFLD5, with only 6% of patients meeting the requirements for weight loss treatment, the first-line treatment for NAFLD6. Furthermore, although excessive lipid accumulation within hepatocytes is the primary pathogenic driver of NAFLD, the fundamental pathogenesis relevant to hepatic lipid overload is still unclear. To combat NAFLD, it is therapeutically advantageous to explore treatments to enhance lipid degradation or inhibit de novo lipogenesis.
Hepatic de novo lipid synthesis in mammalian cells undergoes strict transcriptional regulation7. Sterol regulatory element-binding protein-1c (SREBP-1c), an essential transcriptional factor responsible for triglyceride (TG) synthesis, can bind to the cis-elements of the promoter region of related genes, such as fatty acid synthase (FASN), acetyl coenzyme A carboxylase (ACC), and stearoyl-CoA desaturase-1 (SCD-1), to initiate their transcription8,9,10. Recently, we demonstrated that depletion of Smad ubiquitination regulatory factor 1 (Smurf1) alleviates hepatic lipid accumulation by promoting SREBP-1c ubiquitination and degradation in mice11, indicating that Smurf1 could be a novel determinant governing hepatic lipogenesis. However, whether alleviation of lipid deposition provided by Smurf1 gene knockout is directly associated with SREBP-1c ubiquitination remains unclear.
p53 has been characterized as a potent regulator controlling cell growth and an essential signaling mediator12,13. Recent studies have revealed that p53 plays an important role in regulating lipid metabolism. p53 can transcriptionally repress SREBP-1c expression, thereby inhibiting de novo lipogenesis14 and enhancing lipolysis via stimulating a series of genes15. In fasted mouse livers and C2C12 myoblasts, p53 elevates the activities of key lipolytic enzymes malonyl-CoA decarboxylase (MCD) and lipin 1 (Lpin1)16,17. p53 is also strictly regulated by the ubiquitin-proteasome system18,19. Inhibiting p53 ubiquitination and degradation is conducive to reducing lipid accumulation in steatosis hepatocytes. p53 ubiquitination is restricted by another protein, mouse double minute 2 (MDM2), a p53-specific E3 ubiquitin ligase that functions as the principal cellular antagonist of p53 to limit the p53 growth-suppressive function in unstressed cells20. A previous study demonstrated that Smurf1 interacts with MDM2 to inhibit MDM2 auto-degradation, ultimately promoting p53 ubiquitination and degradation21. Therefore, we speculated that Smurf1 deletion could protect p53 from ubiquitination, relieving lipid accumulation within hepatocytes.
Materials and methods
Human liver samples
NAFLD and control liver samples and blood samples were collected from Beijing Youan Hospital, Capital Medical University, Beijing, China. All procedures involving human liver samples were approved by the Ethics Committee of Beijing Youan Hospital and conducted following the principles of the Declaration of Helsinki. Supplementary Table S1 shows all patients’ information.
Animals and treatment
Wild-type and Smurf1-KO male C57BL/6N mice11 at eight weeks old were reared in standard cages with a 12:12 h light/dark cycle and free access to normal chow diets (CD, D12450B: 10% Kcal fat) or high-fat diets (HFD, D12492: 60% Kcal fat) for eight weeks (n = 6 each). After that, their livers were harvested and stored at −80 °C or fixed in 10% buffered formalin. The study was approved by the Institutional Animal Care and Usage Committee of Capital Medical University. All animal experiments were conducted following the guidelines for Care and Use of Animals.
Cell culture
HepG2 cells from the American Type of Cell Collection (ATCC, Manassas, VA, USA) were cultured at 37 °C in DMEM media (Gibco, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum (FBS, Gibco) in a humidified incubator with 5% CO2.
Antibodies
The study utilized antibodies against Smurf1 (ab57573, Abcam, Cambridge, MA, USA), MDM2 (ab38618, Abcam), SREBP-1c (ab3259, Abcam), p53 (2524s, Cell Signaling, Danvers, MA, USA), Lipin1 (YN1951, Immunaway, Plano, TX, USA), MLYCD (15265-1-AP, Proteintech, Rosemont, IL, USA), β-actin (66009-1-Ig, Proteintech), ubiquitin (10201-2-AP, Proteintech), Lamin A/C (10298-1-AP, Proteintech), GAPDH (60004-1-Ig, Proteintech), flag-tag (M185-3, MBL, Minato-ku, Tokyo, Japan), Myc-tag (M047-3, MBL), and HA-tag (M180-3, MBL).
Lentiviral shRNA vectors, plasmids, and small interfering RNAs (siRNAs)
Lentiviruses carrying Smurf1 shRNA with 90% interference efficiency (5’-GCGTTTGGATCTATGCAAACT-3’) or with 70% interference efficiency (5’-GCCCA GAGATACGAAAGAGAT-3’) were purchased from Sangon Biotech (Shanghai, China). DNA transfections into HepG2 cells were conducted using 8 μg/mL polybrene transfection reagent. Stably transduced HepG2 cells were selected by culture in media with 1 μg/mL puromycin for one week. Flag-Smurf1, myc-MDM2 and HA-ubiquitin plasmids were constructed as described previously21. p53 siRNA (5’-CACCAUCCACUACAACUACAUT T-3’) and MDM2 siRNA (5’-CGAUUAUAUGAUGAGAAGCAATT-3’) were purchased from Sangon Biotech and transfected into HepG2 cells with Lipofectamine™ 3000 transfection reagent (L3000008, Invitrogen, Carlsbad, CA, USA). The interference efficiency was assessed by Western blot.
Primary mouse hepatocyte isolation and culture
Hepatocytes were isolated from C57BL/6N mice at the age of 6–8 weeks using the collagenase perfusion method11. After isolation, hepatocytes were resuspended in DMEM media (Gibco) containing 20% FBS, 100 units/mL penicillin, and 100 μg/mL streptomycin (Gibco), plated on cell culture plates (Corning, Steuben County, NY, USA) at 5 × 103 cells/cm2, and cultured at 37 °C in a humidified incubator with 5% CO2 for 4 h to allow attachment. After refreshing the media to remove the non-attached cells, the attached hepatocytes were further cultured as described above.
Oil-Red O staining
Oil-Red O from Sigma was dissolved in isopropyl alcohol as 0.5% stock solution and stored at 4 °C in the dark. Liver tissue sections or cells were fixed with 4% paraformaldehyde for 10 min at room temperature and stained in a freshly diluted 0.3% Oil-Red O solution for 15 min. After fully washed with PBS, rinsed in 60% isopropyl alcohol for 5 s and rewashed with PBS, the stained samples were observed microscopically. The representative lipid droplet images were obtained and analyzed using ImageJ (Rawak Software Inc., Stuttgart, Germany) and Image-pro plus (Media Cybernetics, Silver Springs, MD, USA).
Intracellular triglyceride assay
Triglyceride (TG) concentrations in liver tissues and hepatocytes were determined using the triglyceride assay kit (GPO-POD, Applygene Technologies Inc., Beijing, China) following the manufacturer’s protocols.
RNA extraction and qRT-PCR
Total RNAs were isolated using Trizol reagent, reversely transcribed into cDNA using the SuperScript™ IV One-Step RT-PCR System (12594100, Invitrogen), and subjected to qRT-PCR using SuperScript™ III Platinum™ SYBR™ Green One-Step qRT-PCR Kit (11736059, Invitrogen). Supplementary Table S2 shows the primers used in these analyses.
Immunohistochemical staining
After deparaffinization, rehydration and blocking, liver tissues were incubated overnight at 4 °C with specific primary antibodies after diluted according to the manufacturers’ recommendation. Thereafter, proteins were visualized and observed under a microscope. Signals in Smurf1-KO mice were compared with those in WT mice.
Immunoprecipitation
Proteins were extracted from liver tissues and HepG2 cells and subjected to immunoprecipitation using the indicated antibodies described previously11.
In vivo ubiquitination assays
HepG2 cells were treated with 10 μM proteasome inhibitor MG132 (HY-13259, MCE, Monmouth Junction, NJ, USA) for 12 h and lysed with RIPA buffer (C1053-100, Applygene). After centrifugation, the supernatant was incubated first with the indicated antibodies for 8 h, then with protein A/G agarose beads (SC-2003, Santa Cruz Biotechnology, Santa Cruz, CA, USA) overnight at 4 °C. Ubiquitination modification was detected by Western blot. Similarly, liver tissues were lysed with RIPA buffer and subjected to Western blot as described above.
Determination of reactive oxygen species
Reactive oxygen species (ROS) in hepatocytes were determined using reactive oxygen species assay kit (CA1410, Solarbio, Beijing, China) following the manufacturer’s protocols.
Statistical analysis
Statistical analyses were performed with GraphPad Prism8 (GraphPad Software, La Jolla, CA, USA). Data are expressed as mean ± standard deviation (SD) and assessed by the two-tailed Student’s t test or the ANOVA test. A difference with p value < 0.05 was considered significant.
Results
Smurf1 is highly expressed in the livers of NAFLD patients
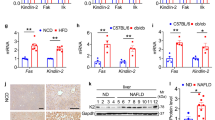
The liver samples were collected from both healthy individuals and patients with mild-moderate NAFLD as diagnosed by hematoxylin-eosin (H&E) staining and NAFLD activity score (NAS) (Fig. 1A, B). The immunohistochemical staining showed that hepatic Smurf1 levels were significantly enhanced in NAFLD patients (Fig. 1C). Q-PCR results showed that serum Smurf1 mRNA levels were increased in NAFLD patients than in healthy individuals (Fig. 1D). Then, we fed wild-type (WT) mice with a high-fat diet (HFD) or a chow diet (CD) for eight weeks. More visible lipid droplets and higher NAS in hepatocytes were observed in the HFD-fed mice than in the CD-fed mice (Fig. 1E, F), indicating that the mouse NAFLD model was successfully established. Consistent with the findings in patients, Smurf1 protein levels were higher in HFD-fed mice than in CD-fed mice (Fig. 1G, H). Moreover, oleic acid (OA)-treatment of mouse hepatocytes for 24 h or HepG2 cells for 48 h increased Smurf1 protein (Fig. 1I, J) and mRNA (Fig. 1K, L) levels as compared with the control hepatocytes, indicating that Smurf1 is overexpressed in NAFLD and potentially involved in the pathogenesis of NAFLD.
A, B Hepatic lipid droplets in NAFLD patients and healthy controls were analyzed using hematoxylin-eosin (H&E) staining and NAFLD activity score (NAS) (n = 6/group, scale bar = 100 μm (left) or 50 μm (right)). C Hepatic Smurf1 levels in NAFLD patients were analyzed using immunohistochemical staining (n = 6/group, scale bar = 100 μm (left) or 50 μm (right)). D Serum Smurf1 mRNA levels in NAFLD patients and healthy controls (n = 6/group). E, F H&E staining and NAS of liver sections of WT male mice fed with CD or HFD for 8 weeks (n = 6 mice/group, scale bar = 100 μm (left) or 50 μm (right)). G, H Hepatic Smurf1 levels in mice were analyzed using immunohistochemical staining and Western blot (n = 6 mice/group, scale bar = 100 μm (left) or 50 μm (right)). I–L Primary hepatocytes from WT male mice and HepG2 cells were treated with 100 μM OA for the indicated times. Smurf1 protein (I, J) and mRNA (K, L) levels were analyzed by Western blot and qRT-PCR, respectively. Liver section images were obtained using an optical microscope with 200/400× magnification and analyzed using Image-pro plus and GraphPad Prism8. All data are presented as mean ± SD (n ≥ 3). *p < 0.05, **p < 0.01, ***p < 0.001.
Smurf1 deficiency attenuates HFD-induced hepatocyte steatosis
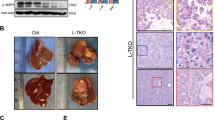
We then questioned if Smurf1 deficiency in hepatocytes could attenuate NAFLD. We found that after HFD feeding, Smurf1-KO mice exhibited less severe hepatic steatosis (Fig. 2A, B), lower NAS (Fig. 2C), and reduced hepatic TG levels (Fig. 2D) than WT mice. Consistent with the in vivo staining results, administration of 100 μM OA increased lipid droplets and TG levels in primary hepatocytes from both Smurf1-KO and WT mice. However, their levels were three-fold lower in Smurf1-KO hepatocytes than in WT hepatocytes (Fig. 2E, F). Moreover, HepG2 cells with Smurf1-shRNA or control-shRNA transfection were treated with 100 μM OA for the indicated times. Oil Red-O staining revealed a gradually increased lipid droplet number with prolonged OA treatment in both control and Smurf1- knockdown cells. However, the number of lipid droplets was significantly lower in Smurf1- knockdown HepG2 cells than in control HepG2 cells at various times (Fig. 2G). We further reintroduced Smurf1 into Smurf1-knockdown cells and found that Smurf1 transfection significantly aggravated lipid accumulation in hepatocytes (Fig. 2H), clearly indicating that Smurf1 is associated with hepatocyte steatosis.
A, B Oil droplets in liver sections of WT and Smurf1-KO male mice fed with CD or HFD for eight weeks were observed using H&E staining and Oil Red-O staining (n = 6 mice/group, scale bar = 100 μm (left) or 50 μm (right)). C, D NAS and triglyceride (TG) of WT and Smurf1-KO male mice were measured. E Oil red O staining of primary hepatocytes from WT and Smurf1-KO mice after BSA or 100 μM OA treatment for 24 h. (n = 6 mice/group, scale bar = 50 μm). F TG levels in primary mouse hepatocytes were measured. G Oil red O staining of Smurf1-shRNA transfected and control hepatocytes after BSA or 100 μM OA treatment for the indicated times (scale bar = 50 μm). H Oil red O staining of control, Smurf1-shRNA transfected, and Smurf1-shRNA and flag-Smurf1 co-transfected hepatocytes after BSA or 500 μM OA treatment for 24 h (scale bar = 50 μm). Liver sections or cells images were obtained using an optical microscope with 200/400× magnification and analyzed using Image-pro plus and ImageJ. All data are presented as mean ± SD (n ≥ 3). *p < 0.05, **p < 0. 01, ***p < 0.001, ****p < 0.0001.
Smurf1 negatively regulates p53 protein by stabilizing MDM2 in hepatocytes
p53 plays a pivotal role in NAFLD progression by promoting lipids oxidation and inhibiting lipids synthesis22,23 and is tightly regulated by ubiquitination via MDM2, the primary E3 ligase of p5319,20. Moreover, studies have confirmed that Smurf1 interacts with its target substrates for ubiquitination, including RhoA, MEKK2, Smad1/5, and JunB24. Thus, we speculated that MDM2 is a Smurf1-modified ubiquitination substrate. To our surprise, immunohistochemical staining showed that hepatic MDM2 protein level was decreased in Smurf1-KO mice than in WT mice (Fig. 3A), suggesting that Smurf1 could stabilize MDM2. Contrarily, hepatic p53 protein was elevated in Smurf1-KO mice (Fig. 3B, C). Consistent with the observations in mouse livers, treatment of primary hepatocytes from Smurf1-KO mice with 100 μM OA for 24 h decreased MDM2 but increased p53 protein levels (Fig. 3D). Moreover, these expression patterns were also confirmed in HepG2 cells treated with 500 μM OA for 24 h (Fig. 3E) or 100 μM OA for the indicated times (Fig. 3F–H). In addition, Smurf1-deficient HepG2 cells with different interference efficiency were used in the rescue experiments (Supplementary Fig. S1A, B). Smurf1 re-delivery into Smurf1-deficient HepG2 cells effectively increased MDM2 level and declined p53 stability (Fig. 3I). Similar to the observation in Smurf1 reintroduced HepG2 cells, MDM2 overexpression in HepG2 cells reduced p53 protein level (Fig. 3J). Contrarily, MDM2 knockdown in Smurf1 overexpressing HepG2 cells eliminated the inhibitory effect of Smurf1 on p53 stability (Fig. 3K). These data indicate that Smurf1 negatively regulates p53 protein via MDM2.
A–C Immunohistochemical staining and Western blot results of MDM2 and p53 expression in the livers of WT and Smurf1-KO mice fed with CD or HFD for eight weeks (n = 6 mice/group, scale bar = 100 μm). D Western blot results of Smurf1, MDM2 and p53 levels in primary mouse hepatocytes from WT and Smurf1-KO mice after BSA or 100 μM OA treatment for 24 h. Western blot results of Smurf1, MDM2, and p53 expression levels in Smurf1-shRNA transfected and control hepatocytes after 500 μM (E) or 100 μM (F) OA treatment for the indicated times. G, H Densitometric quantities of MDM2 and p53 levels in F. Western blot results of Smurf1, MDM2 and p53 expression levels in control, Smurf1-shRNA transfected, Smurf1-shRNA and flag-Smurf1 co-transfected (I) and Smurf1-shRNA and myc-MDM2 co-transfected (J) hepatocytes after BSA or 500 μM OA treatment for 24 h. K Western blot results of Smurf1, MDM2, and p53 expression levels in control, flag-Smurf1 transfected, and flag-Smurf1 and MDM2-siRNAs co-transfected hepatocytes after BSA or 500 μM OA treatment for 24 h. All data are presented as mean ± SD (n ≥ 3). *p < 0. 05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Smurf1 interacts with MDM2 and inhibits MDM2 ubiquitination and degradation in NAFLD livers
To explore the mechanism underlying Smurf1-mediated MDM2 elevation, we explored the interaction of Smurf1 with MDM2 using co-immunoprecipitation (Co-IP) assays. The results indicated that endogenous Smurf1 interacts with endogenous MDM2 (Fig. 4A) and transfected myc-MDM2 (Fig. 4B) in HepG2 cells. Furthermore, the interaction between endogenous Smurf1 and MDM2 was enhanced in HFD-fed mice than in CD-fed mice (Fig. 4C). We then explored the impact of Smurf1 on MDM2 ubiquitination using ubiquitination assays and found that endogenous MDM2 ubiquitination was elevated in HFD-fed Smurf1-KO mice (Fig. 4D). Identical to the observation in vivo, MDM2 ubiquitination was increased in Smurf1-deficient HepG2 cells after OA treatment. However, MDM2 ubiquitination was decreased after reintroducing Smurf1 into Smurf1-deficient HepG2 (Fig. 4E). These data suggest that Smurf1 interacts with MDM2 to prevent its ubiquitination and degradation.
Co-immunoprecipitation (co-IP) assay was performed to detect the interaction of endogenous Smurf1 with MDM2 (A) and myc-MDM2 (B) in HepG2 cells and with endogenous MDM2 (C) in mouse livers. Ubiquitination assay was performed to detect MDM2 ubiquitination in the livers of WT and Smurf1-KO mice fed with CD or HFD for eight weeks (D) and in HepG2 cells transfected with HA-ubiquitin, Smurf1-shRNA, and flag-Smurf1 after treated with 500 μM OA for 24 h (E). Data shown are the representatives of at least three individual experiments with similar results.
Smurf1 promotes MDM2-induced p53 ubiquitination in NAFLD procession
A previous study has demonstrated that Smurf1 promotes p53 ubiquitination and degradation independent of its E3 ligase function21. We further examined whether Smurf1- regulated p53 ubiquitination is dependent on MDM2 in NAFLD mice. As shown in Fig. 5A, endogenous p53 ubiquitination was significantly decreased in HFD-fed Smurf1-KO mice than in HFD-fed WT mice. Moreover, p53 ubiquitination were reduced in the nuclear and cytoplasmic compartments of the livers of HFD-fed Smurf1-KO mice than those from the HFD-fed WT mice (Fig. 5B). Furthermore, cytoplasmic p53 ubiquitination was more prominent (Fig. 5B), consistent with previous reports that p53 ubiquitination and degradation mainly occur in the cytoplasm25,26. Corresponding to the results of p53 ubiquitination, Western blot showed that nuclear p53 expression was increased in Smurf1-KO mice than in WT mice (Fig. 5C), suggesting that Smurf1 deficiency enhances nuclear p53 stability by inhibiting p53 ubiquitination in the cytoplasm. Similarly, p53 ubiquitination level was reduced in Smurf1-knockdown HepG2 cells than in control HepG2 cells with OA treatment. However, this phenomenon was obviously reversed by re-delivering Smurf1 into the cells (Fig. 5D). To further confirm that Smurf1 regulation of p53 ubiquitination is a prerequisite for MDM2 involvement, we further elevated MDM2 expression in Smurf1 knockdown hepatocytes. The results showed that MDM2 overexpression increased p53 ubiquitination level in Smurf1-deficient cells (Fig. 5E). Contrarily, MDM2 knockdown inhibited the promoting effect of Smurf1 on p53 ubiquitination in Smurf1-overexpressing HepG2 cells (Fig. 5F). Overall, Smurf1 promotes p53 ubiquitination and degradation in a MDM2-dependent manner. Thus, we speculated that Smurf1 deficiency could inhibit lipid accumulation by stabilizing p53. We then knocked down p53 in Smurf1 knockdown hepatocytes and treated these hepatocytes with 500 μM OA for 24 h. The Oil Red-O staining showed that p53 knockdown significantly counteracted the inhibition of Smurf1 deficiency on hepatic lipid accumulation (Fig. 5G).
Ubiquitination assay was performed to detect the ubiquitination levels of total p53 (A), nuclear p53 and cytoplasmic p53 (B) in the livers of WT and Smurf1-KO male mice fed with CD or HFD for eight weeks. C Western blot results of the nuclear and cytoplasmic p53 levels in the livers of WT and Smurf1-KO mice fed with CD or HFD for eight weeks. Ubiquitination assay was performed to detect p53 ubiquitination in HepG2 cells transfected with HA-ubiquitin, Smurf1-shRNA, and flag-Smurf1 (D) or myc-MDM2 (E) after treated with 500 μM OA for 24 h, in HepG2 cells transfected with HA-ubiquitin, flag-Smurf1, and MDM2-siRNAs (F) after treated these with 500 μM OA for 24 h. G Shown are the oil red O staining images of control, Smurf1-shRNA transfected, and Smurf1-shRNA and p53-siRNAs co-transfected hepatocytes after BSA or 500 μM OA treatment for 24 h (scale bar = 50 μm) at 400× magnification and the analysis results using Image-pro plus and ImageJ. Data shown are the representatives of at least three individual experiments with similar results.
Smurf1 depletion regulates p53 target genes in lipid metabolism at both mRNA and protein levels
We analyzed mRNA levels of three lipid metabolism-related p53 target genes, SREBP-1c, MCD, and Lpin1, in HFD- and CD-fed Smurf1-KO mice. The results showed that Smurf1 deletion decreased SREBP-1c and increased MCD and Lpin1 mRNA levels in mouse livers (Fig. 6A). Similar results were also found in primary mouse hepatocytes isolated from Smurf1-KO mice regardless OA treatment (Fig. 6B). Additionally, lower SREBP-1c (Fig. 6C) and higher MCD and Lpin1 mRNA levels (Fig. 6D, E) were found in Smurf1-deficient cells than in control cells after incubation with 100 μM OA for different times. We then reintroduced flag-tagged Smurf1 into Smurf1-deficient HepG2 cells and found that Smurf1 re-transfection reversed the changes in SREBP-1c, MCD and Lpin1 caused by Smurf1 deficiency (Fig. 6F).
A, B qRT-PCR analysis of SREBP-1c, MCD, and Lpin1 mRNAs in mouse livers and primary hepatocytes. The WT and Smurf1-KO mice were fed with CD or HFD for eight weeks. Mouse primary hepatocytes were treated with BSA or 100 μM OA 24 h. qRT-PCR analysis of SREBP-1c (C), MCD (D), and Lpin1 (E) mRNA levels in Smurf1-shRNA transfected and control hepatocytes after BSA or 100 μM OA treatment for the indicated times. F qRT-PCR analysis of SREBP-1c, MCD, and Lpin1 mRNA levels in control, Smurf1-shRNA transfected, and Smurf1-shRNA and flag-Smurf1 co-transfected hepatocytes after BSA or 500 μM OA treatment for 24 h. All data are presented as mean ± SD (n ≥ 3). *p < 0.05, **p < 0.01, and ***p < 0.001, ****p < 0.0001.
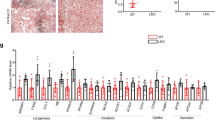
We further analyzed SREBP-1c, MCD, and Lpin1 protein levels in HFD- and CD-fed Smurf1-KO mice. Both immunohistochemical staining and Western blot showed that Smurf1 depletion decreased SREBP-1c but increased MCD and Lpin1 protein levels in mouse livers (Fig. 7A, B). Consistently, these results were also confirmed in primary hepatocytes isolated from Smurf1-KO mice regardless of OA treatment (Fig. 7C). Furthermore, lower SREBP-1c and higher MCD and Lpin1 protein levels were also found in Smurf1-deficient HepG2 cells compared with control HepG2 cells after incubation with 500 μM OA for 24 h (Fig. 7D) or 100 μM OA for the indicated times (Fig. 7E–H). As a transcription factor, SREBP-1c needs to be cleaved and translocated into the nucleus to transcriptionally activate its target genes. We found that both nuclear and cytoplasmic SREBP-1c protein levels were decreased in the livers of HFD-fed Smurf1-KO mice than those in HFD-fed WT mice (Fig. 7I). Combined with the results that Smurf1 deletion reduces SREBP-1c mRNA level (Fig. 6), it is assumed that Smurf1 deficiency could inhibit SREBP-1c synthesis. Considering that MCD and Lpin1 are two important lipolytic enzymes that may affect ROS levels, we further analyzed ROS levels in the control and Smurf1-deficient HepG2 cells upon OA treatment. Although no significant difference in ROS levels was observed between the control and Smurf1-deficient HepG2 cells, ROS levels were gradually reduced in HepG2 cells with prolongation of OA induction (Supplementary Fig. S2A, B), indicating that MCD and Lpin1 mainly promote fatty acid oxidation rather than increase oxidative stress.
A, B Immunohistochemical staining and Western blot were performed to detect SREBP-1c, MCD, and Lpin1 levels in the livers of WT and Smurf1-KO mice fed with CD or HFD for eight weeks (n = 6 mice/group, Scale bar = 100 μm). C Western blot was performed to detect SREBP-1c, MCD, and Lpin1 expression in mouse primary hepatocytes from WT and Smurf1-KO mice after BSA or 100 μM OA treatment for 24 h. Western blot was performed to detect SREBP-1c, MCD, and Lpin1 expression in Smurf1-shRNA transfected and control transfected hepatocytes after 500 μM (D) or 100 μM (E) OA treatment for the indicated times. F–H SREBP-1c, MCD, and Lpin1 expression in E were quantified using densitometry. I Western blot results of the nuclear and cytoplasmic SREBP-1c levels in the livers of WT and Smurf1-KO mice fed with CD or HFD for eight weeks. Liver section images were obtained under an optical microscope at a 400× magnification and analyzed using Image-pro plus and GraphPad Prism8. All data are presented as mean ± SD (n ≥ 3). *p < 0. 05, **p < 0. 01, and ***p < 0. 001, ****p < 0.0001.
Smurf1 depletion regulates p53 target genes in lipid metabolism via the MDM2-p53 pathway
To explore whether Smurf1 deficiency regulates SREBP-1c, MCD, and Lpin1 through the MDM2-p53 pathway, we overexpressed Smurf1 in Smurf1 knockdown HepG2 cells. Western blot showed that Smurf1 overexpression reversed the changes of SREBP-1c, MCD, and Lpin1 caused by Smurf1 deficiency (Fig. 8A). Moreover, we overexpressed MDM2 or knocked down p53 in Smurf1-knockdown hepatocytes to rescue MDM2 reduction and p53 elevation caused by Smurf1 deficiency. Consistent with the observation in Smurf1 reintroduced HepG2 cells, MDM2 overexpression or p53 knockdown reversed the changes of SREBP-1c, MCD, and Lpin1 caused by Smurf1 deficiency (Fig. 8B, C). These results indicate that Smurf1 regulates lipid accumulation through the MDM2-p53 pathway.
A Western blot was performed to detect SREBP-1c, MCD, and Lpin1 expression in control, Smurf1-shRNA transfected, and Smurf1-shRNA and flag-Smurf1 co-transfected hepatocytes after BSA or 500 μM OA treatment for 24 h. B, C Western blot was performed to detect Smurf1, MDM2, p53, SREBP-1c, MCD, and Lpin1 expression levels in control, Smurf1-shRNA transfected, and Smurf1-shRNA and myc-MDM2 co-transfected (B) or Smurf1-shRNA and p53-siRNAs co-transfected (C) hepatocytes after BSA or 500 μM OA treatment for 24 h. Data shown are the representatives of at least three individual experiments with similar results. D The proposed mechanism linking Smurf1 to hepatic lipid accumulation: Smurf1 binds to MDM2 and stabilizes MDM2 to promote p53 ubiquitination and degradation. p53 decrease facilitates SREBP-1c transcription, which is required for lipogenesis, while inhibits MCD and Lpin1 transcription, which is required for lipolysis. As a result, Smurf1 aggravates hepatic lipid accumulation.
Discussion
As a classic tumor-suppressing gene, p53 plays a pivotal role in cell cycle arrest, apoptosis, aging, and other signaling pathways27,28. However, accumulating evidence shows that p53 might also be involved in the pathogenesis of NAFLD16,22,29,30,31. Although the relationship between p53 and NAFLD has been investigated, the specific role of p53 linked to NAFLD is still controversial32. It has been reported that p53 inhibits hepatic lipid accumulation and inflammation under normal or mild stress conditions16,32,33,34,35. Conversely, alternative studies believed that p53 promotes lipid accumulation and oxidative stress under severe stress conditions, such as nonalcoholic steatohepatitis (NASH)36,37,38. It is also reported that p53 expression gradually increases with NAFLD progression39. Hence, p53 plays a crucial and complex role in NAFLD development, exerting a protective role in the early stage but a worsening role in the middle and late stages of NAFLD32. Many studies have shown that p53 is essential in lipid metabolism by reducing lipogenesis via repressing the expression of SREBP-1c and glucose-6-phosphate dehydrogenase (G6PD)14,40 and increasing lipolysis via inducing expression of MCD, Lpin1, sirtuin 1, aromatase, and caveolin 116,17,23,41,42. Thus, maintaining p53 stability in the early stage of NAFLD is likely to be the core element to prevent NAFLD from deteriorating into NASH. p53 stability is primarily regulated by post-translational modifications, including ubiquitination and ubiquitin-like modifications, among which ubiquitination is the main p53 degradation pathway43,44,45,46,47. It seems that inhibition of p53 ubiquitination plays a positive role in the early stage of NAFLD. To simulate mild fatty liver, we used 8-week HFD-fed mice and OA-induced hepatocytes as the NAFLD models in the study.
The ubiquitination process is mediated sequentially by ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2), and ubiquitin ligases (E3)48. During the process, ubiquitin is transferred from E1 to E2 and then to the substrates mediated by E349, the most important ubiquitination enzyme determining the specificity of receptor substrates50. Several E3-ligases have been identified and characterized51,52,53,54. Smurf1, a HECT-type E3 of the Nedd4 family55,56, is closely related to many physiological processes, including bone formation, embryonic development, neural outgrowth, and tumor cells invasion55. Our recent report unveils Smurf1 as one of the key players during NAFLD occurrence. By directly interacting with SREBP-1c in mouse livers, Smurf1 promotes the binding of SREBP-1c with Fbw7a, a major ubiquitin E3 ligase of SREBP-1c, to suppress SREBP-1c ubiquitination11. Smurf1 was also able to bind with MDM2 to inhibit its auto-ubiquitination, ultimately promoting p53 ubiquitination21. Thus, we speculate that the involvement of Smurf1 during NAFLD could possibly be associated with its regulating p53 ubiquitination. It is known that ubiquitination substrates of Smurf1 do not include p5321. Therefore, we speculated that the effect of Smurf1 on p53 ubiquitination is potentially mediated by an intermediate medium, MDM2, one of the most important ubiquitin ligases (E3) of p5318,19. As expected, we found Smurf1 fails to promote p53 degradation without MDM2 (Fig. 3K). Therefore, we believe that the core significance of the current work relays on confirmation of the fact that ubiquitination of p53 offered by Smurf1 is largely dependent upon the presence of MDM2, which affects p53-regulated lipid metabolism during HFD-fed mice (Fig. 5A), which echoes with the high p53 level in the livers of Smurf1-KO mice (Fig. 3). Moreover, our data also clearly demonstrate that Smurf1 interacts with MDM2 in mouse livers, and this interaction appears to be more intense in the livers of HFD-fed mice (Fig. 4C), possibly due to high Smurf1 expression induced by HFD. These results indicate that enhanced p53 ubiquitination by Smurf1 via MDM2 is involved in NAFLD development.
Among the p53 target genes related to lipid metabolism14,15,16,17,23,40,41,42, we found that Smurf1 knockdown significantly affects the transcription of SREBP-1c, MCD, and Lpin1. We further confirmed that Smurf1 deficiency reduces SREBP-1c and increases MCD and Lpin1 expressions in the livers of HFD-fed mice (Figs. 6 and 7) and revealed that these changes are strictly regulated by the MDM2-p53 pathway (Fig. 8). MCD and Lpin1 are mainly involved in fatty acid oxidation. They activate mitochondrial fatty acid oxidation mainly by increasing carnitine palmitoyl transferase (CPT) activity and upregulating peroxisome proliferator activated receptor α (PPARα) level. Fatty acid oxidation is a complex physiological process in NAFLD progression. It could reduce lipid accumulation and may produce oxidative stress in the middle and late stages of NAFLD. How Smurf1 regulates mitochondrial fatty acid oxidation through the MDM2-p53 pathway should be further clarified. Moreover, our current research mainly focuses on the role of Smurf1 in lipid accumulation. In addition to steatosis, the hits of lipotoxicity, oxidative stress, and inflammatory cascade activation all play vital roles in the development of NAFLD. Whether Smurf1 could regulate these hits? Whether the inhibitory effect of Smurf1 on p53 stability would sustain the progression of NAFLD? These scientific questions constitute the ongoing research focus of our future work. Therefore, we plan to dynamically analyze NAFLD development in combination with Smurf1-KO mice and try to figure out essential genes or proteins that substantially promote progression of NAFLD to NASH, liver fibrosis or cirrhosis. We will characterize the regulatory effects of Smurf1 on NAFLD occurrence and development in our future studies.
In summary, this study reveals a possible mechanism of Smurf1 involvement in NAFLD pathogenesis via the MDM-p53 pathway. Mechanically, Smurf1 interacts with and stabilizes MDM2 to promote p53 ubiquitylation and degradation, leading to enhanced lipogenesis by transcriptionally inducing SREBP-1c and inhibited lipolysis by repressing MCD or Lipin1 (Fig. 8D). Our study provides new insight into the possible pathogenesis of NAFLD.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Sharpton SR, Schnabl B, Knight R, Loomba R. Current Concepts, Opportunities, and Challenges of Gut Microbiome-Based Personalized Medicine in Nonalcoholic Fatty Liver Disease. Cell Metab. 33, 21–32 (2021)
Cotter TG, Rinella M. Nonalcoholic Fatty Liver Disease 2020: The State of the Disease. Gastroenterology 158, 1851–1864 (2020)
Wong RJ, Aguilar M, Cheung R, Perumpail RB, Harrison SA, Younossi ZM, et al. Nonalcoholic steatohepatitis is the second leading etiology of liver disease among adults awaiting liver transplantation in the United States. Gastroenterology 148, 547–55 (2015)
Younossi ZM, Koenig AB, Abdelatif D, Fazel Y, Henry L, Wymer M. Global epidemiology of nonalcoholic fatty liver disease-meta-analytic assessment of prevalence, incidence, and outcomes. Hepatology 64, 73–84 (2016)
Sasaki A, Nitta H, Otsuka K, Umemura A, Baba S, Obuchi T, et al. Bariatric surgery and non-alcoholic Fatty liver disease: current and potential future treatments. Front Endocrinol 5, 164 (2014)
Younossi ZM, Loomba R, Rinella ME, Bugianesi E, Marchesini G, Neuschwander-Tetri BA, et al. Current and future therapeutic regimens for nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Hepatology 68, 361–371 (2018)
Buzzetti E, Pinzani M, Tsochatzis EA. The multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD). Metabolism 65, 1038–48 (2016)
Kawano T, Cohen DE. Mechanisms of hepatic triglyceride accumulation in non-alcoholic fatty liver disease. J. Gastroenterol 48, 434–441 (2013)
Eberle D, Hegarty B, Bossard P, Ferre P, Foufelle F. SREBP transcription factors: Master regulators of lipid homeostasis. Biochimie 86, 839–48 (2004)
Kim CW, Addy C, Kusunoki J, Anderson NN, Deja S, Fu X, et al. Acetyl CoA Carboxylase Inhibition Reduces Hepatic Steatosis but Elevates Plasma Triglycerides in Mice and Humans: A Bedside to Bench Investigation. Cell Metab. 26, 394–406 (2017)
Zhang X, Zhan Y, Lin W, Zhao F, Guo CJ, Chen YJ, et al. Smurf1 aggravates non-alcoholic fatty liver disease by stabilizing SREBP-1c in an E3 activity-independent manner. FASEB J. 34, 7631–7643 (2020)
Marcel V, Long FNV, Diaz JJ. 40 Years of Research Put p53 in Translation. Cancers (Basel) 10, 152 (2018)
Levine AJ. p53: 800 million years of evolution and 40 years of discovery. Nat. Rev. Cancer 20, 471–480 (2020)
Yahagi N, Shimano H, Matsuzaka T, Najima Y, Sekiya M, Nakagawa Y, et al. p53 activation in adipocytes of obese mice. J. Biol Chem. 278, 25395–25400 (2003)
Parrales A, Iwakuma T. p53 as a Regulator of Lipid Metabolism in Cancer. Int. J. Mol. Sci. 17, 2074 (2016)
Liu Y, He Y, Jin A, Tikunov AP, Zhou L, Tollini LA, et al. Ribosomal protein-Mdm2-p53 pathway coordinates nutrient stress with lipid metabolism by regulating mcd and promoting fatty acid oxidation. Proc Natl Acad Sci USA. 111, E2414–E2422 (2014)
Assaily W, Rubinger DA, Wheaton K, Lin Y, Ma W, Xuan W, et al. ROS-mediated p53 induction of lpin1 regulates fatty acid oxidation in response to nutritional stress. Mol. Cell 44, 491–501 (2011)
Brooks CL, Gu W. Ubiquitination, phosphorylation and acetylation: the molecular basis for p53 regulation. Curr. Opin. Cell Biol. 15, 164–71 (2003)
Michael D, Oren M. The p53-Mdm2 module and the ubiquitin system. Semin. Cancer Biol. 13, 49–58 (2003)
Fang S, Jensen JP, Ludwig RL, Vousden KH, Weissman AM. Mdm2 is a RING finger-dependent ubiquitin protein ligase for itself and p53. J. Biol. Chem. 275, 8945–51 (2000)
Nie J, Xie P, Liu L, Xing G, Chang Z, Yin Y, et al. Smad ubiquitylation regulatory factor1/2(Smurf1/2) promotes p53 degradation by stabilizing the E3 ligase MDM2. J. Biol. Chem. 285, 22818–30 (2010)
Yahagi N, Shimano H, Matsuzaka T, Sekiya M, Najima Y, Okazaki S, et al. p53 involvement in the pathogenesis of fatty liver disease. J. Biol. Chem. 279, 20571–20575 (2004)
Wang X, Zhao X, Gao X, Mei Y, Wu M. A new role of p53 in regulating lipid metabolism. J. Mol. Cell Biol. 5, 147–150 (2013)
Guo X, Shen S, Song S, He S, Cui Y, Xing G, et al. The E3 ligase Smurf1 regulates Wolfram syndrome protein stability at the endoplasmic reticulum. J. Biol. Chem. 286, 18037–47 (2011)
O’Brate A, Giannakakou P. The importance of p53 location: nuclear or cytoplasmic zip code? Drug Resist Updat. 6, 313–22 (2003)
Stommel JM, Marchenko ND, Jimenez GS, Moll UM, Hope TJ, Wahl GM. A leucine-rich nuclear export signal in the p53 tetramerization domain: regulation of subcellular localization and p53 activity by NES masking. EMBO J. 18, 1660–72 (1999)
Vogelstein B, Lane D, Levine AJ. Surfing the p53 network. Nature 408, 307–310 (2000)
Goldstein I, Marcel V, Olivier M, Oren M, Rotter V, Hainaut P. Understanding wild-type and mutant p53 activities in human cancer: new landmarks on the way to targeted therapies. Cancer Gene Ther. 18, 2-11 (2011)
Sun H, Li L, Li W, Yang F, Zhang Z, Liu Z, et al. p53 transcriptionally regulates SQLE to repress cholesterol synthesis and tumor growth. EMBO Rep. 22, e52537 (2021)
Zhang X, Lin Y, Lin S, Li C, Gao J, Feng Z, et al. Silencing of functional p53 attenuates NAFLD by promoting HMGB1-related autophagy induction. Hepatol Int. 14, 828–841 (2020)
Zhou H, Du W, Li Y, Shi C, Hu N, Ma S, et al. Effects of melatonin on fatty liver disease: The role of NR4A1/DNA-PKcs/p53 pathway, mitochondrial fission, and mitophagy. J Pineal Res. 64, (2018)
Yan Z, Miao X, Zhang B, Xie J. p53 as a double-edged sword in the progression of non-alcoholic fatty liver disease. Life Sci. 215, 64–72 (2018)
Finck BN, Gropler MC, Chen Z, Leone TC, Croce MA, Harris TE, et al. Lipin 1 is an inducible amplifier of the hepatic PGC-1alpha/PPARalpha regulatory pathway. Cell Metab. 4, 199–210 (2006)
Schwabe RF, Brenner DA. Mechanisms of Liver Injury. I. TNF-alpha-induced liver injury: role of IKK, JNK, and ROS pathways. Am. J. Physiol. Gastrointest. Liver Physiol. 290, G583–9 (2006)
Porteiro B, Fondevila MF, Buque X, Gonzalez-Rellan MJ, Fernandez U, Mora A, et al. Pharmacological stimulation of p53 with low-dose doxorubicin ameliorates diet-induced nonalcoholic steatosis and steatohepatitis. Mol. Metab. 8, 132–143 (2018)
Xu Y, Zhu Y, Hu S, Xu Y, Stroup D, Pan X, et al. Hepatocyte Nuclear Factor 4α Prevents the Steatosis-to-NASH Progression by Regulating p53 and Bile Acid Signaling (in mice). Hepatology 73, 2251–2265 (2021)
Tomita K, Teratani T, Suzuki T, Oshikawa T, Yokoyama H, Shimamura K, et al. p53/p66Shc-mediated signaling contributes to the progression of non-alcoholic steatohepatitis in humans and mice. J. Hepatol. 57, 837–43 (2012)
Farrell GC, Larter CZ, Hou JY, Zhang RH, Yeh MM, Williams J, et al. Apoptosis in experimental NASH is associated with p53 activation and TRAIL receptor expression. J. Gastroenterol Hepatol. 24, 443–52 (2009)
Panasiuk A, Dzieciol J, Panasiuk B, Prokopowicz D. Expression of p53, Bax and Bcl-2 proteins in hepatocytes in non-alcoholic fatty liver disease. World J. Gastroenterol. 12, 6198–202 (2006)
Jiang P, Du W, Wang X, Mancuso A, Gao X, Wu M, et al. p53 regulates biosynthesis through direct inactivation of glucose-6-phosphate dehydrogenase. Nat. Cell Biol. 13, 310–316 (2011)
Nemoto S, Fergusson MM, Finkel T. Nutrient availability regulates sirt1 through a forkhead-dependent pathway. Science 306, 2105–2108 (2004)
Bist A, Fielding CJ, Fielding PE. p53 regulates caveolin gene transcription, cell cholesterol, and growth by a novel mechanism. Biochemistry 39, 1966–1972 (2000)
Gu B, Zhu WG. Surf the post-translational modification network of p53 regulation. Int. J. Biol. Sci. 8, 672–84 (2012)
Huang YF, Wee S, Gunaratne J, Lane DP, Bulavin DV. Isg15 controls p53 stability and functions. Cell Cycle. 13, 2200–10 (2014)
Weger S, Hammer E, Heilbronn R. Topors acts as a SUMO-1 E3 ligase for p53 in vitro and in vivo. FEBS. Lett. 579, 5007–12 (2005)
Maki CG, Huibregtse JM, Howley PM. In vivo ubiquitination and proteasome-mediated degradation of p53(1). Cancer Res. 56, 2649–54 (1996)
Honda R, Tanaka H, Yasuda H. Oncoprotein MDM2 is a ubiquitin ligase E3 for tumor suppressor p53. FEBS. Lett. 420, 25–7 (1997)
Hegde AN, Haynes KA, Bach SV, Beckelman BC. Localubiquitin-proteasome- mediated proteolysis and long-term synaptic plasticity. Front. Mol. Neurosci. 7, 96 (2014)
Hershko A, Ciechanover A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–79 (1998)
Metzger MB, Hristova VA, Weissman AM. HECT and RING finger families of E3 ubiquitin ligases at a glance. J. Cell Sci. 125, 531–7 (2012)
Wu HQ, Baker D, Ovaa H. Small molecules that target the ubiquitin system. Biochem. Soc. Trans. 48, 479–497 (2020)
Berndsen CE, Wolberger C. New insights into ubiquitin E3 ligase mechanism. Nat. Struct. Mol. Biol. 21, 301–7 (2014)
Rotin D, Kumar S. Physiological functions of the HECT family of ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 10, 398–409 (2009)
Wang Y, Argiles-Castillo D, Kane EI, Zhou A, Spratt DE. HECT E3 ubiquitin ligases - emerging insights into their biological roles and disease relevance. J. Cell Sci. 133, jcs228072 (2020)
Cao Y, Zhang L. A Smurf1 tale: function and regulation of an ubiquitin ligase in multiple cellular networks. Cell Mol. Life Sci. 70, 2305–2317 (2013)
Ingham RJ, Gish G, Pawson T. The Nedd4 family of E3 ubiquitin ligases: Functional diversity within a common modular architecture. Oncogene 23, 1972–1984 (2004)
Acknowledgements
Pictures of liver, proteasome, and its degradation products in “Fig. 8D” were created with BioRender (https://app.biorender.com/user/signin).
Funding
This work was supported by the Chinese National Natural Science Foundation Projects (Grant no. 81570515).
Author information
Authors and Affiliations
Contributions
Y.Z., P.X. and W.A. performed study concept and design; W.L. and Y.Z. and P.X. performed development of methodology and writing, review and revision of the paper; W.L., X.Z., C. Z. and L.L. provided acquisition, analysis and interpretation of data, and statistical analysis; W.A., P.X., Y.Z. and J.Z. provided technical and material support. All authors read and approved the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All procedures involving human liver samples and blood samples were approved by the Ethics Committee of Beijing Youan Hospital and conducted following the principles of the Declaration of Helsinki. Informed consent in writing was obtained from each patient.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lin, W., Zhang, X., Zhang, C. et al. Deletion of Smurf1 attenuates liver steatosis via stabilization of p53. Lab Invest 102, 1075–1087 (2022). https://doi.org/10.1038/s41374-022-00802-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-022-00802-x
This article is cited by
-
MiR-122-5p regulates the mevalonate pathway by targeting p53 in non-small cell lung cancer
Cell Death & Disease (2023)