Abstract
Patient-derived xenograft (PDX) murine models are employed for preclinical research on cancers, including non-small cell lung cancers (NSCLCs). Even though lung squamous cell carcinomas (LUSCs) show the highest engraftment rate among NSCLCs, half of them nevertheless show PDX failure in immunodeficient mice. Here, using immunohistochemistry and RNA sequencing, we evaluated the distinct immunohistochemical and gene expression profiles of resected LUSCs that showed successful engraftment. Among various LUSCs, including the basal, classical, secretory, and primitive subtypes, those in the non-engrafting (NEG) group showed gene expression profiles similar to the pure secretory subtype with positivity for CK7, whereas those in the engrafting (EG) group were similar to the mixed secretory subtype with positivity for p63. Pathway analysis of 295 genes that demonstrated significant differences in expression between NEG and EG tumors revealed that the former had enriched expression of genes related to the immune system, whereas the latter had enriched expression of genes related to the cell cycle and DNA replication. Interestingly, NEG tumors showed higher infiltration of B cells (CD19+) and follicular dendritic cells (CD23+) in lymph follicles than EG tumors. Taken together, these findings suggest that the PDX cancer model of LUSC represents only a certain population of LUSCs and that CD19- and CD23-positive tumor-infiltrating immune cells in the original tumors may negatively influence PDX engraftment in immunodeficient mice.
Similar content being viewed by others
Introduction
Lung squamous cell carcinoma (LUSC) is a type of non-small cell lung cancer (NSCLC) linked more strongly with smoking than the other common type of NSCLC, lung adenocarcinoma (LUAD) [1]. LUAD originates from peripheral airway cells with TTF1 expression, whereas LUSC is characterized by squamous differentiation with p40 expression, suggesting an origin from basal cells of the bronchial mucosa [1, 2]. Wilkerson et al. have classified LUSCs into the basal, classical, secretory, and primitive subtypes, which show dramatic differences in their mRNA expression profiles due to their different ancestor cells and differentiation stages [3, 4]. They also show distinctive differences in their biological processes associated with cell adhesion, xenobiotic metabolism, immune response, and proliferation, respectively [3]. Recently, genomic alterations in LUSCs have been comprehensively characterized, and they are known to have a high somatic mutation rate similar to the genetic signature of tobacco smoking, with a sensitive response to immunotherapies such as immune checkpoint inhibitors [4]. Unlike LUADs, molecularly targeted agents for specific treatment of LUSCs are still being developed [4, 5].
Patient-derived xenograft (PDX) models can be generated by direct transplantation of human tumors, obtained by surgery or biopsy, into immunodeficient mice for rapid expansion of the specimen [6]. Since PDX tumors mostly retain the principal histological and genetic characteristics of their original tumors and remain stable across passages [7, 8], they can provide a better understanding of cancer biology, allowing investigation of novel anticancer treatments, and development of personalized therapeutic strategies [9]. However, the successful construction of a PDX model (engraftment rate 20–80%) depends on the purity of the original tumor used as the transplanted material, the nature of the transplantation site (e.g., subcutaneous or orthotopic implantation), and various aspects of the recipient mouse strain [9]. Among NSCLCs, LUSCs show the highest engraftment rate, but nevertheless half of such tumors fail to be established as PDXs. Thus, the factors that determine the success of PDX using material from LUSCs remain unclear.
It is possible that factors in the tumor microenvironment, such as patient-derived tumor-infiltrating immune cells, might influence the rate of PDX establishment. The infiltrating immune cells of LUSCs showing a high proliferation rate include a high proportion of lymphocytes but a low proportion of dendritic cells or neutrophils [10, 11]. Particularly, the lymph follicles at peritumoral and intratumoral sites have organized lymphoid aggregates including T cells, B cells, follicular dendritic cells, and mature dendritic cells with or without germinal centers. The high expression of lymph follicles is reportedly associated with a favorable outcome in LUSC patients [12, 13]. Since the infiltrating immune cells within patient-derived specimens are frequently cotransplanted with the original tumor tissues into immunodeficient mice, it would be informative to investigate whether the presence of lymph follicles in tumors affects the engraftment of PDX.
In the present study using RNA sequencing and immunohistochemistry (IHC), we evaluated the distinct gene expression profiles and immunohistochemical features of resected LUSCs that may affect the engraftment success rate. We classified patient-derived and PDX tumors into their LUSC subtypes on the basis of gene expression profiling. Non-engrafting (NEG) patient-derived tumors were of the pure secretory subtype and engrafting (EG) patient-derived and PDX tumors included the four LUSC subtypes. Our findings suggested that tumor-infiltrating immune cells including CD19+ B cells or CD23+ follicular dendritic cells in lymph follicles may be associated with engraftment failure.
Materials and methods
Preparation of clinical samples of tumor tissues for PDX generation and immunohistochemical analysis
Between January 2015 and August 2017, we collected 51 clinical samples of lung carcinomas at the University of Tsukuba Hospital (Tsukuba, Japan). To increase the success rate of PDX generation, we inoculated the tumor specimens into immunodeficient mice immediately after resection. Moreover, to avoid interference of stromal cells such as fibroblasts, we carefully collected small pieces of tumor tissue (2–3 mm3) comprising more than about 95% tumor cells under macroscopic (gross) observation by two pathologists. The samples included 10 squamous cell carcinomas (LUSCs), 37 adenocarcinomas (LUADs), and 4 other histological types.
Clinical information for all of the corresponding patients was obtainable from the medical records, and all of the patients provided informed consent for use of their materials. This study was approved by the Institutional Ethical Review Committee. The LUSC cases were classified according to the World Health Organization (WHO) histological classification of malignant tumors (4th edition) and evaluated pathologically in accordance with the UICC TNM classification of malignant tumors (7th edition) [14, 15].
Animals and establishment of tumor xenografts
For PDX generation we used C.B-17/ICR-SCID immunodeficient mice (Charles River Laboratories, Tsukuba, Japan) aged 6–8 weeks. The mice were subjected to inhalation anesthesia with isoflurane and propylene glycol (3:7, Wako Pure Chemical Industries, Osaka, Japan), and then received subcutaneous transplants of fresh tumor tissue. The PDX tumors that subsequently formed were dissected out when the tumor weight/body weight ratio became 5–8%, and then passaged to new recipients.
RNA sequencing of tumor tissues and PDX tumors
Total RNA was extracted from snap-frozen clinical samples of LUSC tumor tissues and PDXs using TRIZOL reagent in accordance with the manufacturer’s protocol (Thermo Fisher Scientific, Waltham, MA). RNA quality was examined using the Agilent Bioanalyzer RNA 6000 pico kit (Agilent Technologies, Santa Clara, CA). After depletion of rRNA with a NEB Next rRNA Depletion Kit (New England Biolabs, Ipswich, MA) using 500 ng of total RNA, a RNA sequencing library was prepared using a NEB Next Ultra Directional RNA Library Prep Kit (New England Biolabs). Paired-end sequencing of 2 × 36 bases was performed using NextSeq500 (Illumina, San Diego, CA) at Tsukuba i-Laboratory LLP and Tsukuba Transborder Medical Center (Tsukuba, Japan). FASTQ files were imported to the CLC Genomics Workbench software (Version 10.1.1; Qiagen, Hilden, Germany) as paired-end reads. Sequences were mapped to the human genome (hg19) and quantified for annotated genes. Transcription expression was subjected to empirical pairwise analysis using EDGE test in the CLC Genomics Workbench (Fig. S1). To observe the differences in gene expression profile, we selected 295 genes among the total of 57,773 genes, which were considered statistically significant with a false discovery rate (FDR) P value < 0.05 and a fold change of more than 2.0.
Immunohistochemical analysis
We used 4-μm-thick whole sections obtained from formalin-fixed paraffin-embedded tissue blocks. After deparaffinization and rehydration, antigen retrieval was performed using an autoclave in 10 mM Tris-EDTA buffer (pH 9.0) at 105 °C for 15 min or 10 mM citrate buffer (pH 6.1) at 121 °C for 10 min. Endogenous peroxidase was then blocked using 3% H2O2 for 30 min to avoid any non-specific reactions. Immunostaining was performed using a Dako Autostainer Link 48 (Agilent Technologies) with the appropriate primary antibody and REAL Envision HRP rabbit/mouse (Agilent Technologies) as a secondary antibody. The immunoreactivity was detected with DAB (Dako REAL Envision Detection System; Agilent Technologies), and counterstaining was performed with hematoxylin for 1 min. Antibodies against the following were used for immunohistochemistry (IHC): p40 (Abcam, Cambridge, MA, mouse BC28 clone, ab172731, 1:500), TTF1 (Dako, Glostrup, Denmark, mouse 8G7G3/1 clone, IR056, ready to use), CK7 (Nichirei Biosciences Inc., Tokyo, Japan, mouse OV-TL12/39 clone, #413481, ready to use), p63 (Dako, mouse DAK-p63 clone, IR662, ready to use), S100 (Dako, IR504, ready to use), CD19 (Dako, mouse LE-CD19 clone, M7296, 1:100), CD23 (Dako, mouse DAK-CD23 clone, M7312, 1:50). For CD19 and CD23 IHC scoring, we counted the number of lymph follicles that contained clusters of CD19- and CD23-positive cells surrounding the tumor area, but we excluded single-positive cells infiltrating the tumor area. We then constructed a ROC curve to derive a median value that could be used as the cut-off point using the SPSS 22 statistical software package (SPSS, Chicago, IL).
Bioinformatics and statistical analysis
For comparison of gene expression between NEG and EG tumors, we constructed a heatmap using Morpheus (https://software.broadinstitute.org/morpheus/) and performed hierarchical clustering of samples and genes using the Euclidean distance metric. The clustered genes were examined using the Reactome Pathway Database (https://reactome.org/) and the enrichment pathway was determined for each cluster. Based on the gene expression detected by RNA sequencing, LUSC subtypes were determined using Gene Expression Profiling Interactive Analysis (GEPIA2, http://gepia.cancer-pku.cn/). To investigate cellular proliferation-associated genes, we used the Gene Ontology Resource (http://geneontology.org/). The comparison mRNA expression levels of engraftment-associated genes in both normal and tumor tissues of LUSCs were examined using The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/) and the Genotype-Tissue Expression (GTEx) databases (https://gtexportal.org/).
For statistical analysis, we used the SPSS 22 and Prism 8 (GraphPad, San Diego, CA) statistical software packages. Correlations of clinicopathological features were analyzed using the chi-squared test. Group results of IHC were expressed as mean ± SD. Data were compared between groups using the t test to calculate two-tailed distributions and the paired t test.
Results
Significant association between PDX generation and pathological subtype of lung cancer
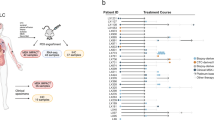
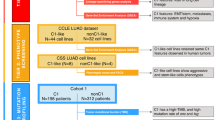
We used 51 surgically resected samples of primary lung cancer tissue obtained at the University of Tsukuba Hospital to generate PDX tumors (Fig. 1). Among these cases, 10 were LUSCs, 37 were LUADs, and 4 were other histological subtypes including large cell neuroendocrine carcinoma, small cell carcinoma, and adenosquamous cell carcinoma. Twenty-three of the 51 samples formed tumors at passage 1 (P1, 45.1%), 7 of the 23 P1 tumors also formed tumors at passage 2 (P2, 30.4%), and only 4 out of the 7 P2 tumors formed tumors at passage 3 tumors (P3, 57.1%). We classified the patient-derived tumor samples as either non-engrafting (NEG) or engrafting (EG), according to whether they were unsuccessfully or successfully transplanted into immunodeficient mice at P1, respectively. PDX formation was significantly associated with histological subtype. The PDX success rate was highest (80%) for LUSCs and lowest (32.4%) for LUADs (Table 1, Table S1). In this study, we focused on LUSCs for further investigation.
The 51 specimens of freshly resected primary lung carcinoma tissue were transplanted into C.B-17/ICR-SCID mice. Depending on whether or not PDX tumors formed, we divided the cancer specimens into the non-engrafting (NEG) and engrafting (EG) groups. Most primary lung carcinomas, except for lung squamous cell carcinoma (LUSC), did not produce a stable PDX tumor beyond passage 3 (P3). To clarify the differences in characteristics between NEG and EG tumors, immunohistochemistry and RNA sequencing were performed. LUAD, lung adenocarcinoma; LCNEC, large cell neuroendocrine carcinoma; Small, small cell carcinoma; Ad+Sq, adenosquamous cell carcinoma.
Association of specific LUSC subtype with successful generation of PDX
To examine the histological features of patient-derived tumors in the NEG group and paired samples of patient-derived and PDX tumors in the EG group, we performed HE staining and IHC staining for p40, which is a specific marker of squamous differentiation. All of the patient-derived tumors were typically moderately differentiated squamous cell carcinoma with intracellular bridges and lacking gland formation, confirming that the tumors in the two groups had no obvious histological differences (Fig. S2). Although the PDX tumors showed slight dedifferentiation relative to each corresponding paired patient-derived tumor (Fig. S2), they were still diagnosed as squamous cell carcinoma in view of their positivity for p40 (data not shown).
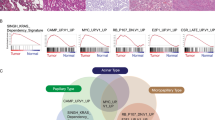
To evaluate differences of gene expression profiles between tumors in the NEG and EG groups, we performed RNA sequencing using snap-frozen samples of patient-derived and PDX tumors. Among 8 cases in the EG group, we excluded 4 cases due to failure of stable PDX formation until passage 3. Using data from mRNA expression profiling, Wilkerson et al. had previously classified LUSCs into four subtypes, which are associated with differentiation stage: “basal (early stage),” “classical (early-mid stage),” “secretory (mid-late stage),” and “primitive (late stage)” [3]. To identify the subtypes of the patient-derived and PDX tumors in the NEG and EG groups, we analyzed their gene expression using the cancer subtype classifier in GEPIA2 based on the Naïve Bayes algorithm to estimate the probability for each subtype. Interestingly, patient-derived tumors in the NEG group were of the pure “secretory” subtype whereas patient-derived and PDX tumors in the EG group included not only the “secretory” subtype but also had characteristics of the “basal,” “classical” and “primitive” subtypes (Fig. 2a). Unlike patient-derived tumors in both the NEG and EG groups that were predominantly of the secretory subtype, PDX tumors in the EG group included the four LUSC subtypes (Fig. 2a). It was possible to characterize these subtypes by IHC using selected marker proteins such as S100 for the basal subtype, p63 (TP63) for the classical subtype, and CK7 (KRT7) for the secretory subtype, with the exception of the primitive subtype [3]. To compare the IHC features of the LUSC subtypes between patient-derived tumors in the NEG and EG groups, we examined the expression of three marker proteins. Although some of the tumor tissues showed heterogeneous expression, most patient-derived tumors in the NEG group showed strong positivity for CK7 and weak positivity for p63, but all were negative for S100 (Figs. 2b, 2c, S3). However, all patient-derived tumors in the EG group showed strong positivity for p63 and negativity for S100, and 3 of the 4 tumors were negative for CK7 (Figs. 2b, 2c, S3). Only one patient-derived tumor in the EG group was CK7-positive. Although it could not be differentiated from other CK7-negative tumors histologically, it may have had functionally poorer differentiation of LUSC than the others in the EG group.
A The patient-derived and PDX tumors in the non-engrafting (NEG) and engrafting (EG) groups were classified in relation to the subtypes of lung squamous cell carcinoma (LUSC) based on RNA sequencing data using GEPIA2. B-C Six cases of primary LUSC were subjected to immunohistochemistry (IHC) to distinguish the subtypes using the following marker proteins: p40, a marker of lung squamous cell carcinoma; TTF1, a marker of lung adenocarcinoma; CK7, a marker of the secretory subtype; p63, a marker of the classical subtype: S100, a marker of the basal subtype. All tissues were positive for p40, negative for TTF1, and negative for S100.
Distinct gene expression profile associated with PDX engraftment
We performed RNA sequencing and filtered 295 genes that showed significant differences of gene expression profiles among 4 groups including patient-derived tumors in the NEG and EG groups and PDX tumors in the P1 and P3 of passage groups (Table S2). Using enrichment pathway analysis, the 295 filtered genes were classified into 6 gene expression pathways, shown by a heatmap with clustering of genes and samples based on their expression levels (Fig. 3). In patient-derived tumors in the NEG group, enriched expression of genes was more significantly associated with programed cell death, immune system, signal transduction, and extracellular matrix organization (Fig. 3). This indicated that patient-derived tumors in the NEG group had characteristics similar to the secretory subtype, which showed a distinct immune cell signature such as a marked inflammatory immune response [11]. Interestingly, enriched expression of genes in the EG group was very limited, and most genes were closely related to the cell cycle and DNA replication pathways (Fig. 3), suggesting that the characteristics of patient-derived tumors in the EG group were relatively similar to those of the primitive subtype than those of NEG tumors. Consistent with the patient-derived tumors in the EG group, PDX tumors (P1 and P3) rarely expressed genes related to immune system pathways (Fig. S4), and instead revealed enrichment of genes related to the cell cycle and DNA replication pathways.
Using RNA sequencing, we detected 295 genes showing significant differences in expression between non-engrafting (NEG) and engrafting (EG) patient-derived tumors (FDR P value < 0.05, fold change >2). Hierarchical clustering was used to produce a 2-tree clustered heatmap. Based on the tree clustering, gene expression pathways were examined using Reactome pathway analysis. PDX engraftment was closely associated with high expression of genes related to the cell cycle and DNA replication but low expression of genes related to immune system and signal transduction. Each clustered group of gene expression pathway was P < 0.001 and FDR < 0.05.
Identification of engraftment-associated genes that can distinguish between non-engrafting and engrafting tumors
To identify engraftment-associated genes that might predict whether or not the original tumors might successfully engraft in immunodeficient mice, we derived two sets of genes based on the fold change in their expression between the NEG and EG groups: expressions of 84 genes and 8 genes were enriched in the NEG group and in the EG group, respectively (Fig. 4a). We also examined genes associated with cellular proliferation using the Gene Ontology database and found 1997 genes that were assumed to contribute to tumorigenic signaling. Among them, only 11 genes were identified as engraftment-associated genes, including the EG-associated gene set in Fig. 4b and the NEG-associated gene set in Fig. 4d and 4f. Moreover, the EG-associated gene set predominantly included genes related to the cell cycle pathway and the DNA replication pathway (Fig. 4b, 4c). The NEG-associated gene set predominantly included genes related to the signal transduction pathway (Fig. 4d, 4e) and immune system pathway including the innate and adaptive immune systems, and cytokine signaling (Fig. 4f-4j, Table S3).
A Cellular proliferation-associated genes (n = 1997) were identified by biological process analysis of the Gene Ontology (GO) database. Among the 295 selected genes detected by RNA sequencing, 92 PDX engraftment-associated genes were filtered with genes that showed a difference in expression of more than double between the non-engrafting (NEG) and engrafting (EG) groups. Two of EG-associated genes (B) and 9 of NEG-associated genes (D, F) belonged to cellular proliferation pathways in the GO biological process. B–J EG-associated gene (n = 8) were limited (B, C), whereas representative NEG-associated genes included signal transduction-related genes (D, E) and immune system-related genes (F, J). The representative immune system-related genes were divided into those involved in the common pathway (innate and adaptive immune systems) (G), innate immune system (H), adaptive immune systems (I), and cytokine signaling pathway (J). All asterisks indicate a P value < 0.05 by two-tailed Student’s t test.
Furthermore, we investigated the distinct levels of expression of engraftment-associated genes in both normal and tumor tissues of LUSCs using the TCGA and GTEx databases. Most of the NEG-associated genes showed higher expression but the EG-associated genes showed lower expression in normal than in tumor tissues (Fig. S5). These results indicated that patient-derived tumors in the NEG group had gene expression profiles relatively closer to those of normal tissues than was the case for tumors in the EG group.
Enriched expression of CD19 and CD23 in lymph follicles is associated with restriction of PDX engraftment
To evaluate further the clinical characteristics of patients whose tumors showed similar NEG or EG features in terms of engraftment-associated genes, we focused the immune system-related genes that showed the greatest difference in expression levels between the NEG and EG groups. We hypothesized that enriched expression of immune system-related genes in patient-derived tumors might influence the rejection of mice against grafted human tumors. Since cell surface molecules interact with receptors and ligands to regulate the immune response and provide information to identify specific immune cells, we focused on clusters of differentiation (CD) molecules and found 6 genes: CD19, LILRA2 (CD85H), and LILRA6 (CD85B) in the common pathway, CR1 (CD35) in the innate immune system, CD22 in the adaptive immune system, and FCER2 (CD23) in the cytokine signaling system (Fig. 4g-4j). The enriched expression of these 6 genes in patient-derived tumors in the EG and NEG groups was validated by IHC. Among them, CD19 and FCER2 (CD23) showed unequivocally distinct differences in positivity in patient-derived tumors between the NEG and EG groups. The CD19- and CD23-positive cells showed aggregation with lymph node-like structures, which were considered to be lymph follicles based on comparison with normal tonsil as an external positive control (Fig. 5a). However, normal cells, including those in alveoli and bronchioles, were totally negative for CD19 and CD23 (Fig. 5a).
A, B Immunohistochemistry (IHC) was performed on patient-derived tumors in the non-engrafting (NEG) and engrafting (EG) groups using anti-CD19 and anti-CD23. A Lung alveoli and bronchiole were used as internal negative controls and tonsil as a positive control for CD19 and CD23 IHC. B Representative patient-derived tumors in the NEG group detected positivity of tumor-infiltrating immune cells in CD19+ lymph follicles and CD23+ secondary lymph follicles, whereas those in the EG group were mostly negative or showed single cell staining pattern. C The numbers of CD19+ lymph follicles and CD23+ secondary lymph follicles were counted, and the results for each specimen were presented as a grayscale heatmap.
Consistent with the results of RNA sequencing, we found that patient-derived tumors in the NEG group showed the clustered positive cells, and we counted each spherical cluster of lymphocytes corresponding to a lymph follicle. The lymph follicles were divisible into two types—primary and secondary—without and with a germinal center (representing proliferating B cells), respectively. The patient-derived tumors in the NEG group had a higher number of CD19+ lymph follicles with or without germinal centers than those in the EG group. Positivity for CD19 showed a single-cell staining pattern, indicating that the CD19+ cells may have been mature and active B cells (Fig. 5b, 5c) [16]. Similar to CD19+ lymph follicles, patient-derived tumors in the NEG group showed a higher number of CD23+ secondary lymphoid follicles, containing proliferating B cells and follicular dendritic cells in their germinal centers, than those in the EG group. Interestingly, the CD23 staining appeared to have a mesh-like membranous pattern, suggesting that the CD23+ cells might be follicular dendritic cells rather than active B cells (Fig. 5b, 5c) [17, 18]. Taken together, these findings suggest that CD19+ and CD23+ lymph follicles in patient-derived tumors might be critical factors that potentially elicit a more robust immune response in the murine microenvironment.
Discussion
Cancer models such as PDXs have been established to identify new tumor biomarkers and clarify the mechanisms of drug resistance [19, 20]. Recently, PDX models are being used increasingly as a valuable platform for assessing the most effective therapeutic regimens of individual drugs or drug combinations in parallel preclinical in vivo and clinical trials for personalized cancer therapy [21].
In this study, we examined the success rate of PDX production from lung carcinomas. Among various histologic subtypes, LUSC is that which produces PDX tumors most readily (Table 1, Fig. 1), and clinicopathological characteristics such as patient sex, age, smoking history, pathologic stage, lymphatic permeation, and vascular invasion do not appear to impact the success of PDX production (Table 1). To investigate potential engraftment-associated factors for the LUSC-PDX model, we compared the expression of various genes between tumors that engrafted successfully and those that did not (Figs. 2, 3).
The non-engrafting (NEG) group of patient-derived tumors, which failed to produce PDX, showed a secretory subtype with positivity for CK7 (Fig. 2), a marker of peripheral pneumocytes that can also be used to identify the primary site of origin of metastatic lesions [22]. However, they showed weak and focal expression p63, which can be a potential indicator of glandular differentiation [23], even though no gland formation was evident histologically (Fig. S2). Consistently, the secretary subtype of LUSC is considered to be more closely associated with adenocarcinoma-like differentiation than the other LUSC subtypes such as the basal, classic, and primitive subtypes [2, 11]. Tumors of the secretory subtype have low proliferation activity reflecting their poor response to EGFR or FGFR targeting therapy [24] and high expression of genes associated with immune response [2, 11]. In this context, it can be suggested that patient-derived tumors in the NEG group might be unable to form PDX tumors due to their low cellular proliferation and similarity to LUADs, which showed a lower engraftment rate than LUSCs (Fig. 1).
The engrafting (EG) group of patient-derived tumors for which PDX was successful was strongly positive for transcription factor p63 (Fig. 2c), which is a marker of squamous epithelium basal cells and has a pivotal function in epithelial development and morphogenesis. The p63 locus is frequently coamplified with SOX2, which directs dividing cells toward an aggressive phenotype and promotes differentiation to LUSC [23, 25]. However, one patient-derived tumor in the EG group showed positivity for CK7, suggesting relatively poorer differentiation of LUSC than other tumors in the EG group. The EG group also showed various characteristics including those of the “secretory”, “basal”, “classical”, and “primitive” subtypes (Fig. 2b), with a relatively higher preponderance of the primitive subtype relative to the NEG group (Fig. 2a). This result indicates that patient-derived tumors with a mixed subtype tend to generate PDX more successfully due to the increased proliferation activity and aggressive characteristics of the tumor cells relative to the pure secretory subtype [3, 26, 27].
Given the similarity of the gene expression profiles between PDX P1 and P3, PDX P2 may be comparable to P1 and P3 as a mixed subtype. Interestingly, PDX P1 and P3 tumors were classified as a more mixed subtype than their paired original tumors and a reduction of secretory-type features. Although the patient-derived and PDX tumors did not exhibit morphological differences, the difference in their gene expression subtypes may have been attributable to replacement of human stromal cells by the murine counterparts upon passaging, or the presence of a tumor microenvironment markedly different from that in humans [8]. Indeed, the murine microenvironment modifies the composition of subclones through murine-specific selection of clonal dynamics or evolution, unlike the situation in humans [28, 29].
In addition to the distinct characteristics of tumor cells, the tumor immune microenvironment exerts important effects on tumor malignancy and the reprogramming of immune cells [30, 31]. Here, we demonstrated that tumor-infiltrating immune cells such as CD19+ cells in lymph follicles or CD23+ cells in secondary lymph follicles were detectable mainly in the peritumoral area of patient-derived tumors in the NEG group but not that of tumors in the EG group (Fig. 5). These results suggest that the tumor-infiltrating immune cells may be considered as factors that affect the cell survival of patient-derived tumors in a murine tissue microenvironment [18]. Intriguingly, the patient-derived tumors in the NEG group showed high expression of genes related to the chemokine (CCL19 and CCL21) and proinflammatory cytokine (IL-10, IL-12, TNF, and IFNγ) pathways (Tables S2 and S3), suggesting that these cytokines recruit B cells, T cells, and follicular dendritic cells to form lymph follicles in the tumor microenvironment [32,33,34,35]. However, further studies will be needed to clarify in detail the molecular mechanisms operating in tumor-infiltrating immune cells in CD19+ or CD23+ lymph follicles via crosstalk with tumor cells in vitro and in vivo as well as the clinical relevance using large-scale human samples.
Taken together, our present results suggest that PDX tumors derived from LUSCs exhibit a specific gene expression profile characterized by a greater degree of squamous epithelial differentiation. The pure secretory subtype of patient-derived LUSC tumors is particularly associated with enrichment of tumor-infiltrating immune cells in CD19+ or CD23+ lymph follicles, possibly representing a major phenotype responsible for impaired tolerance in an immunodeficient murine model. It can therefore be inferred that LUSC-PDX tumors may not completely recapitulate the characteristics of human LUSCs, especially those that contain abundant tumor-infiltrating immune cells in the tumor microenvironment.
References
Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK. Non-small-cell lung cancers: a heterogeneous set of diseases. Nat Rev Cancer. 2014;14:535–46.
Yatabe Y, Dacic S, Borczuk AC, Warth A, Russell PA, Lantuejoul S, et al. Best practices recommendations for diagnostic immunohistochemistry in lung cancer. J Thorac Oncol. 2019;14:377–407.
Wilkerson MD, Yin X, Hoadley KA, Liu Y, Hayward MC, Cabanski CR, et al. Lung squamous cell carcinoma mRNA expression subtypes are reproducible, clinically important, and correspond to normal cell types. Clin Cancer Res. 2010;16:4864–75.
Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature. 2012;489:519–25.
Gandara DR, Hammerman PS, Sos ML, Lara PN Jr., Hirsch FR. Squamous cell lung cancer: from tumor genomics to cancer therapeutics. Clin Cancer Res. 2015;21:2236–43.
Day CP, Merlino G, Van, Dyke T. Preclinical mouse cancer models: a maze of opportunities and challenges. Cell. 2015;163:39–53.
Hidalgo M, Amant F, Biankin AV, Budinska E, Byrne AT, Caldas C, et al. Patient-derived xenograft models: an emerging platform for translational cancer research. Cancer Discov. 2014;4:998–1013.
Jiang Y, Zhao J, Zhang Y, Li K, Li T, Chen X, et al. Establishment of lung cancer patient-derived xenograft models and primary cell lines for lung cancer study. J Transl Med. 2018;16:138.
Morgan KM, Riedlinger GM, Rosenfeld J, Ganesan S, Pine SR. Patient-derived xenograft models of non-small cell lung cancer and their potential utility in personalized medicine. Front Oncol. 2017;7:2.
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH, et al. The immune landscape of cancer. Immunity. 2018;48:812–30.e14.
Faruki H, Mayhew GM, Serody JS, Hayes DN, Perou CM, Lai-Goldman M. Lung adenocarcinoma and squamous cell carcinoma gene expression subtypes demonstrate significant differences in tumor immune landscape. J Thorac Oncol. 2017;12:943–53.
Sautes-Fridman C, Petitprez F, Calderaro J, Fridman WH. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat Rev Cancer. 2019;19:307–25.
Colbeck EJ, Ager A, Gallimore A, Jones GW. Tertiary lymphoid structures in cancer: drivers of antitumor immunity, immunosuppression, or bystander sentinels in disease? Front Immunol. 2017;8:1830.
Travis WD, Brambilla E, Noguchi M, Nicholson AG, Geisinger KR, Yatabe Y, et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol. 2011;6:244–85.
Travis WD, Brambilla E, Nicholson AG, Yatabe Y, Austin JHM, Beasley MB, et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol. 2015;10:1243–60.
Wang K, Wei G, Liu D. CD19: a biomarker for B cell development, lymphoma diagnosis and therapy. Exp Hematol Oncol. 2012;1:36.
Henningsson F, Ding Z, Dahlin JS, Linkevicius M, Carlsson F, Gronvik KO, et al. IgE-mediated enhancement of CD4+ T cell responses in mice requires antigen presentation by CD11c+ cells and not by B cells. PLoS One. 2011;6:e21760.
Silina K, Soltermann A, Attar FM, Casanova R, Uckeley ZM, Thut H, et al. Germinal centers determine the prognostic relevance of tertiary lymphoid structures and are impaired by corticosteroids in lung squamous cell carcinoma. Cancer Res. 2018;78:1308–20.
Stewart EL, Mascaux C, Pham NA, Sakashita S, Sykes J, Kim L, et al. Clinical utility of patient-derived xenografts to determine biomarkers of prognosis and map resistance pathways in EGFR-mutant lung adenocarcinoma. J Clin Oncol. 2015;33:2472–80.
Fichtner I, Rolff J, Soong R, Hoffmann J, Hammer S, Sommer A, et al. Establishment of patient-derived non-small cell lung cancer xenografts as models for the identification of predictive biomarkers. Clin Cancer Res. 2008;14:6456–68.
Yoshida GJ. Applications of patient-derived tumor xenograft models and tumor organoids. J Hematol Oncol. 2020;13:4.
Kummar S, Fogarasi M, Canova A, Mota A, Ciesielski T. Cytokeratin 7 and 20 staining for the diagnosis of lung and colorectal adenocarcinoma. Br J Cancer. 2002;86:1884–7.
Gurda GT, Zhang L, Wang Y, Chen L, Geddes S, Cho WC, et al. Utility of five commonly used immunohistochemical markers TTF-1, Napsin A, CK7, CK5/6 and P63 in primary and metastatic adenocarcinoma and squamous cell carcinoma of the lung: a retrospective study of 246 fine needle aspiration cases. Clin Transl Med. 2015;4:16.
Wu D, Pang Y, Wilkerson MD, Wang D, Hammerman PS, Liu JS. Gene-expression data integration to squamous cell lung cancer subtypes reveals drug sensitivity. Br J Cancer. 2013;109:1599–608.
Watanabe H, Ma Q, Peng S, Adelmant G, Swain D, Song W, et al. SOX2 and p63 colocalize at genetic loci in squamous cell carcinomas. J Clin Invest. 2014;124:1636–45.
Ben-David U, Ha G, Tseng YY, Greenwald NF, Oh C, Shih J, et al. Patient-derived xenografts undergo mouse-specific tumor evolution. Nat Genet. 2017;49:1567–75.
Moro M, Bertolini G, Caserini R, Borzi C, Boeri M, Fabbri A, et al. Establishment of patient derived xenografts as functional testing of lung cancer aggressiveness. Sci Rep. 2017;7:6689.
Nguyen LV, Cox CL, Eirew P, Knapp DJ, Pellacani D, Kannan N, et al. DNA barcoding reveals diverse growth kinetics of human breast tumour subclones in serially passaged xenografts. Nat Commun. 2014;5:5871.
Belderbos ME, Koster T, Ausema B, Jacobs S, Sowdagar S, Zwart E, et al. Clonal selection and asymmetric distribution of human leukemia in murine xenografts revealed by cellular barcoding. Blood. 2017;129:3210–20.
Choi H, Sheng J, Gao D, Li F, Durrans A, Ryu S, et al. Transcriptome analysis of individual stromal cell populations identifies stroma-tumor crosstalk in mouse lung cancer model. Cell Rep. 2015;10:1187–201.
Lambrechts D, Wauters E, Boeckx B, Aibar S, Nittner D, Burton O, et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat Med. 2018;24:1277–89.
Hillinger S, Yang SC, Batra RK, Strieter RM, Weder W, Dubinett SM, et al. CCL19 reduces tumour burden in a model of advanced lung cancer. Br J Cancer. 2006;94:1029–34.
Lin Y, Sharma S, John MS. CCL21 cancer immunotherapy. Cancers. 2014;6:1098–110.
Engelhard VH, Rodriguez AB, Mauldin IS, Woods AN, Peske JD, Slingluff CL Jr. Immune cell infiltration and tertiary lymphoid structures as determinants of antitumor immunity. J Immunol. 2018;200:432–42.
Fukuyama T, Ichiki Y, Yamada S, Shigematsu Y, Baba T, Nagata Y, et al. Cytokine production of lung cancer cell lines: correlation between their production and the inflammatory/immunological responses both in vivo and in vitro. Cancer Sci. 2007;98:1048–54.
Acknowledgements
We express our appreciation to Professor Masafumi Muratani (Department of Genome Biology, Faculty of Medicine, University of Tsukuba and Tsukuba Transborder Medical Center) for research support and kind advice. We thank also Tsukuba Human Biobank Center for sample supply.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Kim, Y., Shiba-Ishii, A., Nakagawa, T. et al. Gene expression profiles of the original tumors influence the generation of PDX models of lung squamous cell carcinoma. Lab Invest 101, 543–553 (2021). https://doi.org/10.1038/s41374-021-00529-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-021-00529-1