Abstract
Most biomedical datasets, including those of ‘omics, population studies, and surveys, are rectangular in shape and have few missing data. Recently, their sample sizes have grown significantly. Rigorous analyses on these large datasets demand considerably more efficient and more accurate algorithms. Machine learning (ML) algorithms have been used to classify outcomes in biomedical datasets, including random forests (RF), decision tree (DT), artificial neural networks (ANN), and support vector machine (SVM). However, their performance and efficiency in classifying multi-category outcomes of rectangular data are poorly understood. Therefore, we compared these metrics among the 4 ML algorithms. As an example, we created a large rectangular dataset using the female breast cancers in the surveillance, epidemiology, and end results-18 database, which were diagnosed in 2004 and followed up until December 2016. The outcome was the five-category cause of death, namely alive, non-breast cancer, breast cancer, cardiovascular disease, and other cause. We analyzed the 54 dichotomized features from ~45,000 patients using MatLab (version 2018a) and the tenfold cross-validation approach. The accuracy in classifying five-category cause of death with DT, RF, ANN, and SVM was 69.21%, 70.23%, 70.16%, and 69.06%, respectively, which was higher than the accuracy of 68.12% with multinomial logistic regression. Based on the features' information entropy, we optimized dimension reduction (i.e., reduce the number of features in models). We found 32 or more features were required to maintain similar accuracy, while the running time decreased from 55.57 s for 54 features to 25.99 s for 32 features in RF, from 12.92 s to 10.48 s in ANN, and from 175.50 s to 67.81 s in SVM. In summary, we here show that RF, DT, ANN, and SVM had similar accuracy for classifying multi-category outcomes in this large rectangular dataset. Dimension reduction based on information gain will increase the model’s efficiency while maintaining classification accuracy.
Similar content being viewed by others
Introduction
Most of the biomedical data are rectangular in shape, including those of ‘omics, large cohorts, population studies, and surveys. Few missing data were present in these datasets. An increasing number of human genomic and survey data have been produced in recent years [1]. Rigorous analyses on these large datasets demand considerably more efficient and more accurate algorithms, which are poorly understood.
Machine learning (ML) algorithms are aimed to produce a model that can be used to perform classification, prediction, estimation, or any other similar task [2, 3]. The unknown dependencies/associations are estimated based on a given dataset and later can be used to predict the output of a new system or dataset [2,3,4]. Therefore, ML algorithms have been used to analyze large biomedical datasets [5,6,7]. Studies have compared the accuracy of several ML algorithms for classifying microarray or genomic data [8,9,10], and show a superior performance of random forests (RF). However, only a few studies accessed the accuracy of ML algorithms in classifying multi-category outcomes, which are important for the more-in-depth understanding of biological and clinical processes. Only when we can accurately predict or identify the factors linked to multicategory outcomes, we can provide patients with more targeted prevention and treatment for the most likely outcome. Hence, we aimed to understand the performance and efficiency of RF, decision tree (DT), artificial neural networks (ANN), and support vector machine (SVM) algorithms in classifying multi-category outcomes of rectangular-shaped biomedical datasets.
As an example, we used a large population-based breast cancer dataset with long-term follow-up data to create a large rectangular dataset. The reasons were: (1) cancer is one of the most common diseases [11]. Knowledge gained by studying cancer can be easily generalized to other biomedical fields; (2) breast cancer is the second most common cancer in U.S. women [11, 12] and would provide a sufficiently large sample size (n = 45,085); (3) breast cancers diagnosed in 2004 had 12 years of follow-up on average and very few missing outcomes/follow-ups were anticipated; (4) breast cancers diagnosed in 2004 would have a moderately good prognosis and their outcomes will be rather diversified (i.e., many patients might not die of breast cancer). Therefore, this study is designed to systematically compare the performance and efficiency of DT, RF, SVM, and ANN algorithms in classifying multicategory causes of death (COD) in a large biomedical dataset (breast cancers).
Methods
Data analysis
We obtained individual-level data from the surveillance, epidemiology, and end results-18 (SEER-18) (www.seer.cancer.gov) SEER*Stat database with treatment data using SEER*Stat software (Surveillance Research Program, National Cancer Institute SEER*Stat software (seer.cancer.gov/seerstat) version <8.3.6>) as we did before [13,14,15]. SEER-18 is the largest SEER database including cases from 18 states and covering near 30% of the U.S. population. The SEER data were de-identified and publicly available. Therefore, this was an exempt study (category 4) and did not require an institutional review board (IRB) review. All incidental invasive breast cancers of SEER-18 diagnosed in 2004 were included and had the follow-up to December 2016. The individual deaths were verified via certified death certificates (2019 data-release). We chose the diagnosis year of 2004 with the consideration of the implementation of the sixth edition of the tumor, node, and metastasis staging manual (TNM6) of the American joint commission on cancer (AJCC) in 2004. Moreover, we only included the primary-cancer cases that had a survival time >1 month, age of 20+ years, and known COD.
The features (i.e., variables) were dichotomized for more efficiency and slightly better performance [7], while the actual values were also tuned for and analyzed using RF models. A total of 54 features were included (Supplementary Table 1). We conducted correlation analyses and produced a correlation matrix (Supplementary Fig. 1) to identify the closely correlated factors. The outcomes of the classification models were the patient’s five-category COD. The COD was originally classified using SEER’s recodes of the causes of death, which were collected through death certificates of deceased patients (https://seer.cancer.gov/). We simplified the SEER COD into five categories based on the prevalence of COD [16,17,18], including alive, non-breast cancer, breast cancer, cardiovascular disease (CVS), and other cause.
The most common task of ML techniques in the learning process is classification [19]. The tenfold cross-validation approach was used to tune all models, which is also termed as model optimization (see Supplementary methods) as described before [20, 21]. This approach is an approximation of leave-p-out cross-validation has the advantage of repeat the process to reduce variance and being less sensitive to the partitioning of the data than the holdout method. Briefly, the samples were randomly divided into ten same-size subsamples, among which one subsample remained as the validation set and the other nine as the test set. The (cross-)validation would be performed ten times as the test subsample was shuffled among the ten subsamples. The mean of the cross-validation’s performances would be calculated and used as the performance of the model. Multinomial logistic regression (MLR) and the ML analyses were carried out using MATLAB (version 2018a, MathWorks, Natick, MA).
Model tuning
The detailed model tuning process is described in Supplementary material. Several DT methods, such as CHAID, CART, and exclusive CHAID are available with MATLAB [22]. We used classification and regression tree(CART) to predict the categories, using the Gini index as the split criterion and 100 iterations for each run (Supplementary Fig. 2). There are no default RF packages/toolboxes in MATLAB’s own toolbox. We thus used the Randomforest-Matlab open-source toolbox developed by Jaiantilal et al. [23, 24].
In tuning RF models, the parameter nTrees, which was to set how many trees in a random forest, and may have an impact on the classification results. We set the value of nTrees from 1 to 600 separately, the results show that if this parameter is not too few (greater than ten), the accuracy of recognition can reach 69–70% (Supplementary Fig. 3). Therefore, we set this parameter to 136 in RF-based analyses. The value of Mtry node for the best model performance was identified by setting the parameter value from 1 to 20 with 1 as the interval. We found 5 was the value, which was indeed consistent with the default value generated using the MATLAB model’s default (i.e., Mtry = floor(sqrt(size(P_train,2)))).
Among different training algorithms of ANN, we used the Trainscg algorithm because it is the only conjugate gradient method that did not require a linear search. The number of input layer nodes is 54 and the number of output layer nodes is 6. To tune the ANN model, we conducted experiments of either single hidden or double hidden layers, with the node numbers ranged from 5 to 100. According to the tuning results of accuracy and mean squared error (MSE), we would set the model to double hidden layers, and the number of layers for the highest accuracy and lowest MSE.
We used the multi-class error-correcting output codes (ECOC) model the SVM modeling which allows classification in more than two classes; and the MATLAB fitcecoc function that creates and adjusts the template for SVM [25]. The Kernel functions considered in the SVM were: linear, radial basis function, Gaussian, and polynomial.
Performance analysis
We analyzed the performance metrics of each proposed model, including accuracy, recall, precision, F1 score, and specificity [26, 27]. They were defined as follows:
True positive (TP) and true negative (TN) were defined as the number of samples that are classified correctly. False positive (FP) and false negative (FN) were defined as the number of samples that are misclassified into the other mutational classes [26, 27]. The specificity or true negative rate (TNR) is defined as the percentage of mutations that are correctly identified:
The receiver operating curve (ROC) is a graph where recall is plotted as a function of 1-specificity. It can more objectively measure the performance of the model itself [25]. The model performance was also evaluated using the area under the ROC, which is denoted the area under the curve (AUC). An AUC value close to 1 highlight a high-performance model, while an AUC value close to 0 demonstrate a low-performance model [28, 29]. AUC is independent of the class prior distribution, class misclassification cost, and classification threshold, which can more stably reflect the model’s ability to sort samples and characterize the overall performance of the classification algorithm. The formula used to determine the AUC can be written as follows [29, 30]:
where P is the total number of positive class and N is the total number of negative class.
Dimension reduction based on the information entropy and information gain
Information entropy is an indicator to measure the purity of the sample set. The formula is as follows:
The measure of information gain is to see how much information a feature can bring to the classification system. The more information it brings, the more important the feature. Information gain IG(A) is the compute of the difference in entropy from start to end the set S is split on an attribute A, the information gain is defined as follows [31]:
where H(S) is the entropy of set S, T is the subsets created from splitting set S by attribute A such that \(S = U_{t \in T}t\), p(t) is the proportion of the number of elements in t to the number of elements inset S, and H(t) is the entropy of subset T.
The information gain of a feature can indicate how much information it brings to the classification system and can be used as a feature weight. When the model uses more features the classification time will be longer. Arbitrarily reducing the characteristics will likely reduce classification accuracy. Therefore, we screened the features based on the calculated information gains to achieve a balance between run time and classification accuracy. We then step-wise deleted the features of the least information of gain, which were likely the least important. Because DT and RF were both ensemble-based algorithms and had similar performances, we conducted dimension reduction with RF, ANN, and SVM models and expect similar results with DT models.
Results
Dataset characteristics and the model tuning
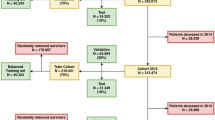
Of the 52,818 samples, we step-wise excluded the 5294 (~10%) cases missing tumor-grade data (_grade), 1770 (3.4%) missing survival-time and 352 (0.6%) missing laterality data (lat_bi), and 317 (0.6%) missing data of TNM6 N category. Overall, we included 45,085 cases of breast cancer diagnosed in 2004 that were included in the SEER-18 and had no missing data in all features (Table 1). The variables were dichotomized into 54 features. Likely due to the unique nature of the pathological and socioeconomic factors, the correlation analyses showed only a few features had a correlation coefficient >0.95 or <−0.95 (Supplementary Fig. 1).
For DT models, the minimum number of leaf nodes with the best performance of the DT range 20–300 and peak at 93 (Supplementary Fig. 2). Therefore, the minimum number of samples contained in the leaf node was 93 for optimization. The overall classification accuracy could reach 67.3%, which was about 4% higher than the original DT, and the cross-validation error has also decreased. However, due to the uneven distribution of the data samples, some categories such as non-breast cancer and other cause were pruned, causing the loss of some data information.
For RF models, the parameter nTrees set how many trees in a RF and may have an impact on the classification results. The Mtry is the number of features randomly sampled as candidates at each split and was optimized (Supplementary methods). After tuning the models, we set the nTrees parameter to 136 in RF-based analyses with the best Mtry node value of 6 (Supplementary Fig. 3).
According to the tuning results of accuracy and MSE in ANN models, when the number of layers was greater than 20, the models’ performance appeared stabilized (Table 2). Therefore, we set the model to double hidden layers, and the number of layers is 40.
For the SVM models of linear, radial basis function, Gaussian or polynomial function, we found the linear kernel function had the highest accuracy (69.06%) and shortest run-time (175.50 s, Table 3), with a one-vs-one approach.
Performance analysis results
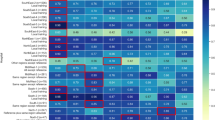
Based on the confusion matrices (Fig. 1), the 4 ML models appeared to have similar performance (Table 4) and all were more accurate than the MLR. The best classification accuracy of DT, RF, ANN, and SVM models in this study was 69.21%, 70.23%, 70.16%, and 69.06%, respectively, and higher than 68.12% of a conventional statistical algorithm (MLR). However, to evaluate the pros and cons of a model, it is not enough to just look at the accuracy. The values of recall, TNR, and F1 can specifically reflect the classification of each category. Further comparing the metrics among the four models show that the precision-recall and F1 in classifying breast cancer by RF model were better than the DT, ANN, and SVM models. This is mainly because the RF model is an integrated learning algorithm; through the voting mechanism, one can balance a certain error. Compared with ML algorithms, MLR had lower overall accuracy and a much lower recall (1.13% vs ~50%) and a lower specificity (Table 4) for predicting the death of breast cancer.
ROC analysis results
After a comparative analysis of the above performance metrics, we found that the RF model was superior to the DT, ANN, and SVM models. However, in the classification of other causes, the ANN model has a higher recognition rate than the RF model, and the F1 values of non-breast cancer cannot be calculated. Therefore, we used the ROC curve to further analyze non-breast cancer and other cause in RF and ANN models.
The AUC of the 5 COD in the RF model was overall lower than those in the ANN model (Fig. 2). Because the AUC index can measure the performance of the model more objectively, the results show that the overall performance of the ANN model is similar or better than that of the RF model.
Dimension reduction based on the information entropy and information gain
This dataset had 54 categorical/binary features. Based on the training datasets, the information entropy and information gain of 54 features were calculated (Supplementary Table 3). Using information entropy and information gain, we obtained the following important features: age >65; surgery; TNM6 metastasis subgroup1; AJCC stage 4; TNM6 metastasis subgroup2; surgery–other; TNM6 tumor subgroup1; TNM6 tumor subgroup2; AJCC stage 1; surgery-lumpectomy; TNM6 lymph node subgroup5; and TNM6 lymph node subgroup1. Then we characterized the key features in a step-wise fashion (Supplementary Fig. 4).
We successfully reduced the data dimension based on information gain and shortened the run times in RF, ANN, and SVM models, while maintaining the overall classification accuracy (Table 5 and Supplementary Tables 3–5). Removal of features with low information gain (0.0000–0.0005) in RF models led to a slight increase in the alive class and the overall accuracy rates, while no accuracy changes in CVS and breast cancer classes. The classification of CVS was always 100%; accuracy rate of alive class and the overall had a slight improvement, respectively about 0.5% and 0.3%; breast cancer and CVS had no significant changes; non-breast cancer accuracy was slightly reduced, and the classification effect is unstable. Therefore, the features with low information gain (0.0000–0.0005) may be considered as redundant features, and deleted in the models, while the running times were scientifically reduced. We also found similar changes in ANN and SVM models.
Feature importance in RF models using the data of convention encoding or one-hot encoding
Our previous works have shown that one-hot encoding (dichotomization of features) led to a slight increase in prediction accuracy of the RF model on prostate cancer using Stata [7]. Consistent with that finding, our tuned RF model on actual values had a prediction accuracy of 69.70%, which was slightly lower than the accuracy of 70.23% on dichotomized features (Supplementary Table 6). However, the top-five important features were different in the two models, except age 65+ years (vs <65 years, Supplementary Figs. 5 and 6).
Discussion
We here compared the performance and efficacy of DT, RF, ANN, and SVM in classifying five-category outcomes of a large rectangular database (54 features and ~45,000 samples). The accuracy in classifying five-category COD with DT, RF, ANN, and SVM was 69.21%, 70.23%, 70.16%, and 69.06%, respectively. It is noteworthy that the accuracy in classifying five-category outcomes is much more difficult than classifying binary-category outcomes since it depends on four sequential classification processes in one-over-all or one-over-rest algorithms (i.e., final accuracy is the accuracy multiplication for four times). Moreover, based on the information entropy and information gain of feature values, we could reduce the feature number in a model to 32 and maintained a similar accuracy, while the running time decreased from 55.57 s for 54 features to 25.99 s for 32 features in RF, from 12.92 s to 10.48 s in ANN and from 175.50 s to 67.81 s in SVM. The DT algorithm was not tested after dimension reduction for its lower performance than RF and its theoretical framework (ensemble-based) like RF.
Few studies to our knowledge investigated the dimension reduction of multicategory classification. A two-stage approach has been shown to effectively and efficiently select features for balanced datasets [32], but no specific reduction in run time was reported. We here show that dimension reduction and efficiency improvement can be achieved by removing features of low to medium information gain (<0.0005) in RF, ANN, and SVM models, which apparently have little effect on the overall classification performance. Such a strategy may be applied to other ML models in classifying unbalanced large rectangular datasets, while caution should be used when classifying outcomes in a balanced dataset.
Consistent with a previous report [7], one-hot encoding (i.e., dichotomization of the features) produced slightly better prediction accuracy in our data than conventional encoding and also has the advantage of no need for normalization. The top-five important features in the models based on the dichotomized and actual-value features differed considerably except age. This finding further confirms the important role of age in modeling five-category COD. However, five of the top ten important features shared in these two models, including advanced age, pathologic staging, surgery status, N category, and histology type. These features thus should be carefully examined, recorded, and considered for predicting or preventing five-category COD in breast cancer patients.
The four included ML algorithms each have their own theoretical frameworks. DT is a logical-based ML approach [33]. The structure of the DT is similar to a flowchart. Using top-down recursion, the classification tree produces the category output. Starting from the root node of the tree, test and compare property values on its internal node, then determine the corresponding branch, and finally reach a conclusion in the leaf node of the DT. This process is repeated at each node of the tree by selecting the optimal splitting features until the cut-off value is reached [34]. A leafy tree tends to overtrain, and its test accuracy is often far less than its training accuracy. By contrast, a shallow tree can be more robust and be easy to interpret [35].
DT works by learning simple decision rules extracted from the data features. But RF is a combination of tree predictors such that each tree depends on the values of a random vector sampled independently, and with the same distribution for all trees in the forest [36]. When RF used for a classification algorithm, the deeper the tree is, the more complex the decision rules and the fitter the model [36, 37]. The generalization error of a forest of tree classifiers depends on the strength of the individual trees in the forest and the correlation between them. Random decision forests overcome the problem of overfitting of the DTs and are more robust with respect to noise.
ANN technique is one of the artificial intelligence tools. It is a mathematical model that imitates the behavior characteristics of animal neural networks [38]. This kind of network carries on distributed parallel information processing by adjusting the connection between a large number of internal nodes, so as to achieve the purpose of processing information. After repeated learning and training, network parameters corresponding to the minimum error are determined, and the ANN model classifies the output automatically from the dataset.
SVM is another popular ML tool based on statistical learning theory, which was first proposed by Vapnik and his colleagues [39, 40]. Unlike traditional learning methods, SVMs are approximate implementations of structural risk minimization methods. The input vector is mapped to a high-dimensional feature space through some kind of non-linear mapping which was selected in advance. An optimal classification hyperplane is constructed in this feature space, to maximize the separation boundary between the positive and negative examples [39, 40]. Support vectors are the data points closest to the decision plane, and they determine the location of the optimal classification hyperplane.
The 4 ML algorithms had different strengths and weaknesses, while all outperformed conventional statistical algorithm (MLR). The RF algorithm in our study seems to have the best overall performance for its lack of being unable to classify some CODs and the best overall accuracy. Despite the similar classification accuracy (~70%), the DT algorithm could not accurately classify the non-breast cancer group. Given the similar theoretical framework, we did not access its performance after dimension reduction. The ANN algorithm in our study is most efficient before and after dimension reduction. Surprisingly, we also notice a small increase in accuracy after dimension reduction which warrants further investigation. The SVM algorithm in our study appears very sensitive to the subgroup size (i.e., number of samples) and was not able to classify two of the five COD, although it also acceptable classification accuracy.
The study’s limitations should be noted when applying our findings to other databases. First, this type of rectangular database is typical in survey and population-study, but not so in computational biology. The major difference is the large p in ‘omics datasets vs the large n in epidemiological datasets, which referred to feature number and sample number, respectively. Second, some of the outcomes were not accurately classified. It is likely owing to the unbalanced outcome distribution. On the other hand, such an undesired situation reflexes real-world evidence/experience. Further studies are needed to improve the classification accuracy in the classes of fewer samples. Third, our works were exclusively based on the MATLAB platform and may not be applicable to other platforms such as R or python. Fourth, a molecular subtype of the cancer was not available due to the lack of human epidermal growth factor receptor 2 (Her2) data. Fifth, the prediction accuracy of the ML algorithms was moderately acceptable (~70%), although ML had much better recall and sensitivity than MLR. The challenge in predicting multi-category outcomes remains outstanding, despite the use of tuned ML algorithms. Thus, future works are needed to improve the prediction performance. Finally, ideally, we should use a large database to validate our models, but it is very difficult to curate and apply the tuned models to another large database that is similar to the SEER database.
In summary, we here show that RF, DT, ANN, and SVM algorithms had similar accuracy, but outperformed MLR, for classifying multi-category outcomes in this large rectangular dataset. Dimension reduction based on information gain will significantly increase the model’s efficiency while maintaining classification accuracy.
References
Liu DD, Zhang L. Trends in the characteristics of human functional genomic data on the gene expression omnibus, 2001–2017. Lab Investig. 2019;99:118–27.
Cruz JA, Wishart DS. Applications of machine learning in cancer prediction and prognosis. Cancer Inf. 2007;2:59–77.
Kourou K, Exarchos TP, Exarchos KP, Karamouzis MV, Fotiadis DI. Machine learning applications in cancer prognosis and prediction. Comput Struct Biotechnol J. 2015;13:8–17.
Bishop CM. Pattern recognition and machine learning. New York, NY, USA: Springer; 2006.
Chow ZL, Thike AA, Li HH, Nasir NDM, Yeong JPS, Tan PH. Counting mitoses with digital pathology in breast phyllodes tumors. Arch Pathol Lab Med. 2020;144:1397–400.
Koo J, Zhang J, Chaterji S. Tiresias: context-sensitive approach to decipher the presence and strength of MicroRNA regulatory interactions. Theranostics. 2018;8:277–91.
Wang J, Deng F, Zeng F, Shanahan AJ, Li WV, Zhang L. Predicting long-term multicategory cause of death in patients with prostate cancer: random forest versus multinomial model. Am J Cancer Res. 2020;10:1344–55.
Maniruzzaman M, Jahanur Rahman M, Ahammed B, Abedin MM, Suri HS, Biswas M, et al. Statistical characterization and classification of colon microarray gene expression data using multiple machine learning paradigms. Comput Methods Programs Biomed. 2019;176:173–93.
Pirooznia M, Yang JY, Yang MQ, Deng Y. A comparative study of different machine learning methods on microarray gene expression data. BMC Genom. 2008;9:S13.
Haibe-Kains B, Desmedt C, Sotiriou C, Bontempi G. A comparative study of survival models for breast cancer prognostication based on microarray data: does a single gene beat them all? Bioinformatics. 2008;24:2200–8.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Goetz MP, Gradishar WJ, Anderson BO, Abraham J, Aft R, Allison KH, et al. NCCN guidelines insights: breast cancer, version 3.2018. J Natl Compr Canc Netw. 2019;17:118–26.
Chavali LB, Llanos AAM, Yun JP, Hill SM, Tan XL, Zhang L. Radiotherapy for patients with resected tumor deposit-positive colorectal cancer: a surveillance, epidemiology, and end results-based population study. Arch Pathol Lab Med. 2018;142:721–9.
Yang M, Bao W, Zhang X, Kang Y, Haffty B, Zhang L. Short-term and long-term clinical outcomes of uncommon types of invasive breast cancer. Histopathology. 2017;71:874–86.
Mayo E, Llanos AA, Yi X, Duan SZ, Zhang L. Prognostic value of tumour deposit and perineural invasion status in colorectal cancer patients: a SEER-based population study. Histopathology. 2016;69:230–8.
Bevers TB, Helvie M, Bonaccio E, Calhoun KE, Daly MB, Farrar WB, et al. Breast cancer screening and diagnosis, version 3.2018. J Natl Compr Cancer Netw. 2018;16:1362–89.
Afifi AM, Saad AM, Al-Husseini MJ, Elmehrath AO, Northfelt DW, Sonbol MB. Causes of death after breast cancer diagnosis: a US population-based analysis. Cancer. 2020;126:1559–67.
Clough-Gorr KM, Thwin SS, Stuck AE, Silliman RA. Examining five- and ten-year survival in older women with breast cancer using cancer-specific geriatric assessment. Eur J Cancer. 2012;48:805–12.
Amrane M, Oukid S, Gagaoua I, Ensari T. Breast cancer classification using machine learning. stanbul: Electric Electronics, Computer Science, Biomedical Engineeringsʼ Meeting (EBBT); 2018. p. 1–4. https://doi.org/10.1109/EBBT.2018.8391453.
Mao Y, Fu Z, Dong L, Zheng Y, Dong J, Li X. Identification of a 26-lncRNAs risk model for predicting overall survival of cervical squamous cell carcinoma based on integrated bioinformatics analysis. DNA Cell Biol. 2019;38:322–32.
Dong RZ, Yang X, Zhang XY, Gao PT, Ke AW, Sun HC, et al. Predicting overall survival of patients with hepatocellular carcinoma using a three-category method based on DNA methylation and machine learning. J Cell Mol Med. 2019;23:3369–74.
Grzesiak W, Zaborski D. Examples of the use of data mining methods in animal breeding. Adem Karahoca, editor. Data mining applications in engineering and medicine. London, UK: IntechOpen Limited; 2012; 303–24.
Wang XC, Shi F, Yu L, Li Y. Cases analysis of MATLAB neural network. Beijing: Beijing University of Aeronautics and Astronautics. 2009. p. 59–62.
Jaiantilal A. Classification and regression by randomforest-matlab. (2009, 2012). https://code.google.com/archive/p/randomforest-matlab/ Accessed 22 July 2020.
Gonçalves CB, Leles ACQ, Oliveira LE, Guimaraes G, Cunha JR, Fernandes H. Machine learning and infrared thermography for breast cancer detection. Multidiscipl Digit Publish Inst Proc. 2019;27:45.
Sokolova M, Japkowicz N, Szpakowicz S. Beyond accuracy, F-score and ROC: a family of discriminant measures for performance evaluation. Australasian joint conference on artificial intelligence. 2006; 1015–21.
Aruna S, Rajagopalan SP, Nandakishore LV. Knowledge based analysis of various statistical tools in detecting breast cancer. Comput Sci Inf Technol. 2011;2:37–45.
Youssef AM, Pradhan B, Jebur MN, El-Harbi HM. Landslide susceptibility mapping using ensemble bivariate and multivariate statistical models in Fayfa area, Saudi Arabia. Environ Earth Sci. 2015;73:3745–61.
Costache R, Hong H, Wang Y. Identification of torrential valleys using GIS and a novel hybrid integration of artificial intelligence, machine learning and bivariate statistics. Catena. 2019;183:104179.
Hong H, Liu J, Bui DT, Pradhan B, Acharya TD, Pham BT, et al. Landslide susceptibility mapping using J48 decision tree with AdaBoost, bagging and rotation forest ensembles in the Guangchang area (China). Catena. 2018;163:399–413.
Anyanwu MN, Shiva SG. Comparative analysis of serial decision tree classification algorithms. Int J Comput Sci Secur. 2009;3:230–40.
Chung D, Keles S. Sparse partial least squares classification for high dimensional data. Stat Appl Genet Mol Biol. 2010;9: Article17. https://www.degruyter.com/document/doi/10.2202/1544-6115.1492/html.
Safavian SR, Landgrebe D. A survey of decision tree classifier methodology. IEEE Trans Syst Man Cybern. 1991;21:660–74.
Lan T, Hu H, Jiang C, Yang G, Zhao Z. A comparative study of decision tree, random forest, and convolutional neural network for spread-F identification. Adv Space Res. 2020;65:2052–61.
Garcia Leiv R, Fernandez AnA, Mancus V, Casari P. A novel hyperparameter-free approach to decision tree construction that avoids overfitting by design. IEEE Access. 2019;7:99978–87.
Breiman L. Random forests. Mach Learn. 2001;45:5–32.
Nguyen C, Wang Y, Nguyen HN. Random forest classifier combined with feature selection for breast cancer diagnosis and prognostic. J Biomed Sci Eng. 2013;6:551–60.
Jain AK, Jianchang M, Mohiuddin KM. Artificial neural networks: a tutorial. Computer. 1996;29:31–44.
Cortes C, Vapnik V. Support-vector networks. Mach Learn. 1995;20:273–97.
Fradkin D, Schneider D, Muchnik I. Machine learning methods in the analysis of lung cancer survival data. DIMACS technical report 2005–35. 2006.
Acknowledgements
We thank Lingling Han at Shenzhen Horb Technology Corporate, Ltd. for invaluable discussions and comments.
Author information
Authors and Affiliations
Contributions
FD, CC, and LZ designed the study, FD and JH conducted the study and drafted the manuscript, all authors discussed, revised, and edited the manuscript, and LZ supervised the work.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Deng, F., Huang, J., Yuan, X. et al. Performance and efficiency of machine learning algorithms for analyzing rectangular biomedical data. Lab Invest 101, 430–441 (2021). https://doi.org/10.1038/s41374-020-00525-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-020-00525-x
This article is cited by
-
A Review of Machine Learning Algorithms for Biomedical Applications
Annals of Biomedical Engineering (2024)
-
Uncovering potential diagnostic biomarkers of acute myocardial infarction based on machine learning and analyzing its relationship with immune cells
BMC Cardiovascular Disorders (2023)
-
Multimetric feature selection for analyzing multicategory outcomes of colorectal cancer: random forest and multinomial logistic regression models
Laboratory Investigation (2022)
-
Assessment of agricultural prospects in relation to land use change and population pressure on a spatiotemporal framework
Environmental Science and Pollution Research (2022)