Abstract
This study focuses on gene expression differences between early retinal states that ultimately lead to normal development, late onset retinoblastoma, or rapid bilateral retinoblastoma tumors. The late-onset and early-onset retinoblastoma tumor cells are remarkably similar to normally proliferating retinal progenitor cells, but they fail to properly express differentiation markers associated with normal development. Further, early-onset retinoblastoma tumor cells express a robust immune gene expression signature followed by accumulation of dendritic, monocyte, macrophage, and T-lymphocyte cells in the retinoblastoma tumors. This characteristic was not shared by either normal retinae or late-onset retinoblastomas. Comparison of our data with other human and mouse retinoblastoma tumor gene expression significantly confirmed, that the immune signature is present in tumors from each species. Strikingly, we observed that the immune signature in both mouse and human tumors was most highly evident in those with the lowest proliferative capacity. We directly assessed this relationship in human retinoblastoma tumors by co-analyzing proliferation and immune cell recruitment by immunohistochemistry, uncovering a significant inverse relationship between increased immune-cell infiltration in tumors and reduced tumor cell proliferation. Directly inhibiting proliferation with a PI3K/mTOR inhibitor significantly increased the number of CD45+ immune cells in the retina. This work establishes an in vivo model for the rapid recruitment of immune cells to tumorigenic neural tissue.
Similar content being viewed by others
Introduction
Retinoblastoma is a rare pediatric tumor of the retina that develops within a narrow window during retinal tissue development [1]. Deep analysis of the human retinoblastoma genome indicates that loss of the RB1 tumor suppressor gene is by far the most significant driving mutation in this tumor [1,2,3]. The retinoblastoma protein (pRb) is a key regulator of E2F activity and as such, these models fail to arrest E2F transcriptional activity and proliferation at the proper time during development leaving it in a growth promoting state [4,5,6,7]. E2F transcription factors are critical inducers of DNA synthesis and cell cycle progression through their ability to coordinate, and integrate cell cycle events with the transcription machinery and high E2F activity is noted in diverse aggressive and advanced cancers [8,9,10]. E2Fs simultaneously activate the expression of pro-apoptotic genes, which eliminate cells that have lost function of pRB and proliferate outside of an appropriate developmental context [11,12,13,14,15,16,17,18]. E2F-driven apoptosis can be suppressed by trophic factors, which permeate developing tissue or by mutations in cancer pathways such as the phosphatidyl-inositol 3 kinase (PI3K), Pten, or p53 which control cell-intrinsic apoptosis [19,20,21,22,23,24,25,26].
A number of relevant mouse models have been generated to recapitulate retinoblastoma, and these have served as important tools to drive optimization of clinically viable strategies [19,20,21, 27,28,29,30,31]. Mouse models of retinoblastoma almost always inactivate the retinoblastoma tumor suppressor gene and one or both of the related pocket proteins Rbl1 (p107) or Rbl2 (p130), since unlike for the human disease, Rb1 loss alone is insufficient to drive tumorigenesis in murine retina [27, 30, 32, 33]. Deregulated E2F activity causes massive apoptosis that eliminates most retinal cells; however the few that remain are highly prone to late-onset uni-lateral and bi-lateral tumors [34]. The remaining cells later display significantly higher unilateral and bilateral retinoblastoma emergence than control, indicating additional mutations picked up late may contribute to tumor initiation [27, 30, 33]. Direct co-alteration of members of the p53 pro-apoptotic target, including Cdkn2b (p19ARF), Mdm2, Mdm4, or Tp53 itself, accelerate tumor formation in these models [19,20,21, 30].
We utilized several mouse models, ranging from normal development (wild-type), late-onset retinoblastoma, all the way to rapid, early-onset bilateral retinoblastoma to investigate early events during retinoblastoma tumor initiation. Each of these models utilized the Vsx2 (formerly called Chx10) promoter to drive an expression of a CRE:GFP fusion protein (our control, hereafter referred to as VSX2-CRE) in retinal progenitor cells [35]. The previously described, late-onset retinoblastoma model (Vsx2-Rb1flfl;p107−/−) was also used [34]. This model exhibits high levels of E2F-induced apoptosis and greatly reduced retinal cell number by P30. The third model we used co-deleted the Pten tumor-suppressor gene with Rb1 and p107. Although PTEN is not mutated in human retinoblastoma, PTEN regulates retinal progenitor cell (RPC) proliferation and differentiation in vivo and can be a potent neural tumor suppressor [22, 36,37,38,39]. Pten co-deletion blocks E2F-driven apoptosis in this context, and induces rapid fully penetrant metastatic bilateral retinoblastoma emergence [31]. The retinoblastomas that develop in ΔRb/p107/Pten-deficient retinae are highly similar to other related murine retinoblastoma models in that they express primarily interneuron, but not retinal progenitor cells (RPC)-related markers [30, 31].
We used these models to compare gene expression differences from early, age-matched retina tissue that develop normally (control), a late-onset model of tumor formation, or a model that we developed and which rapidly generates bilateral tumors. We observed that both retinoblastoma tumor models are highly similar to normally proliferating RPCs, but they later fail to express differentiation markers as control retinal cells do. Instead, purified bilateral tumor cells express an immune gene expression signature not shared by control or the late-onset tumor cells. In expression profiles derived from bulk retinoblastoma tumors, this gene expression pattern was co-expressed with cluster of transcripts which define infiltrating immune cells. We utilized a dimensional reduction approach called Gene Cluster Expression Summary Scoring (GCESS) to observe a strong negative correlation between the immune cluster and the cell cycle cluster in our purified tumor cells and in both human and mouse bulk tumors, which we further validated by immunohistochemistry in human tumors. To determine whether cell cycle inhibition could lead to increased immune infiltration, we treated the early retinoblastoma model with cell cycle inhibitors and observed increased immune infiltration.
Materials and methods
Experimental animals
Vsx2-Cre mice [35], Rb1lox/lox mice [5], p107−/− mice [40], and Ptenlox/lox mice [41] and related PCR genotyping protocols have been described [31, 42]. All mouse experiments were performed in accordance with University of Minnesota Institutional Animal Care and Use Committee procedures and guidelines. BEZ235, a dual PI3K/mTOR inhibitor, was administered as described in the ref. [26]. Briefly, vehicle or BEZ235 was administered by oral gavage daily (50 µg/kg) on post-natal days 2–6 (P2–P6) and 22–26 and retinae harvested at P30 for histological analysis.
Retinal cell sorting and RNA isolation
Mice of the appropriate genotype and age were sacrificed and retinal tissue harvested. Each contained the Vsx2-Cre:GFP transgene. Retinal cells were trypsinized into single cell mixtures and sorted into GFP+ and GFP− populations. These cells were retested by flow cytometry after cell sorting to assess purity of GFP+ and GFP− populations (Fig. S1). RNA was isolated from GFP+ cell populations and used for library generation for RNA-seq.
RNA-sequencing (RNA-Seq)
Sequencing libraries were prepared from each sample using the TruSeq Library Preparation Kit (Illumina, San Diego, CA). Paired end sequencing (30–40 million reads per sample) was done at the University of Minnesota Genomics Center (UMGC) on a High Seq 2000 (Illumina, San Diego, CA, USA). The raw FASTQ files are available at the NCBI Sequence Read Archive and linked to from Gene Expression Omnibus SuperSeries GSE. Briefly, mapping was carried out using Tophat, Samtools and Cufflinks to generate FPKM values using the MM10 genome [43, 44]. To minimize the effects of dividing FPKM values by numbers close to 0 and stochastic noise, 0.1 was added to each FPKM value [45, 46]. The FPKM files are available at NCBI GEO (GSE162344). Mapping statistics are summarized in Supplemental Table S1.
RNA-Seq transcriptomic and GCESS analysis
GCESS is a method to systematically identify similar gene clusters across multiple complex transcriptional datasets [47, 48]. Transcriptional profile datasets are separately log transformed and mean centered, filtered for highly variant genes, and clustered using average linkage hierarchical clustering using Pearson correlation as the metric. The resulting trees are parsed to identify gene clusters with node correlation greater than a given value and greater than a given number of genes. The identified cluster members are then statistically compared across datasets to identify clusters, that are enriched for common genes present greater than expected by random chance using the Fisher exact test. Using this method allows facile identification of conserved transcriptional patterns observed across complex datasets without prior biological knowledge. From this dataset GCESS scores were calculated using correlation 0.20 and 50 genes selected, ten clusters were obtained. From GSE87043, eight retinal development time points from embryonic day 17.5 (E17.5) up to P21 were obtained for analysis. Genes with standard deviation (SD) > 0.6 were selected, GCESS run with correlation 0.20 and 50 genes selected, and three clusters were obtained. From GSE29683, Affymetrix array data from 62 human retinoblastoma tumors, genes with SD > 1.0 were selected. GCESS run with correlation 0.20 and 50 genes selected, and six clusters were obtained. From GSE29685 Affymetrix array, data from 132 mouse GEMM tumors was selected; genes with SD > 0.7 were selected. GCESS run with correlation 0.20 and 50 genes selected, and 12 clusters were obtained. The FISHER.pl scripts were used to identify significant overlap between clusters from our dataset and GSE87043, GSE29683, and GSE29685. We performed pathway and upstream regulator analyses using QIAGEN’s Ingenuity® Pathway Analysis (IPA®, QIAGEN Redwood City, www.qiagen.com/ingenuity) tools and protein–protein interaction (PPI) network analysis. This software predicts upstream molecules, such as microRNA, transcription factors, or chemicals, which cause similar gene expression changes elsewhere and thus may be causing the observed gene expression changes. Principle component analysis (PCA) and t-distributed Stochastic Neighbor Embedding (tSNE) analyses were carried out using the R programing language using the prcomp function, the Rtsne and and ggplot2 libraries on mean centered, log-base2 transformed datasets.
Immunofluorescence, immunohistochemistry, and immunoblotting
Unfixed murine control and tumor eyes were prepared for sectioning by freezing in optimal cutting temperature (OCT, Sakura Finetek USA, Torrance, CA). Tissue was sectioned at 6 μm on a cryostat and placed on positively charged glass slides. Tissue sections were fixed in acetone for 5 min, rehydrated in 1× PBS and blocked with 10% normal donkey serum (Jackson Immunoresearch, West Grove, PA). Avidin/Biotin blocking kit was applied per company instructions on all sections, that used a biotinylated antibody (Vector Labs, Burlingame, CA) followed by blocking with 10% normal donkey serum for 1 h (Jackson Immunoresearch, West Grove, PA). Primary antibodies CD11c biotin 1:100, CD11b FITC 1:100 (BD Biosciences, San Jose, CA) F4/80 FITC 1:100 (eBioscience, San Diego, CA) and CD3 1:400 (EMD Millipore, Burlington, MA) were applied for 1 h. Corresponding secondary antibodies were used for the unconjugated antibodies donkey anti-rabbit cy3 1:500 and streptavidin Cy3 1:500 (Jackson Immunoresearch, West Grove, PA) were applied for 1 h. All incubations were done at room temperature. Slides were coverslipped with hard set DAPI, 4,6-diamidino-2-phenylindole, (Vector Labs, Burlingame, CA) and examined by confocal fluorescence microscopy (Olympus BX61, Olympus Optical, Tokyo, Japan). Isotype negative control antibodies did not detect background staining (Supplemental Fig. S4). CD45 antibody (Clone 30-F11, BD Biosciences) was used at a 1:100 titer for IHC. IHC detection used Vector biotinylated secondary (1:250), tertiary was streptavidin-horseradish peroxidase (Covance #SIG-32000) and chromagen 3,3′-diaminobenzidine substrate (Covance #SIG-31043). Human retinoblastoma tissue microarray were obtained from US Biomax (#BC35111a). H&E staining, Ki67 (Sp6 Biocare medical, #CRM325) & TUNEL (EMD Millipore, #s7101), and Caspase-3 (Cell Signaling Technology, #9661) IHC staining were performed by the BIONET shared resource [14]. Immunoblotting was performed as previously described [49].
Results
Immunohistochemistry (IHC) of model states
To explore mechanisms associated with early tumor formation in the retina, we examined retinae from P0 and P7 for H&E staining, Ki67 IHC to detect proliferation and Caspase-3 IHC to detect apoptotic cells. Retinae from P0 control mice display high levels of Ki67+ proliferative cells in the developing neuroblastic layer with very few apoptotic cells at P0 (Fig. 1A). By P7, nearly all proliferation had ceased in control retinae, and modest levels of apoptotic cells were detected in the inner nuclear layer (INL). Hematoxylin and eosin (H&E) staining displays normal retinal development at P30.
Immunohistochemical and principal component (PC) analysis indicate that early retinoblastoma tumors are highly similar to normal retinal progenitor cells. A Post-natal day 0 (P0) and P7 retinae were collected from control (Vsx2+), Vsx2+; Rbfl/fl; p107−/− (late-onset retinoblastoma model), and from Vsx2+; Rbfl/fl; p107−/−; Ptenfl/fl (early-onset bilateral tumors). Sections were stained with hematoxylin and eosin (H&E), or by immunohistochemistry with antibodies that stain Ki67 to detect proliferation or active, cleaved caspase-3 (Casp3) to detect apoptosis. B Principle component analysis of all samples, displaying PC #1 and PC #2. Three retinae from P7 Vsx2+; Rbfl/fl (SKO) mice were also harvested for simultaneous analysis. Post-mitotic, differentiated control (P7) retinae separated from all proliferating samples on PC #1, with ΔRb showing an intermediate phenotype on this axis. ΔRb/p107/Pten-deficient P7 retinae were the only samples separated from all other samples, particularly control, on PC #2. The individual genotypes are represented by color and the timepoints are represented by shape for the control Vsx2+ (CHX), Vsx2+; Rbfl/fl (SKO), Vsx2+; Rbfl/fl; p107−/− (DKO), and from Vsx2+; Rbfl/fl; p107−/−; Ptenfl/fl (TKO) sorted retinal tissues.
ΔRb/p107-deficient retinae combines a conditional deletion of the retinoblastoma tumor suppressor gene (Rb1) driven by Vsx2-Cre:GFP with a germline deletion of p107 (p107−/−) [30, 31, 34]. These retinae express high levels of the E2F transcription factors owing to deletion of both restrictive pocket proteins pRB and p107. ΔRb/p107-deficient retinae are abnormally disorganized and display Ki67+ proliferating cells with low apoptosis. However, in contrast to control retinae, ΔRb/p107-deficient retinal cells remain highly proliferative and apoptotic at P7. By P30, excess apoptosis has eliminated most retinal cells. But the remaining cells are highly susceptible to increased late-onset, unilateral, and bilateral retinoblastoma tumor formation [30, 33].
Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox mice conditionally co-deletes Pten in the Vsx2-Cre+; Rb1lox/lox; p107−/− background. Pten is a tumor suppressor gene in neural and other tissue types, and its deletion in the retina exerts strong anti-apoptotic effects and differentiation defects [23, 38, 50, 51]. Like other P0 retinae, the ΔRb/p107/Pten-deficient retinae are also highly proliferative with minimal apoptosis. This proliferation remains high at P7 but Pten-deletion significantly reduced the high-level apoptosis observed in the ΔRb/p107 retinae. Deregulated proliferation combined with suppression of apoptosis resulted in large, bilateral retinoblastoma tumors that filled the ocular cavity by P30 (Fig. 1A). The consequences of altered proliferation and apoptotic phenotypes in these models are clear by P30. Control retina look normal, the ΔRb/p107-deficient (center) retina has lost most retinal cells owing to apoptosis, and ΔRb/p107/Pten-deficient mice develop aggressive bilateral tumors.
RNA-seq analyses
To uncover transcriptional differences between these early stage retina states, we used fluorescence-activated cell sorting to collect CRE::GFP positive cells at P0 and P7 from retinae of each of these genotypes, isolated RNA and performed RNA-sequencing using Illumina Hi-Seq. We also isolated RNA from Vsx2-Cre; Rb1fl/fl (ΔRb) at P7 as a putative intermediate between the control and ΔRb/p107 genotypes. Sorting allowed identification of a pure retinal (normal/tumor) cell transcription, minimizing signal derived from other cell types in the tumor microenvironment. GFP+ FACS sorted retinal cells averaged over 97% purity when reanalyzed after the sort (Fig. S1). Following mapping to the MM10 genome (Supplemental Table S1), we were able to directly validate our genotypes by visualizing the engineered constructs using the Integrative Genomics Viewer (IGV) [52]. Genotype visualization confirmed that our cell population was pure: almost all cells in our sequenced population had undergone Cre-mediated recombination at the expected locations (Fig. S2).
Fragments per kilobase of transcript per million mapped reads (FPKM) values were generated based on the mapped files, and principal component analysis (PCA) was used to compare the samples based on genotype and developmental timepoint [53]. The control P7 retinae were most separated from a cloud of the remaining tissues on PC#1 (Fig. 1B). The Vsx2-Cre; Rb1fl/fl (ΔRb) P7 retinae were intermediate on the principle component (PC) #1 spectrum between the non-proliferative P7 CHX and the proliferating P7 ΔRb/p107 samples. This indicates that the additional loss of p107 further separated ΔRb/p107 samples away from ΔRb and the growth-arrested control (P7) on this axis. Remarkably, proliferating P0 control retinae clustered with both the ΔRb/p107-deficient and ΔRb/p107/Pten-deficient samples on the PC#1 axis. Importantly, the P7 ΔRb/p107/Pten-deficient tumors separated from all other samples on the PC#2 axis. Overall, these data indicate that as retinoblastoma tumors initiate at P0 they are remarkably similar to proliferating RPCs, but later acquire a unique expression component by P7 not observable in any of the other genotypes.
The dataset was log transformed and mean centered. From the ~24,000 starting transcripts, 4044 showed a log base 2 change of absolute value greater than 2 across the sample set. These genes were selected and used for hierarchical gene clustering (Fig. 2). We also analyzed each gene cluster using Ingenuity Pathway Analysis (IPA) software to deduce enriched functions in each cluster (Fig. S3). This clustering revealed five major, distinct clusters. We observed a pair of clusters (#1 and #4) with inversely expressed genes whose expression did not depend on underlying genotype (Fig. 2). Cluster #1 contained genes that were expressed to a higher degree at P0 than at P7, whereas genes in cluster #4 were uniformly high at P7 and low at P0. Genes in cluster #1, which we named “Progenitor”, scored the highest for gene components involved in the hedgehog signaling pathway such as smoothened (1e−9), GLI3 (1e−6), and sonic hedgehog (SHH) (1e−6) and the “positive regulation of cell proliferation” gene annotation ranked highest for this set (7.6e−5) (Fig. S3). The hedgehog pathway transmits signaling information critical for proper retinal cell differentiation [54]. Transcripts within cluster #4, named “Mature”, were inversely expressed compared to transcripts in cluster #1, containing genes with the highest expression at P7 regardless of genotype. IPA analysis of the “mature” retinal cluster indicated enrichment of genes in categories such as cell adhesion; 7e−6, cell differentiation; 2e−4, and nervous system development 2e−4 (Fig. S3).
Four thousand and transcripts identified by RNA-sequencing were log transformed, mean centered and used for hierarchical clustering of gene transcripts. Five unique clusters were identified and labeled progenitor, cell cycle, immune, mature, and differentiation to categorize them. Relative expression levels of transcripts is denoted by the blue (low) to yellow (high) color code.
A second set of clusters (#2 and #5) also showed inverse expression patterns. Cluster #2 is comprised of genes that are highly expressed in proliferating retinae but not in the post-mitotic P7 control retinae. Cluster #2 highly expressing samples included control P0 proliferating retina, and both P0 and P7 ΔRb/p107-deficient and ΔRb/p107/Pten-deficient retinae. P7 ΔRb samples were intermediate between P7 control and ΔRb/p107 samples. Cluster #2, was named “Cell Cycle” because it primarily contains cell-cycle related E2F transcriptional targets that are highly expressed in proliferating retina with GO annotations, such as cell cycle (p = 10−87) and cell division (p = 10−60) (Fig. S3). Conversely, the expression pattern of Cluster #5, called “Differentiation” was generally opposite that of Cluster #2, being most highly expressed in post-mitotic P7 control retinae but low in each of the proliferating retinal samples. IPA analysis of Cluster #5 scored highest for retinal transcriptional regulators that control differentiation such as Crx (3e−12), Rest (1e−13), and Nrl (6e−12) by “Upstream regulator” analysis, and visual perception (10−37) by gene annotation [55,56,57]. Thus, cluster #2 and #5 appear to define a proliferation to differentiation axis. This axis is strongly shifted towards proliferation in both the ΔRb/p107-deficient and ΔRb/p107/Pten-deficient samples, with ΔRb-deficient samples showing an intermediate phenotype.
Finally, cluster #3 was expressed solely in P7 ΔRb/p107/Pten-deficient tumor samples. Genes in this cluster were most highly related to immune response-related annotations such as interferon gamma; 4e−45, lipopolysaccharide; 4e−39, interleukin-10; 1e−26, and interleukin-6 treatment; 1e−25. Several of the individual genes located in cluster #3, such as Ccl2, Ccl3, Ccl4, and Cxcl2, are well known immune cell chemoattractants (p < 0.05) [58,59,60]. Intriguingly, these genes were not notably induced in the ΔRb/p107-deficient, late-onset retinoblastoma tumor cells. ΔRb/p107/Pten-deficient tumor samples showed some variability in the intensity of gene expression in cluster #3, with three out of eight samples more highly expressing them than the other five, although each were notably stronger than observed in other clusters. Together, these findings suggest that the ΔRb/p107/Pten-deficient early-onset retinoblastoma tumors express a robust immune gene expression signature not shared by the ΔRb/p107-deficient apoptotic retinae.
Bilateral tumors accumulate immune cells, but late-onset and control retinae don’t
It was surprising to detect an immune response gene enrichment signature in the early-onset bilateral tumors, because the RNA had been isolated with high purity from Cre:GFP+ sorted retinal tumor cells rather than from bulk tumors, which might be expected to also contain immune cells. We determined if this signature was associated with immune cell accumulation within the ΔRb/p107/Pten-deficient samples relative to all other samples. ΔRb/p107/Pten-deficient retinoblastoma tumors were assessed over a timecourse for the presence the CD45+ marker (protein tyrosine phosphatase receptor type C), a transmembrane protein present on most immune cell types (Fig. 3A) [61]. We noted large numbers of CD45+ cells in the center of the retina at P7. CD45+ cells were qualitatively reduced over the time course as the tumor continued to fill the ocular cavity. By P30, CD45+ cells were reduced in number and by P60, only small groups of CD45+ cells were observed. We used flow cytometry to quantify the total number of CD45+ cells in P15 control retinae and ΔRb/p107/Pten-deficient bulk retinoblastoma tumors (Fig. 3B). Control retinae contained around 1% CD45+ cells, equivalent to numbers reported previously [62]. The total number of CD45+ cells present in bulk ΔRb/p107/Pten tumors was around 8%. To gain further insight into the origins of the CD45+ cells in the tumors, we co-analyzed for GFP expression along with CD45 by flow cytometry. Tumor and immune cells derived from retinal progenitor cells were exposed to Cre:GFP fusion protein, whereas external immune cells were not. We found that ~93% of CD45+ immune cells isolated from bulk retinoblastoma tumors were GFP−, and only 7% were both CD45+ and GFP+. This suggests that the total immune cell composition in these retinoblastomas derive primarily from outside the retina, although CD45+ cells that originate from within the retina, such as Müller glia cells, comprise a small but significant portion of total tumor resident immune cells. Further cell fate lineage tracing will be required to fully and accurately define the origins of the immune cells [63]. ΔRb/p107-deficient retinae undergo significant E2F-induced cell death, significantly altering the retinal architecture by P30, however these retinae failed to express the immune signature. To determine if they also failed to recruit CD45+ immune cells, mouse control retina, ΔRb/p107-deficient, and ΔRb/p107/Pten-deficient retinoblastoma tumors were analyzed by H&E, and IHC for TUNEL as a marker of cell death, Ki67 to detect proliferating cells, and CD45+ cells (Fig. 3C). Control (P15) and ΔRb/p107-deficient retinae showed no CD45+ reactive cells, despite displaying augmented proliferation, but we observed high numbers of CD45+ cells within proliferating ΔRb/p107/Pten-deficient retinoblastoma tumors.
A Retinae were harvested from P7, P15, P30 and P60 Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox mice and stained by IHC for CD45+ cells. B Fifteen-day old control and Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox tumor samples were isolated, trypsinized to single cells, processed for flow cytometry to quantify CD45+ cell numbers, and graphed as percent of total cells. C Cells (P15) from 3B were further analyzed for GFP by flow cytometry to detect Cre:GFP+/CD45+ cells native to retina, or Cre:GFP−/CD45+ cells external to the retina, and graphed as the percent of total cells. D Retinae were harvested from P15 control, Vsx2-Cre+; Rb1lox/lox; p107−/− and Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox and stained by H&E, or stained by IHC to detect apoptotic cells by TUNEL, proliferation by Ki67, and CD45+ immune cells. E The numbers of CD45+ cells detected by IHC were counted and graphed as the % CD45+ cells compared to total. *p < 0.05; **p < 0.01; ***p < 0.001.
We next used immunofluorescence to identify the immune cell composition present in retinoblastoma tumors. Three P15 control retina and three P15 ΔRb/p107/Pten-deficient tumors were harvested and frozen in optimal cutting media (OCT) for sectioning and immunofluorescence analysis. Retinae were tested for the presence of dendritic cells (α-CD11c), monocytes (α-CD11b), tumor-associated macrophages (TAMs) (α-F4/80), and T-lymphocytes (CD3). We observed each of these immune cell types in P15 tumor sections but not control retinae (Fig. 4). CD11c+ dendritic cells were present in 3/3 of the tumors and comprised slightly under 2% of the retinal bulk tumor cell population (Fig. 4B). CD11b+ monocytes and CD3+ T cells were present in tumors #2 and #3 but not #1, and CD3+ cells comprised around 2% of total cells, and CD11b+ around 4%, in the tumor population (Fig. 4B). F4/80+ TAMs were only present in tumor #3 at around 2% of the total tumor cell population. Natural killer (NK) cells were not detected with α-NK1.1 antibodies (not shown). Isotype negative control antibodies did not detect background staining in either control or tumor samples (Supplemental Fig. S4). Immune cell presence is thus also a variable phenotype and ranged from one extreme, with 4/4 immune cell types in tumor #3, to the other extreme, with only 1/4 immune cell types represented in tumor #1. These studies indicate that ΔRb/p107/Pten-deficient retinoblastoma tumors uniquely express an immune gene expression signature and also similarly accumulate CD45+ cells comprised of dendritic cells, monocytes, macrophages, and T-lymphocytes.
A P15 control or Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox tumor samples were processed for H&E staining and immunofluorescence to detection of dendritic cells (α-CD11c), monocytes (α-CD11b), tumor associated macrophages (α-F4/80), and T-lymphocyte immune cells (α-CD3). Isotype-matched negative control antibodies are shown in Supplemental Figure S4 and did not detect background staining. B Dendritic cells (α-CD11c), monocytes (α-CD11b), tumor associated macrophages (α-F4/80), and T-lymphocyte immune cells (α-CD3) were counted and graphed as the % cell type against the total cell population (n = 3 control and tumor retinae each).
Greater immune gene expression in less proliferative human and mouse retinoblastomas
Our data indicates that both late-onset and early-onset bilateral tumor prone cells fail to properly express normal retinal differentiation transcripts compared to control retinae. We used a dimensional reduction technique called GCESS [47, 48] to systematically compare the transcriptional patterns present in our dataset with a recently published retinal developmental RNA-Seq timecourse (GSE87043) (Fig. 5A, B) [64]. GCESS identifies potentially related gene clusters across multiple complex transcriptional datasets, and performs direct statistical comparison to assess the significance of the findings. GCESS analyses showed that each of the five clusters that we identified in our control and retinoblastoma RNA-seq data (Fig. 5A) were each present within the retinal developmental timecourse (Fig. 5B). These clusters fell into two main categories. First, the progenitor (blue) and cell cycle (red) signatures were highly represented in proliferating retinae from embryonic day 14.5 (E14.5), E17.5, and P0. Both of these signatures were lost between P0 and P3 as retinal proliferation ceased and differentiation commenced. In contrast, the other three clusters, which included differentiation (green), “mature” retina (orange) and “immune” (yellow) clusters, were low in proliferating progenitors but accumulated at P3 and persisted through later timepoints when proliferation ceased and differentiation occurred. These analyses further confirmed the usefulness of GCESS to successfully identifying like-clusters in related datasets [47, 48]. They also confirmed that many transcriptional features associated with normal development were delayed or missing altogether in the early-onset and late-onset tumor models.
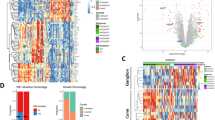
Comparison of our identified clusters with those from several other mouse developmental or mouse and human retinoblastoma studies. A Clusters of genes identified in our RNA-Seq transcriptional data are displayed as heatmaps with yellow showing increased transcription and blue decreased transcript levels. B Gene expression pattern comparisons with a mouse retinal development timecourse (GSE87043). C A human retinoblastoma dataset (GSE29683)  -Y79 and Weri1 human retinoblastoma cell lines;
-Y79 and Weri1 human retinoblastoma cell lines;  -human patient derived xenograft samples and D a mouse retinoblastoma dataset (GSE29685). Each of the five clusters identified by our RNA-seq analyses are marked with color coded sidebars to the right of the heatmap, and their identifiers are shown below.
-human patient derived xenograft samples and D a mouse retinoblastoma dataset (GSE29685). Each of the five clusters identified by our RNA-seq analyses are marked with color coded sidebars to the right of the heatmap, and their identifiers are shown below.  -ΔRb/p107/p130-deficient murine tumors. Color-coded clusters in related cross-species databases denote shared genes at a highly significant level (p < 1e−5).
-ΔRb/p107/p130-deficient murine tumors. Color-coded clusters in related cross-species databases denote shared genes at a highly significant level (p < 1e−5).
Several different mouse models have been used to form retinoblastoma tumors. We wondered if the gene expression patterns we identified, particularly the immune-related expression signature, were unique to our model or if they were also detectable in other mouse and human retinoblastomas with related gene expression analysis [30]. Array based datasets from bulk retinoblastoma (mouse; GSE29685) and (human; GSE29683) were analyzed against each other as well as our gene expression data using GCESS to identify significantly-related clusters. Cell cycle, immune, and differentiation state transcripts corresponding with our data were clearly and significantly (p < 1e−10) identified in both mouse and human retinoblastoma tumors (Fig. 5C, D).
The cell cycle and differentiation cluster are relatively straightforward to interpret and indicate that the variation in our dataset are directly relevant to both human and mouse retinoblastoma tumors. The situation with the immune cluster is more complex. Much of the observable immune transcript cluster was derived from transcripts likely expressed from infiltrating immune cells present in the bulk tumors analyzed, but were not present in our RNA-Seq dataset because it is made up of highly purified retinal cells. Further evidence for this hypothesis is that many more immune-infiltrate transcripts are present in the immune cluster from the other human and murine bulk tumor datasets (Fig. 5C, D).
The immune infiltrate signature was expressed at the lowest levels in gene expression profiles derived from the highly proliferative Y79 and Weri1 human retinoblastoma cell lines, due to tissue culture conditions, indicating that this signature primarily originates within tumors in vivo (see  -mark, Fig. 5C). The infiltrating immune signature was also very low in human xenograft tumors serially passaged in immunocompromised mice, perhaps due to the failure of the human-based array to identify the mouse immune cell transcripts or because the human cells were grown in mice without a resident immune system (see
-mark, Fig. 5C). The infiltrating immune signature was also very low in human xenograft tumors serially passaged in immunocompromised mice, perhaps due to the failure of the human-based array to identify the mouse immune cell transcripts or because the human cells were grown in mice without a resident immune system (see  -mark, Fig. 5C).
-mark, Fig. 5C).
Intriguingly, the “progenitor” cluster was not significantly detected in human retinoblastomas, and was only represented in ΔRb/p107/p130-deficient murine tumors, which lacks all three pocket proteins (see  -mark, Fig. 5D) [30]. This further supports the model that retinoblastoma tumor emergence is more tightly associated with the loss of key retinal differentiation genes, rather than the acquisition or retention of gene expression tied to an immature progenitor state, as has been suggested [64].
-mark, Fig. 5D) [30]. This further supports the model that retinoblastoma tumor emergence is more tightly associated with the loss of key retinal differentiation genes, rather than the acquisition or retention of gene expression tied to an immature progenitor state, as has been suggested [64].
Strong negative correlation between the cell cycle and the immune-modulatory transcripts in bulk human and mouse tumors
Expression of E2F cell cycle transcripts is generally a feature of early development that is also maintained in early-onset and late-onset tumor states. But these cell cycle transcripts are lost in normal, older tissues that have increased transcripts with a role in retinal cell differentiation. Differentiation and proliferation are generally thought to be mutually exclusive, but it was also reported that proliferating retinoblastoma tumors can simultaneously retain expression of transcripts associated with differentiation of multiple retinal cell types [64]. The inter-relationship between proliferative and immune transcripts in tumors is less well defined. The GCESS method allows calculation of a single summary score for each tumor representative of the transcriptional cluster. This technique allows an unbiased dimensional reduction of transcriptional datasets allowing meaningful intratumor statistical analyses to be conducted. To further examine the association between proliferation, differentiation, and immune transcripts within individual tumors, we used GCESS to calculate scores for each of these three states across individual samples within our dataset, and across individual tumors within the mouse and human retinoblastomas datasets.
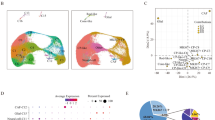
We first compared the GCESS scores for the cell cycle transcripts to the differentiation score within individual samples to determine if a statistically significant correlation existed. We observed a clear negative correlation across our full dataset [−0.96] (Fig. 6A). This negative correlation was also noted during analysis of only the ΔRb/p107/Pten-deficient tumors [−0.95] (Fig. 6B) and the mouse retinal developmental timecourse (GSE87043) [−0.9] (Fig. 6C). We saw further evidence for this negative correlation in the mouse retinoblastoma datasets (−0.39) (Fig. 6D), but not in the human tumor retinoblastoma datasets [0.41] (Fig. 6E). Surprisingly, we also observed a strong negative correlation between the cell cycle and the immune GCESS scores [−0.78] within our early-onset ΔRb/p107/Pten-deficient tumor samples at P7 (Fig. 6F). To determine if this negative correlation between cell cycle and immune transcripts was a fundamental feature of retinoblastoma, we again used GCESS to examine the bulk human and mouse tumors. Importantly, this negative correlation was clearly observable and statistically significant in both the mouse [−0.65] and human [−0.79] tumor datasets (Fig. 6G, H).
Analysis of cell-cycle, differentiation, and immune Gene Cluster Expression Summary Scoring (GCESS) scores to determine intra-tumor gene expression pattern relationships in related cross-species developmental and retinoblastoma datasets. Cell cycle versus Differentiation GCESS scores were compared within A our full dataset, B only the early-onset tumors from our dataset, C the mouse retinal developmental dataset (GSE87043), D mouse RB tumors (GSE29685), and E human RB tumors (GSE29683). Intratumor comparison between Cell cycle and immune GCESS scores for F early-onset tumors from our dataset, G mouse RB tumors (GSE29685), and H human RB tumors (GSE29683).
Cell cycle and immune transcripts negatively correlate in human retinoblastoma
We directly tested if cell cycle and immune infiltration are also inversely correlated at the protein level in human retinoblastoma. We used IHC to stain human retinoblastoma samples (n = 14) in duplicate for Ki67 to identify dividing cells, and for CD45 to detect immune infiltration, and show five representative examples (Fig. 7A). None of the tumors were treated with chemotherapy before removal, which could have affected proliferation and immune cell levels. Tumor #1 was highly positive for Ki67 with few CD45+ cells. Ki67 levels are reduced in tumor #2 and #3, and increased numbers of CD45+ cells are noted. Tumors #4 and #5 show regions heterogeneous for Ki67 and CD45. #4 is mostly Ki67+, but a large Ki67− region in the tumor has acquired CD45+ cells in a serial section. Similarly, tumor #5 also shows several regions in serial sections with high numbers of Ki67+ and few CD45+ cells, or vice versa.
Human retinoblastoma samples display a significant inverse relationship between proliferation and immune cell infiltration. A Fourteen human retinoblastoma samples on a tissue microarray were stained simultaneously by immunohistochemistry for the proliferation marker Ki67 or for CD45, the “leukocyte common antigen” to detect immune cells. Five representative samples are shown: two have high Ki67 and low CD45+ cells, and two show the reverse phenotype. B Each sample was scored based on the number of Ki67 and CD45 positive cells. Tumors with greater than 50% Ki67+ cells were marked “high” and those with less than 50% as “low”. Tumors with greater than 5% immune cells were marked high, and less than 5% low. Scores for Ki67 and CD45 were compared against each other and found significant using Fisher’s exact test (p < 0.01).
Ki67 and CD45 levels were scored as “high” or “low” based on comparative levels across all samples and tested for statistical significance (Fig. 7B). The tumor cells with highest Ki67 levels (>50% cells Ki67 positive) were significantly associated with decreased CD45+ immune infiltration (less than 5% CD45 positive) (8/8). Tumors with decreased Ki67 staining (<50% cells positive) showed high levels of immune infiltration (more than 5% positive) (5/6). This inverse relationship between cell cycle and immune-related IHC was highly significant relative to random (Fisher’s exact test; p < 0.01) and overall is consistent with what we observed from the transcriptional data.
Increasing immune infiltration into tumors is a highly sought after clinical goal, because checkpoint inhibition is unlikely to succeed without it. Because of the noted inverse correlation between cell cycling and immune cell accumulation, we hypothesized that inhibiting proliferation via pharmaceutical treatment may lead to increased immune infiltration. PI3K and mTOR both become highly active following the loss of Pten in this model since PTEN protein is their natural negative regulator, so we tested BEZ235, a dual PI3K/mTOR inhibitor, for inhibiting proliferation [23, 50]. To test this, we treated five control and five Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox mice with BEZ235, a dual PI3K/mTOR inhibitor, by daily oral gavage on post-natal days 2–6 and 22–26 [26, 65]. Retinae from vehicle and BEZ235-treated mice were harvested at P30 for analysis by IHC (Fig. 8A). We observed decreased tumor cell proliferation via Ki67 staining and Ki67+ cell counts, which were reduced from ~17% positive cells on control to ~5% in BEZ235-treated retinae, indicating that the drug was working to decrease cell division (Fig. 8B). The BEZ235-treated tumors also showed increased CD45+ immune cell accumulation via IHC (Fig. 8A). CD45+ cell counts increased from 2.5% in control retinal to ~18% in tumors (Fig. 8B), indicating that the negative correlation between proliferation and immune cell accumulation may provide a therapeutic vulnerability. These results indicate that decreasing cell division via pharmaceutical treatment in retinoblastoma surprisingly can lead to increased immune infiltration, which may potentially then be activated by checkpoint inhibition therapy.
Vehicle or BEZ235, a dual PI3K/mTOR inhibitor, was administered to Vsx2-Cre+; Rb1lox/lox; p107−/−; Ptenlox/lox mice by oral gavage daily on post-natal days 2–6 and 22–26 and harvested at P30 for histological analysis. A IHC was used to detect Ki67 and CD45 positive cells. Dual PI3K/mTOR inhibition with BEZ235 inhibits tumor cell proliferation and increases CD45+ immune cells in the tumor. Results are representative of analysis of five mice from each group. B Ki67+ and CD45+ cells were counted from vehicle and BEZ235-treated retinae and graphed as the percent of positive cells (n = 5, each group). ***p < 0.001.
Discussion
Most adult tumors originate in fully developed tissue after a stepwise accumulation of multiple mutations or chromosomal breakages [66]. Pediatric tumors in contrast usually originate in developing tissue through the misappropriation of normal epigenetic and differentiation proteins, and as such, they often require a far lower mutational burden to elicit cancer [2, 42, 67,68,69,70,71]. This effectually limits the number of potential druggable targets in pediatric malignancy, even though many of the driver genetic alterations are conserved between adult and childhood cancers. Retinoblastoma, as an example, is frequently driven by the loss of only the Rb1 tumor suppressor gene, with biallelic loss of Rb1 serving as the key driver of aggressive, bilateral tumors [72]. Retinoblastoma has typically been treated by combinations of chemotherapeutic agents such as carboplatin, vincristine, and etoposide [73]. Early detection and improved treatment regimens have significantly improved survival rates of patients with retinoblastoma, but therapy still often results in enucleation, loss of vision in the affected eyes, and treatment is often also followed by refractory disease or co-morbidities [1]. It is also devastating if not detected early as metastatic disease is nearly 100% fatal. Thus, better models and treatments have the potential to eliminate the cancer while maintaining vision. Immunotherapies are having a major impact in oncology, but it is increasingly clear that effective immunomodulatory strategies depend on unleashing a sufficient baseline immune response [74]. The recent success of cancer immunotherapies also made clear that manipulating different forms of cell death can influence the subsequent immune response [75]. In this study, we investigated the immune response mounted against retinoblastoma, and this has potential therapeutic applications for both pediatric retinoblastoma and for modulating inflammation-related retinal degeneration.
A previous study explored the gene expression differences between six different mouse models of retinoblastoma, which included the Rb/p107-deficient model we included here [30]. Retinoblastoma bulk tumors from each of those models were subjected to gene expression, and also characterized by immunostaining for proliferation and macrophages and other markers [30]. Although macrophages were detected in each of these models, no significant differences were observed for Ki67 or macrophage numbers across tumors of distinct genotypes. The Rb/p107-deficient retinae did not produce significant numbers of immune cells at early timepoints in our hands. In contrast, co-deleting the Pten tumor suppressor gene in the context of Rb and p107 deficiency blocked E2F-dependent intrinsic apoptosis in retinal cells rapidly generated bilateral tumors [31], and accumulated a large number of adaptive and innate immune cells. These gene expression studies differed in several important regards. Our gene expression analysis was conducted in retinoblastoma tumors at early time-points, that fall within the retinal developmental window when tumors normally originate. These developmental timepoints picked up the expected differences in control cell gene expression states relating to proliferation and differentiation. This feature may be particularly relevant for retinoblastoma, which most often initiates during the timeframe of retinal cellular differentiation. Further, these timepoints also allowed for the temporal detection of the immune response at P7 but not P0. Ki67 scoring also may not fully recapitulate our “cell-cycle” signature and related GCESS score, which was comprised of many genes. Likewise, we noted accumulation of a variety of immune cell types, and macrophage cell numbers may not fully recapitulate our immune GCESS score. It seems unlikely that the increased immune cell accumulation in the Pten-deficient tumors was simply correlated with increased tumor burden, as even larger tumors observed at P60 and beyond did not have more immune cells than smaller tumors at P7 or P15.
The underlying mechanistic basis for the difference in immune cell recruitment between these models is not entirely clear. It is possible that the ∆Rb/p107/Pten genotype itself alters retinal gene expression to attract immune cells. ∆Rb/p107 deficiency activates E2F transcription factor function. The concept that different E2F functions and transcriptional outputs are regulated, and may be separable has not been extensively explored and has potentially important consequences for tumor suppression and immunity. For example, an E2F dependent “proliferation signature” is common in many different tumor types owing to inactivation of the retinoblastoma pathway. Pten deletion leads to unrestrained PI3K activity, and we previously demonstrated that PI3K signaling alters Rb/E2F-mediated gene expression to reduce pro-apoptotic target genes [25]. Further, an E2F/FOXO transcriptional complex promotes apoptotic gene expression and cell death in the retinal context, and this complex may be disrupted by AKT signaling downstream of Pten deficiency [31]. Rb deficiency also induces expression of CCL2 and other cytokines that modulate the tumor microenvironment [76, 77]. Pten deficiency also causes numerous cellular changes to differentiation, apoptosis, metabolism, and immunity [50, 78, 79]. Pten-deletion in melanoma cell lines also induces CCL2 and other cytokines but promotes resistance to immunotherapy in that context [80]. Similarly, direct interactions between E2F and NF-κB factors directly control the timing of expression of inflammatory-signaling gene targets [81]. pRb also binds specifically to the p65 subunit of the NF-κB protein to inhibit expression of the programmed death ligand-1 (PD-L1) to suppress cancer immunity [82]. These studies underscore the critical relationship between these pathways and immunity, and provide further focus and strategies for pharmacologic manipulation.
Alternatively, co-deleting the Pten gene may have blocked intrinsic apoptosis and caused cancer so rapidly and destructively, that primary retinal damage caused an inflammatory or immune response to deal with the damage. The observation that the immune signature was more strongly activated in some P7 tumors than others, and likewise that some tumors contained more immune cells support this idea. Different types of stress in normal retina and retinoblastomas have different effects on immune cell invasion. For example, treating retinoblastoma tumors with therapeutic adenoviral vectors led to few changes to the immune environment [83, 84]. Direct physical damage to primary retinal cells in vivo, as through damage with a laser, ischemia, or optic nerve crush, leads to a rapid immune response [85,86,87]. Different modes of cell death and signaling from dying cells also result in distinct effects on the nature of the subsequent immune response [88]. For example, rampant E2F activity following the loss of Rb and p107 caused a robust apoptotic programmed cell death (PCD) response with little early immune cell accumulation. We observed necrotic tissue in the central retina at P7 in bilateral tumors, and similar necrosis has been observed in other highly aggressive murine retinoblastoma models [89]. Necrosis was originally described as an alternate form of cell death caused after apoptosis is blocked, and causes cells to swell and rupture. Dying retinal cells release pathogen-associated or damage-associated molecular patterns (PAMPs/DAMPs) [90]. These trigger the production of cytokines that alert and recruit the immune system [91]. Forced induction of necroptosis, or programmed necrosis, capably synergizes with immune checkpoint blockade to kill tumor cells [75].
In summary, the results of this work are unexpected because a clear connection between reduced proliferation and increased immune infiltration, or suppression of immune infiltration in highly proliferative tumor has not been identified. By directly testing this model in vivo, we found that decreasing proliferation by targeted therapy with BEZ235 increased immune cell accumulation. These data leads to several pertinent follow-up questions. For example, does immune ablation or augmentation change tumor cell proliferative capacity? Do agents that promote differentiation also affect immune response directly or indirectly by altering proliferation? Differentiation therapy may be useful for two different rationalizations. It may force tumor cells to stop dividing via the Rb/E2F cell cycle signaling pathway and reduce tumor cell proliferation, indirectly increasing immune cell infiltration. However differentiation therapy may also decrease Rb/E2F transcriptional activity and reduce expression of immune-modulatory targets [81]. The observation that inhibition of tumor cell proliferation strongly associates with increased immune cell presence has important implications for anti-tumor therapies. These results indicate that utilization of such approaches to activate the immune system may have anti-tumor utility in retinoblastoma. Inflammatory pathways can be chronically overactive in the retina, causing a variety of human diseases linked to inflammation and retinal cell degeneration. These studies will further elucidate how apoptotic and immune signaling integrate crosstalk with E2F-dependent transcriptional output, and may lead to development of strategies that manipulate the immune responses to influence retinal inflammation, immune cell recruitment, and retinal cell proliferation.
Data availability
Related datasets are available at NCBI GEO (GSE162344).
References
Dimaras H, Corson TW, Cobrinik D, White A, Zhao J, Munier FL, et al. Retinoblastoma. Nat Rev Dis Prim. 2015;1:15021.
Dyer MA. Lessons from retinoblastoma: implications for cancer, development, evolution, and regenerative medicine. Trends Mol Med. 2016;22:863–76.
Rushlow DE, Mol BM, Kennett JY, Yee S, Pajovic S, Theriault BL, et al. Characterisation of retinoblastomas without RB1 mutations: genomic, gene expression, and clinical studies. Lancet Oncol. 2013;14:327–34.
Zamanian M, La Thangue NB. Adenovirus E1a prevents the retinoblastoma gene product from repressing the activity of a cellular transcription factor. EMBO J. 1992;11:2603–10.
Sage J, Miller AL, Perez-Mancera PA, Wysocki JM, Jacks T. Acute mutation of retinoblastoma gene function is sufficient for cell cycle re-entry. Nature. 2003;424:223–8.
Dyson NJ. RB1: a prototype tumor suppressor and an enigma. Genes Dev. 2016;30:1492–502.
Dick FA, Goodrich DW, Sage J, Dyson NJ. Non-canonical functions of the RB protein in cancer. Nat Rev Cancer. 2018;18:442–51.
Nevins JR. Toward an understanding of the functional complexity of the E2F and Retinoblastoma families. Cell Growth Diff. 1998;9:585–93.
Dimova DK, Dyson NJ. The E2F transcriptional network: old acquaintances with new faces. Oncogene. 2005;24:2810–26.
van den Heuvel S, Dyson N. Conserved functions of the pRB and E2F families. Nat Rev Mol Cell Biol. 2008;9:713–24.
DeGregori J, Leone G, Miron A, Jakoi L, Nevins JR. Distinct roles for E2F proteins in cell growth control and apoptosis. Proc Natl Acad Sci USA. 1997;94:7245–50.
Iaquinta PJ, Lees JA. Life and death decisions by the E2F transcription factors. Curr Opin Cell Biol. 2007;19:649–57.
Lu H, Hallstrom TC. The nuclear protein UHRF2 is a direct target of the transcription factor E2F1 in the induction of apoptosis. J Biol Chem. 2013;288:23833–43.
Lu H, Bhoopatiraju S, Wang H, Schmitz NP, Wang X, Freeman MJ et al. Loss of UHRF2 expression is associated with human neoplasia, promoter hypermethylation, decreased 5-hydroxymethylcytosine, and high proliferative activity. Oncotarget. 2016;7:76047.
Yavorski JM, Blanck G. TCGA: increased oncoprotein coding region mutations correlate with a greater expression of apoptosis-effector genes and a positive outcome for stomach adenocarcinoma. Cell Cycle. 2016;15:2157–63.
Hallstrom TC, Nevins JR. Balancing the decision of cell proliferation and cell fate. Cell Cycle. 2009;8:532–5.
Lu H, Liang X, Issaenko OA, Hallstrom TC. Jab1/CSN5 mediates E2F dependent expression of mitotic and apoptotic but not DNA replication targets. Cell Cycle. 2011;10:1–10.
Fisler DA, Sikaria D, Yavorski JM, Tu YN, Blanck G. Elucidating feed-forward apoptosis signatures in breast cancer datasets: higher FOS expression associated with a better outcome. Oncol Lett. 2018;16:2757–63.
Laurie N, Donovan S, Shih C, Zhang J, Mills N, Fuller C, et al. Inactivation of the p53 pathway in retinoblastoma. Nature. 2006;444:61–66.
Xu XL, Fang Y, Lee TC, Forrest D, Gregory-Evans C, Almeida D, et al. Retinoblastoma has properties of a cone precursor tumor and depends upon cone-specific MDM2 signaling. Cell. 2009;137:1018–31.
Conkrite K, Sundby M, Mu D, Mukai S, MacPherson D. Cooperation between Rb and Arf in suppressing mouse retinoblastoma. J Clin Invest. 2012;122:1726–33.
Stambolic V, Suzuki A, de la Pompa JL, Brothers GM, Mirtsos C, Sasaki T, et al. Negative regulation of PKB/Akt-dependent cell survival by the tumor suppressor PTEN. Cell. 1998;95:29–39.
Cully M, You H, Levine AJ, Mak TW. Beyond PTEN mutations: the PI3K pathway as an integrator of multiple inputs during tumorigenesis. Nat Rev Cancer. 2006;6:184–92.
Nath S, Chowdhury A, Dey S, Roychoudhury A, Ganguly A, Bhattacharyya D, et al. Deregulation of Rb-E2F1 axis causes chromosomal instability by engaging the transactivation function of Cdc20-anaphase-promoting complex/cyclosome. Mol Cell Biol. 2015;35:356–69.
Hallstrom TC, Mori S, Nevins JR. An E2F1-dependent gene expression program that determines the balance between proliferation and cell death. Cancer Cell. 2008;13:11–22.
Xie C, Freeman MJ, Lu H, Wang X, Forster CL, Sarver AL et al. Retinoblastoma cells activate the AKT pathway and are vulnerable to the PI3K/mTOR inhibitor NVP-BEZ235. Oncotarget. 2017;8:38084.
Chen D, Livne-bar I, Vanderluit J, Slack R, Agochiya M, Bremner R. Cell-specific effects of RB or RB/p107 loss on retinal development implicate an intrinsically death-resistant cell-of-origin in retinoblastoma. Cancer Cell. 2004;5:539–51.
Conkrite K, Sundby M, Mukai S, Thomson JM, Mu D, Hammond SM, et al. miR-17~92 cooperates with RB pathway mutations to promote retinoblastoma. Genes Dev. 2011;25:1734–45.
Xu XL, Singh HP, Wang L, Qi DL, Poulos BK, Abramson DH, et al. Rb suppresses human cone-precursor-derived retinoblastoma tumours. Nature. 2014;514:385–8.
McEvoy J, Flores-Otero J, Zhang J, Nemeth K, Brennan R, Bradley C, et al. Coexpression of normally incompatible developmental pathways in retinoblastoma genesis. Cancer Cell. 2011;20:260–75.
Xie C, Lu H, Nomura A, Hanse EA, Forster CL, Parker JB, et al. Co-deleting Pten with Rb in retinal progenitor cells in mice results in fully penetrant bilateral retinoblastomas. Mol Cancer. 2015;14:93.
Robanus-Maandag E, Dekker M, van der Valk M, Carrozza M, Jeanny J, Dannenberg J, et al. p107 is a suppressor of retinoblastoma development in pRb-deficient mice. Genes Dev. 1998;12:1599–609.
Donovan SL, Schweers B, Martins R, Johnson D, Dyer MA. Compensation by tumor suppressor genes during retinal development in mice and humans. BMC Biol. 2006;4:14.
Zhang J, Schweers B, Dyer M. The first knockout mouse model of retinoblastoma. Cell Cycle. 2004;3:952–9.
Rowan S, Cepko CL. Genetic analysis of the homeodomain transcription factor Chx10 in the retina using a novel multifunctional BAC transgenic mouse reporter. Dev Biol. 2004;271:388–402.
Sakagami K, Chen B, Nusinowitz S, Wu H, Yang XJ. PTEN regulates retinal interneuron morphogenesis and synaptic layer formation. Mol Cell Neurosci. 2011;49:171–83.
Cantrup R, Dixit R, Palmesino E, Bonfield S, Shaker T, Tachibana N, et al. Cell-type specific roles for PTEN in establishing a functional retinal architecture. PLoS ONE. 2012;7:e32795.
Jo HS, Kang KH, Joe CO, Kim JW. Pten coordinates retinal neurogenesis by regulating Notch signalling. EMBO J. 2012;31:817–28.
Furnari FB, Lin H, Huang H-JS, Cavenee WK. Growth suppression of glioma cells by PTEN requires a functional phosphatase catalytic domain. Proc Natl Acad Sci USA. 1997;94:12479–84.
Lee M-H, Williams BO, Mulligan G, Mukai S, Bronson RT, Dyson N, et al. Targeted disruption of p107: functional overlap between p107 and Rb. Genes Dev. 1996;10:1621–32.
Trotman L, Niki M, Dotan Z, Koutcher J, Di Cristofano A, Xiao A. et al. Pten dose dictates cancer progression in the prostate. PLoS Biol. 2003;1:385–96.
Filtz EA, Emery A, Lu H, Forster CL, Karasch C, Hallstrom TC. Rb1 and Pten co-deletion in osteoblast precursor cells causes rapid lipoma formation in mice. PLoS ONE. 2015;10:e0136729.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25:1105–11.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Consortium SM-I. A comprehensive assessment of RNA-seq accuracy, reproducibility and information content by the Sequencing Quality Control Consortium. Nat Biotechnol. 2014;32:903–14.
Sarver AL. Toward understanding the informatics and statistical aspects of micro-RNA profiling. J Cardiovasc Transl Res. 2010;3:204–11.
Beckmann PJ, Larson JD, Larsson AT, Ostergaard JP, Wagner S, Rahrmann EP, et al. Sleeping beauty insertional mutagenesis reveals important genetic drivers of central nervous system embryonal tumors. Cancer Res. 2019;79:905–17.
Scott MC, Temiz NA, Sarver AE, LaRue RS, Rathe SK, Varshney J, et al. Comparative transcriptome analysis quantifies immune cell transcript levels, metastatic progression, and survival in osteosarcoma. Cancer Res. 2018;78:326–37.
Lu H, Hallstrom TC. Sensitivity to TOP2 targeting chemotherapeutics is regulated by Oct1 and FILIP1L. PLoS ONE. 2012;7:e42921.
Di Cristofano A, Pandolfi P. The multiple roles of PTEN in tumor suppression. Cell. 2000;100:387–90.
Breuksch I, Welter J, Bauer HK, Enklaar T, Frees S, Thuroff JW, et al. In renal cell carcinoma the PTEN splice variant PTEN-Delta shows similar function as the tumor suppressor PTEN itself. Cell Commun Signal. 2018;16:35.
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, et al. Integrative genomics viewer. Nat Biotechnol. 2011;29:24–26.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28:511–5.
Jensen AM, Wallace VA. Expression of sonic hedgehog and its putative role as a precursor cell mitogen in the developing mouse retina. Development. 1997;124:363–71.
Furukawa T, Morrow EM, Li T, Davis FC, Cepko CL. Retinopathy and attenuated circadian entrainment in Crx-deficient mice. Nat Genet. 1999;23:466–70.
Mao CA, Tsai WW, Cho JH, Pan P, Barton MC, Klein WH. Neuronal transcriptional repressor REST suppresses an Atoh7-independent program for initiating retinal ganglion cell development. Dev Biol. 2010;349:90–99.
Mears AJ, Kondo M, Swain PK, Takada Y, Bush RA, Saunders TL, et al. Nrl is required for rod photoreceptor development. Nat Genet. 2001;29:447–52.
De Filippo K, Dudeck A, Hasenberg M, Nye E, van Rooijen N, Hartmann K, et al. Mast cell and macrophage chemokines CXCL1/CXCL2 control the early stage of neutrophil recruitment during tissue inflammation. Blood. 2013;121:4930–7.
Bystry RS, Aluvihare V, Welch KA, Kallikourdis M, Betz AG. B cells and professional APCs recruit regulatory T cells via CCL4. Nat Immunol. 2001;2:1126–32.
Charmoy M, Brunner-Agten S, Aebischer D, Auderset F, Launois P, Milon G, et al. Neutrophil-derived CCL3 is essential for the rapid recruitment of dendritic cells to the site of Leishmania major inoculation in resistant mice. PLoS Pathog. 2010;6:e1000755.
Trowbridge IS, Thomas ML. CD45: an emerging role as a protein tyrosine phosphatase required for lymphocyte activation and development. Annu Rev Immunol. 1994;12:85–116.
Gregerson DS, Yang J. CD45-positive cells of the retina and their responsiveness to in vivo and in vitro treatment with IFN-gamma or anti-CD40. Invest Ophthalmol Vis Sci. 2003;44:3083–93.
Reyes NJ, Mathew R, Saban DR. Fate mapping in vivo to distinguish bona fide microglia versus recruited monocyte-derived macrophages in retinal disease. Methods Mol Biol. 2019;1834:153–64.
Aldiri I, Xu B, Wang L, Chen X, Hiler D, Griffiths L, et al. The dynamic epigenetic landscape of the retina during development, reprogramming, and tumorigenesis. Neuron. 2017;94:550–568 e510.
Serra V, Markman B, Scaltriti M, Eichhorn PJ, Valero V, Guzman M, et al. NVP-BEZ235, a dual PI3K/mTOR inhibitor, prevents PI3K signaling and inhibits the growth of cancer cells with activating PI3K mutations. Cancer Res. 2008;68:8022–30.
Wood L, Parsons D, Jones S, Lin J, Sjoblom T, Leary R, et al. The genomic landscapes of human breast and colorectal cancers. Science. 2007;318:1108–13.
Calo E, Quintero-Estades JA, Danielian PS, Nedelcu S, Berman SD, Lees JA. Rb regulates fate choice and lineage commitment in vivo. Nature. 2010;466:1110–4.
Ku SY, Rosario S, Wang Y, Mu P, Seshadri M, Goodrich ZW, et al. Rb1 and Trp53 cooperate to suppress prostate cancer lineage plasticity, metastasis, and antiandrogen resistance. Science. 2017;355:78–83.
Solin SL, Shive HR, Woolard KD, Essner JJ, McGrail M. Rapid tumor induction in zebrafish by TALEN-mediated somatic inactivation of the retinoblastoma1 tumor suppressor rb1. Sci Rep. 2015;5:13745.
Naert T, Colpaert R, Van Nieuwenhuysen T, Dimitrakopoulou D, Leoen J, Haustraete J, et al. CRISPR/Cas9 mediated knockout of rb1 and rbl1 leads to rapid and penetrant retinoblastoma development in Xenopus tropicalis. Sci Rep. 2016;6:35264.
Naert T, Van Nieuwenhuysen T, Vleminckx K. TALENs and CRISPR/Cas9 fuel genetically engineered clinically relevant Xenopus tropicalis tumor models. Genesis 2017;55:e23005.
Friend SH, Bernards R, Rogers J, Weinberg RA, Rapin AMC, Albert DM, et al. A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature. 1986;323:643–6.
Nemeth KM, Federico S, Carcaboso AM, Shen Y, Schaiquevich P, Zhang J, et al. Subconjunctival carboplatin and systemic topotecan treatment in preclinical models of retinoblastoma. Cancer. 2011;117:421–34.
Topper MJ, Vaz M, Marrone KA, Brahmer JR, Baylin SB. The emerging role of epigenetic therapeutics in immuno-oncology. Nat Rev Clin Oncol. 2020;17:75–90.
Snyder AG, Hubbard NW, Messmer MN, Kofman SB, Hagan CE, Orozco SL et al. Intratumoral activation of the necroptotic pathway components RIPK1 and RIPK3 potentiates antitumor immunity. Sci Immunol. 2019;4:36.
Li F, Kitajima S, Kohno S, Yoshida A, Tange S, Sasaki S, et al. Retinoblastoma inactivation induces a protumoral microenvironment via enhanced CCL2 secretion. Cancer Res. 2019;79:3903–15.
Kitajima S, Li F, Takahashi C. Tumor milieu controlled by RB tumor suppressor. Int J Mol Sci. 2020;21:2450.
Li J, Yen C, Liaw D, Podsypanina K, Bose S, Wang SI, et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science. 1997;275:1943–7.
Horie Y, Suzuki A, Kataoka E, Sasaki T, Hamada K, Sasaki J, et al. Hepatocyte-specific Pten deficiency results in steatohepatitis and hepatocellular carcinomas. J Clin Invest. 2004;113:1774–83.
Peng W, Chen JQ, Liu C, Malu S, Creasy C, Tetzlaff MT, et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 2016;6:202–16.
Ankers JM, Awais R, Jones NA, Boyd J, Ryan S, Adamson AD, et al. Dynamic NF-kappaB and E2F interactions control the priority and timing of inflammatory signalling and cell proliferation. Elife. 2016;5:e10473.
Jin X, Ding D, Yan Y, Li H, Wang B, Ma L, et al. Phosphorylated RB promotes cancer immunity by inhibiting NF-κB activation and PD-L1 expression. Mol Cell. 2019;73:22–35.
Ildefonso CJ, Kong L, Leen A, Chai SJ, Petrochelli V, Chintagumpala M, et al. Absence of systemic immune response to adenovectors after intraocular administration to children with retinoblastoma. Mol Ther. 2010;18:1885–90.
Pascual-Pasto, G, Bazan-Peregrino, M, Olaciregui, NG, Restrepo-Perdomo, CA, Mato-Berciano, A, Ottaviani, D et al. Therapeutic targeting of the RB1 pathway in retinoblastoma with the oncolytic adenovirus VCN-01. Sci Transl Med. 2019;11:476.
Minhas G, Sharma J, Khan N. Cellular stress response and immune signaling in retinal ischemia-reperfusion injury. Front Immunol. 2016;7:444.
Karlstetter M, Kopatz J, Aslanidis A, Shahraz A, Caramoy A, Linnartz-Gerlach B, et al. Polysialic acid blocks mononuclear phagocyte reactivity, inhibits complement activation, and protects from vascular damage in the retina. EMBO Mol Med. 2017;9:154–66.
Akhtar-Schäfer, I, Wang, L, Krohne, TU, Xu, H & Langmann, T Modulation of three key innate immune pathways for the most common retinal degenerative diseases. EMBO Mol Med 2018;10:e8259 (2018).
Fuchs Y, Steller H. Live to die another way: modes of programmed cell death and the signals emanating from dying cells. Nat Rev Mol Cell Biol. 2015;16:329–44.
Nittner D, Lambertz I, Clermont F, Mestdagh P, Köhler C, Nielsen SJ, et al. Synthetic lethality between Rb, p53 and Dicer or miR-17-92 in retinal progenitors suppresses retinoblastoma formation. Nat Cell Biol. 2012;14:958–65.
Murakami Y, Matsumoto H, Roh M, Giani A, Kataoka K, Morizane Y, et al. Programmed necrosis, not apoptosis, is a key mediator of cell loss and DAMP-mediated inflammation in dsRNA-induced retinal degeneration. Cell Death Differ. 2014;21:270–7.
Tanzer MC, Frauenstein A, Stafford CA, Phulphagar K, Mann M, Meissner F. Quantitative and dynamic catalogs of proteins released during apoptotic and necroptotic cell death. Cell Rep. 2020;30:1260–70.
Acknowledgements
This work was funded by grants to T.C.H. from N.I.H. (R01CA168622) and the Children’s Cancer Research Fund, Minneapolis, MN.
Author information
Authors and Affiliations
Contributions
A.L.S. and T.C.H. designed and supervised this study, interpreted data, and wrote the paper. C.X. and M.J.R. contributed to study design, performed experiments, collected, and analyzed data. J.T. contributed to study design and analyzed data. C.L.F., X.W., H.L., W.W., and J.B.P. performed experiments.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Sarver, A.L., Xie, C., Riddle, M.J. et al. Retinoblastoma tumor cell proliferation is negatively associated with an immune gene expression signature and increased immune cells. Lab Invest 101, 701–718 (2021). https://doi.org/10.1038/s41374-021-00573-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-021-00573-x