Abstract
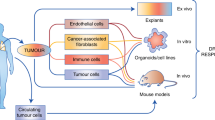
Patient-derived explants (PDEs) represent the direct culture of fragments of freshly-resected tumour tissue under conditions that retain the original architecture of the tumour. PDEs have advantages over other preclinical cancer models as platforms for predicting patient-relevant drug responses in that they preserve the tumour microenvironment and tumour heterogeneity. At endpoint, PDEs may either be processed for generation of histological sections or homogenised and processed for ʻomic’ evaluation of biomarker expression. A significant advantage of spatial profiling is the ability to co-register drug responses with tumour pathology, tumour heterogeneity and changes in the tumour microenvironment. Spatial profiling of PDEs relies on the utilisation of robust immunostaining approaches for validated biomarkers and incorporation of appropriate image analysis methods to quantitatively and qualitatively monitor changes in biomarker expression in response to anti-cancer drugs. Automation of immunostaining and image analysis would provide a significant advantage for the drug discovery pipeline and therefore, here, we have sought to optimise digital pathology approaches. We compare three image analysis software platforms (QuPath, ImmunoRatio and VisioPharm) for evaluating Ki67 as a marker for proliferation, cleaved PARP (cPARP) as a marker for apoptosis and pan-cytokeratin (CK) as a marker for tumour areas and find that all three generate comparable data to the views of a histomorphometrist. We also show that Virtual Double Staining of sequential sections by immunohistochemistry results in imperfect section alignment such that CK-stained tumour areas are over-estimated. Finally, we demonstrate that multi-immunofluorescence combined with digital image analysis is a superior method for monitoring multiple biomarkers simultaneously in tumour and stromal areas in PDEs.
Similar content being viewed by others
Introduction
Robust methods for preclinical testing of novel anti-cancer drugs that have strong predictive value for clinical translation are lacking [1]. While numerous in vitro, in vivo and ex vivo models are used such as organoids, cell lines and patient-derived xenograft models, many of these rely on deconstruction of the original tumour and segregation of the patient-specific stromal cells. Patient-derived explants (PDEs) on the other hand, do not involve tumour deconstruction and therefore allow for drug testing in intact human tumours, thus allowing for patient-specific responses, while retaining the tumour architecture [1,2,3,4]. Historically, most endpoints used to analyse drug responses have involved digestion or homogenisation of the PDEs post drug treatment and assessment of response by methods such as the MTT (3-(6)-2, 5-diphenyl tetrazolium bromide) assay [5, 6] or molecular approaches such as qRT-PCR [7, 8]. These approaches, however, remove intra-tumour heterogeneity and cell-specific spatial analysis of drug response.
Immunohistochemical (IHC) analysis of biomarker expression of formalin fixed paraffin embedded (FFPE) sections has been an approach more recently used to assess in situ drug responses in PDEs [2, 4]. Previous work from our laboratory using quantitative assessment of IHC staining for cPARP as a marker for apoptosis and Ki67 as a marker for proliferation has shown that non-small cell lung cancer (NSCLC) PDEs respond to the chemotherapeutic drug cisplatin in a manner that is predictive of clinical outcome [2]. The workflow used to quantitate cPARP or Ki67 staining in this study utilised a manual and laborious process involving the imaging of multiple fields of view (FoV) by bright-field microscopy followed by manually stitching the images together and calculation of the staining index using ImmunoRatio. Non-tumour stromal regions were segregated by manual means and were not included in the analysis such that changes in the tumour microenvironment in response to the drug could not be monitored [2].
There is now a plethora of whole-slide scanners and analysis software available to digitise and analyse whole tissue sections as opposed to imaging FoVs. Also, multiplexed immunofluorescence (mIF) has emerged as a powerful tool in place of traditional chromogenic immunohistochemistry (IHC) in identifying and validating predictive biomarkers [9, 10]. For PDEs to be adopted more widely in the drug discovery pipeline, advances in slide scanning and analysis methodologies need to be coupled to the PDE workflow. Here, we investigate the advantages of digitally capturing images of PDE sections stained with key biomarkers using a whole-slide scanner, and compare images using three digital Pathology solutions: VisioPharm [11], ImmunoRatio [12] and QuPath [13]. We evaluate the inter-platform agreeability of the three software platforms and compare these to the opinions of a trained histomorphometrist. Further to this, we have compared traditional IHC staining with DAB to mIF approaches for monitoring biomarker outputs in the analysis of PDEs at endpoint.
Materials and methods
PDE processing
NSCLC samples were collected fresh from surgery (Ethical approval: LREC: 07/MRE08/07). PDEs were generated from fresh tumours and cultured as previously described [2]. PDEs were incubated for 24 h with 50 μM cisplatin in 0.1% (v/v) dimethylformamide (DMF) or DMF alone as vehicle control. Following treatment, PDEs were transferred to fresh 6-well plates containing 4% (w/v) paraformaldehyde for 20–24 h, transferred onto sponges soaked in 70% (v/v) IMS and placed in histology cassettes. PDEs were embedded in paraffin blocks and 4 μm sequential serial sections generated.
Immunostaining
FFPE tissue sections were stained with standard H&E approaches. Where sequential sections were used, no trimming occurred between tissue sections. For both IHC and mIF, primary antibodies used were: Ki67 (Clone MIB-1: DAKO 1:1000); pan-Cytokeratin (CK; Clone MNF116: DAKO 1:100); cPARP (Clone [E51]: Abcam 1:2000) diluted in PBS. IHC sections that utilised DAB for detection were processed as described previously using the Novolink Polymer Detection System (Leica Biosystems) [2]. For mIF, tissue sections were dewaxed and rehydrated following standard procedures. All steps were performed at room temperature. Heat-induced epitope retrieval was performed by submerging slides in 20 mM citrate buffer (pH 6) and microwaving on full power (800 W) for 20 min. Slides were then washed in PBS, and incubated with Protein Block (Perkin Elmer, ARD1001EA) for 10 min, followed by 30 min with the Ki67 primary antibody. Slides were then washed with PBS twice and incubated with HRP-Polymer (secondary anti-Mouse and anti-Rabbit antibodies; Perkin Elmer ARH1001EA) for 30 min. Slides were washed twice with PBS, followed by incubation with Opal-520 (1:100) diluted in 1× Amplification Diluent (Perkin Elmer) for 10 min and then further washed twice in PBS. For subsequent antibody-fluorophore pairs, previous antibodies were stripped by microwaving slides in citrate buffer at full power for 10 min. Successive antibody incubations were: CK paired with Opal-570, and cPARP paired with Opal-690. After the final antibody-fluorophore pair, slides were washed thoroughly in deionised water, and counterstained for 5 min with 6 μM DAPI, followed by two washes in deionised water. Slides were mounted with VectaShield Antifade mounting medium, cover slips added and sealed with nail varnish. Whole-slide scans of all stained slides were captured with a Hamamatsu NanoZoomer XR. For mIF slides, Opal-520, Opal-570 and Opal-690 were detected using FITC, TRITC, and Cy-5 excitation and emission filters respectively. Standard DAPI filters were used for excitation/emission of DAPI. DAB-stained slides were imaged using standard bright-field procedures. All scans were acquired with a ×40 objective lens.
Image analysis
Chromogenic whole-slide scans of Ki67 and cPARP-stained sections were manually evaluated by a histomorphometrist to generate % positivity [14]; these observations were made across the digitised whole-slide scans. DAB-stained images were imported into QuPath or VisioPharm and individual nuclei were segmented based on haematoxylin staining in the case of QuPath, and haematoxylin and DAB staining in the case of VisioPharm. Individual cells were then identified as positive or negative based on Ki67 or cPARP labelling as previously described [11, 13]. The same histomorphometrist as above oversaw the outputs from the digital platforms and agreed that the classification settings used yielded true and representative results within the limitations of each platform, with algorithm variables tuned where appropriate to make results more accurate. ImmunoRatio uses a colour deconvolution algorithm to calculate the percentage of positively stained Ki67 and cPARP nuclear area and was used as described previously [12]. For Virtual Double Staining (VDS), Ki67 and cPARP-stained slides were aligned with an adjacent slide stained with CK allowing the segregation of tissue on the Ki67- and cPARP-stained slides into tumour and stroma [11]. For mIF-stained sections, whole-slide images were imported and analysed in VisioPharm. PDEs were identified and regions separated into tumour and stroma based on CK staining. The samples used in this study displayed very little necrosis with these areas also being separated into tumour and stroma based on cytokeratin staining. Nuclear segmentation was achieved by using a customised application. Ki67 and cPARP positive cells were identified based on fluorescent nuclear signal intensity and manually tuned to obtain accurate results.
Statistical analysis
Unless otherwise stated, all statistics were calculated using R for windows (v 3.6.1). For bivariate linear regression, Pearson’s correlation was performed using the ggpubr package, with p < 0.05 considered as statistically significant. For paired analysis, boxplots were generated using ggpubr and evaluated using Wilcoxon-paired test, with p < 0.05 considered as statistically significant. To compare platforms and describe agreement between two methods, Bland–Altman (BA) plots were generated [15]. To evaluate inter-platform agreeability, Passing–Bablok (PB) regression was fitted to the data [16]. PB was fitted and plotted in R using the mcr package. BA plots were generated using Stata version 15.1 and R using the ggplot package. For PB analysis, if the range in the confidence intervals (CIs) from the estimated intercept contained a 0, it was accepted that there was no constant difference between analytical methods. If the range in the CIs did not span 0, it was concluded that there was a constant difference between analytical methods. If the range in the CIs from the estimated slope spanned 1, it was concluded that there was no proportional difference between platforms. PB statistics expressed as X.XX(Y.YY_Z.ZZ) where X.XX indicates intercept or slope, and Y.YY_Z.ZZ indicates the lower and upper confidence intervals.
Results
Patient-derived explants and staining methodologies
We have previously shown that sensitivity of NSCLC PDEs to cisplatin predicts patient outcome [2]. In this previous study, cell death (cPARP) or proliferation (Ki67) responses in PDEs were evaluated using IHC and bright-field microscopy of separately captured multiple FoVs. These separate FoVs were then ‘stitched’ together to generate an image of a whole PDE, the flanking stromal tissue was removed and the ImageJ plugin ImmunoRatio was used to calculate the percentage of brown (DAB) over blue (haematoxylin) to generate a proliferation index and percentage value for apoptosis. This approach was both labour-intensive and time-consuming and therefore we have sought to improve the PDE workflow.
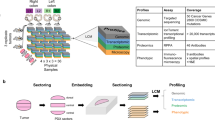
To compare image analysis approaches, in the present study, serial sections were generated from NSCLC PDEs. As indicated in Fig. 1A, parallel sections were then stained sequentially with either H&E, with three different antibodies (Ki67, cPARP, CK) using IHC or with the same three antibodies combined on one section using mIF. Chromogenic IHC stains are typically limited to the staining and imaging of one or two biomarkers per section. While this can be overcome by ‘multiplexed IHC consecutive staining on a single slide (MICSSS)’, this process is typically low throughput, with correspondingly high turnaround times [17, 18]. Although automated instrumentation can improve staining throughput with MICSSS somewhat, the time taken for analysis is still rate limiting [17, 18]. By contrast, mIF allows for easy detection of multiple markers on one section, generating a ‘merged’ image that integrates the staining for all biomarkers (Fig. 1A).
A Serial sections and sequential staining of parallel sections with: H&E, Ki67 using IHC, CK using IHC, cPARP using IHC and all three biomarkers together with DAPI by mIF. B, C Serial sections from vehicle (B) or cisplatin (C) treated NSCLC PDEs of the lung adenocarcinoma subtype were stained with H&E, with the three biomarkers (Ki67, CK, cPARP) on sequential sections using IHC, or with the three biomarkers on the same section using mIF. The merged image generated from mIF shows integration of Ki67 (green), CK (yellow), cPARP (red) with DAPI (blue) staining.
Representative FoVs for both vehicle and cisplatin-treated PDEs with the three approaches are shown in Fig. 1B and C. As shown previously [2], using both IHC and mIF, vehicle-treated NSCLC PDEs displayed high levels of Ki67 staining compared to cisplatin-treated samples, indicating suppression of proliferation by cisplatin. There were low levels of cPARP staining in the vehicle-treated samples compared to cisplatin-treated samples, indicating strong apoptosis induction by cisplatin.
Comparison of digital pathology solutions for endpoint PDE analysis
To improve the PDE endpoint analysis workflow, we investigated the use of digital pathology platforms for biomarker evaluation and quantitation. Most image analysis platforms initially require the accurate identification (segmentation) of cell nuclei for subsequent biomarker evaluation. However, different software packages use differing strategies for this step, leading to discrepancies in final quantification. We therefore sought to evaluate data obtained from three different packages to identify those that most closely matched a histomorphometrist’s judgement.
Slides stained by IHC for the three biomarkers (Ki67, cPARP, CK) were digitised using a whole-slide scanner at ×40 magnification (Fig. 1) and then analysed using three available software packages: QuPath, ImmunoRatio and VisioPharm. For simplicity, these platforms have been labelled as Platforms 1, 2 and 3, respectively. Platforms-1 and Platform-3 can both segment individual nuclei, which allows for per cell level analyses. In contrast, Platform-2 uses a colour deconvolution algorithm for separating the staining components and generates a ratio of DAB/haematoxylin stained areas only. Platform-2 (Immunoratio) was used in our previous study [2] and served as a benchmark from which to improve PDE endpoint analysis. Raw images of a Ki67-stained section along with digitised images generated with each of the three platforms are shown in Fig. 2A. For all three platforms, blue-labelled cells indicate Ki67 negative cells/areas, and orange/red-labelled cells indicate Ki67 positive cells/areas. Platform-1 additionally allows for cytoplasmic expansion around the nucleus, thus allowing the investigator to evaluate cytoplasmic-stained cells.
A Digitisation of Ki67-stained IHC raw images using three software packages. PDEs were stained for Ki67 by IHC and visualised with DAB. The slides were then imaged using a Hamamatsu Nanozoomer XR and analysed using three digital pathology image analysis platforms (QuPath, ImmunoRatio and VisioPharm; Platforms 1–3 respectively). For all platforms, red/orange indicates DAB positivity and blue indicates DAB negative cells. B Pearson’s correlation coefficient comparing Platform-1 and Platform-3 values for total cells counted across 56 NSCLC PDEs. C Bland–Altman plot comparing total cell counts as determined by Platform-1 and Platform-3. The blue line indicates the mean difference between total counts, the green and red lines illustrate the upper and lower limits of agreement based on 95% CIs, respectively. D Passing–Bablok regression model illustrating a proportional difference in total cell counts between Platform-1 and Platform-3. E Pearson’s correlation coefficient comparing Platform-3 and Platform-1 for Ki67+ve and cPARP+ve cell counts across 56 PDEs. F Bland–Altman plot comparing Platform-3 and Platform-1 for Ki67+ve/cPARP+ve cell counts. The blue line indicates the mean difference between positive counts while the green and red lines illustrate the upper and lower limit of agreement based on 95% CIs, respectively. G Passing–Bablok regression model illustrates a proportional difference in positive cell counts between Platform-3 and Platform-1.
As Platform-1 and Platform-3 are able to count individual cells, whereas Platform-2 cannot, we first assessed inter-method concordance between Platforms 1 and 3. In terms of total cells counted, there was a significant linear relationship in the number of cells between Platforms (Fig. 2B). However, Platform-3 consistently counted more cells than Platform-1 (Fig. 2C). The increased cell counts in Platform-3 were proportionally different, but not constantly different (Fig. 2D). We then compared Platform-1 and Platform-3 with regard to Ki67 and cPARP staining. There was a significant linear relationship between Ki67+ve and cPARP+ve quantitation using both Platforms (Fig. 2E). However, as with total cells counted, Platform-3 consistently counted more positive cells (Fig. 2F) and Platform-3 counted proportionally more positive cells than Platform-1 (Fig. 2G). The proportionally different counts between Platform-3 and Platform-1 resulted in ~39% more cells counted in Platform-3 and, as the cell density increased, these differences became substantially larger (>10,000 cells).
Further to this, we compared DAB positive cells/mm2 identified by Platform-1 and Platform-3. There was a significant linear relationship in positive counts using both platforms (Supplementary Fig. 1Ai). When expressed as positive cells/mm2, Platform-3 counted more positive cells than Platform-1 (Supplementary Fig. 1Aii). The increase in cell density per mm2 observed with Platform-3 was statistically proportionally different to Platform-1, and this method of analyses also consistently counted ~39% more cells using Platform-3 (Supplementary Fig. 1Aiii). To determine which Platform detected positive cells closest to the ʻground truth’, eight FoVs were selected and positive cells were identified by Platform-1 and Platform-3 and compared to cells manually counted by a histomorphometrist (Supplementary Fig. 1B). Both platforms detected more positive cells than the histomorphometrist, with Platform-1 performing closer to the ʻground truth’.
Because the proportional difference of ~39% was observed for both total cells counted and number of Ki67+ve/cPARP+ve cells with Platforms 1 and 3, we hypothesised that converting the cell counts into % positivity would normalise the comparable values. We therefore asked a trained histomorphometrist to score the slides for % positive Ki67 and % positive cPARP in vehicle and cisplatin-treated PDEs and compared the opinions to data generated from all three Platforms. There was no statistically significant difference (Wilcoxon-paired test) between the histomorphometrist score and any Platform (Fig. 3A). We fitted PB regression to all % positive normalised data from all platforms, stains and treatment combinations to generate more stringent statistical data and found that, despite the observed statistically significant differences with PB regression (Fig. 2B–G), differences between Platforms were not large (Fig. 3B). For example, the range of constant difference between all comparisons for cPARP+ve cells in vehicle-treated PDEs is −0.63% to 0.69%.
A Comparison of % Ki67+ve and cPARP+ve staining across platforms and, in comparison, to histomorphometrist (HM) scoring. Vehicle and cisplatin-treated NSCLC PDEs were evaluated using the different approaches. Wilcoxon-paired test shows that there was no significant difference (ns) in mean positivity identified between the histomorphometrist and any software platform. B Passing–Bablok regression analysis. PB regression was fitted to all combinations of data obtained from Platforms 1–3 and from the histomorphometrist (HM) to test for inter-platform agreeability. Green highlights a constant difference between methods. Yellow highlights fields indicating a proportional difference between methods.
Overall, these data highlight that there are excellent inter-platform and histomorphometrist agreements when quantitating both Ki67 and cPARP staining, thus encouraging the use of the software platforms for evaluating PDEs at endpoint. Although statistical significance was reached, only small differences in values were observed and are unlikely to be of clinical significance. In fact, these differences are to be expected as, for example, a recent study [19] evaluated inter-observer reproducibility between 14 Pathologists for scoring Ki67 positivity in malignant breast tumours. They found median values ranged from 20 to 40%, highlighting far larger differences than those observed here. Indeed, our previous experience with NSCLC PDEs demonstrated that a difference of 28.45% in cPARP induction in cisplatin resistant/sensitive PDEs was necessary to distinguish clinical outcomes [2].
Evaluation of virtual double staining
Multiplexing of IHC generally involves staining of sequential sections with different biomarkers and then comparison of images. A known inherent problem with the use of sequential sections is the change in ʻtissue geography’ between sections leading to errors in cell and tissue classification. However, with digital pathology software packages, the option to integrate images generated from staining of sequential sections by computational alignment using ʻVirtual Double Staining’ (VDS) becomes possible and some of these inherent problems, such as section stretching, can be partially mitigated. Here we have evaluated the accuracy of VDS.
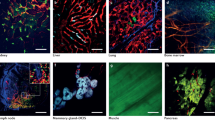
We first investigated the extent to which VDS identifies CK-stained tumour areas. Figure 4A shows raw images for CK staining of tumour, lung airways and parenchymal regions within a NSCLC PDE (Fig. 1A) generated by VisioPharm software. As can be seen, the VDS images resulted in the false positive identification of both normal lung parenchyma and lung airways within the NSCLC PDEs (Fig. 4A). To account for inherent alignment artefacts, the CK tumour mask was expanded to ensure that, when overlaid onto neighbouring slides, the tumour islands were completely covered with the tumour mask. This step, however, resulted in over-estimation of tumour areas (Fig. 4A).
A VDS alignment of CK-stained regions in NSCLC PDEs. PDEs were stained with CK by IHC. Raw images of tumour, parenchymal and airway regions are shown as well as the corresponding VDS alignments using Platform-3. CK-stained cells/regions are highlighted in blue and stroma regions/cells highlighted in green. B Pearson’s correlation of VDS with histomorphometrist scoring of CK-positive areas within Ki67 or cPARP-stained sections. C Pearson’s correlation of VDS with histomorphometrist scoring of CK-positive areas within sections generated from vehicle or cisplatin-treated PDEs. D Pearson’s correlation of total cPARP+ve cells in tumour and stroma regions after VDS or manual segmentation of CK-positive areas. The dotted lines represent x = y + 0 (identity line). E Pearson’s correlation of total Ki67+ve cells in tumour and stroma regions after VDS or manual segmentation of CK-positive areas. The dotted lines represent x = y + 0 (identity line). F % positivity for Ki67+ve and cPARP+ve staining in tumour and stroma areas of vehicle and cisplatin-treated PDEs using either Platform-3 for VDS identification of tumour area or manual judgement. Statistical comparisons were undertaken using the Wilcoxon-paired test.
The area of the tumour regions identified by VisioPharm by VDS was then directly compared to areas circumscribed by a histomorphometrist. In Ki67- and cPARP-stained slides, there was a significant linear relationship between manually identified tumour areas and VDS (Fig. 4B). Additionally, when areas were assessed based on PDE treatment type, there was also a significant correlation between VDS and manual tumour area detection. However, VDS identified more CK-positive areas in cisplatin-treated PDEs compared to vehicle-treated PDEs (Fig. 4C). This was due to VisioPharm identifying extensive areas of small, diffuse CK-positive apoptotic bodies/regions in cisplatin-treated PDEs, some of which were excluded by the histomorphometrist when manually annotating tumour areas on the whole slides.
As the computationally detected tumour areas were consistently larger than areas circumscribed by the histomorphometrist due to the tumour mask expansion, Ki67+ve and cPARP+ve cells were counted in both the tumour and stroma tissue regions identified by each method. Despite the large differences in identified tumour area (Fig. 4B), there was a significant correlation between tissue segmentation methods when counting cells, with them lying close to the identity line (Fig. 4D, E).
We also compared the values obtained for % Ki67+ve and % cPARP+ve staining in tumour and stromal areas using VDS and manual alignment (Fig. 4F). There was no significant difference in % cPARP+ve cells in the tumour between the two methods (Fig. 4F, upper left panel). However, in the stroma, significantly more cPARP+ve cells were identified by VDS alignment (Fig. 4F upper right panel). % Ki67+ve was significantly higher in the tumour areas identified by manual methods (Fig. 4F, lower left panel) and was mirrored by significantly lower Ki67 positivity in the stroma identified by manual methods (Fig. 4F, lower right panel). These differences are likely attributable to the over-estimation of CK-positive areas using VDS (Fig. 4A–D) and highlight a significant limitation with VDS that needs to be considered when using this approach to quantitate PDE responses to anti-cancer drugs at endpoint.
Comparison of mIF and IHC methods for endpoint PDE analysis
Due to the limitations described above with VDS for sequentially stained IHC images, we transitioned to using mIF to visualise all three biomarkers (Ki67, CK, cPARP) simultaneously on a single tissue section. Unlike VDS, an expansion constant was not required, as CK-positive tumour areas can be seen using the ‘tumour mask’ panel (Fig. 5A). Identification of Ki67 and cPARP positive cells can then be achieved by thresholding on nuclear fluorescence signal intensity, allowing the visualisation of both apoptosing and proliferating cells on the same slide (Fig. 5A).
A Representative example of mIF-stained PDEs for Ki67 (green), cPARP (red), cytokeratin (yellow) and DAPI (blue). Individual stains and merged images are shown. A tumour mask (yellow) and stromal tissue (blue) were identified following cellular segmentation, allowing positive stain identification on a single slide. A lower power view of mIF-stained explants on a single slide is shown in the upper left panel. B Pearson’s correlation of the total cells identified in tumour and stroma in mIF slides versus virtual double staining (IHC). C Pearson’s correlation of cPARP+ve cells in the tumour and stroma in mIF sections versus virtual double staining (IHC). D Pearson’s correlation of Ki67+ve cells identified in tumour and stroma in mIF sections versus virtual double staining (IHC). E % Ki67+ve and % cPARP+ve cells identified in stroma and tumour regions when analysed by mIF or by IHC using the VDS approach. The Wilcoxon-paired test was used to test for significance. Each point represents a single PDE.
We compared total cell counts in PDEs stained with mIF to the VDS generated from IHC-stained sections (Fig. 4) and found that there was a significant linear relationship between the methods in both tumour and stromal areas (Fig. 5B). There was also significant concordance between the two methods in the number of cPARP+ve cells in tumour and stromal regions (Fig. 5C). However, there was no significant linear relationship in the number of Ki67+ve cells in the tumour regions of the mIF and IHC sections. This is likely attributable to the physical distance between stained sections in the case of the IHC staining i.e. >12 μm (Fig. 1A).
When cPARP+ve cells were expressed as % positivity, there was significantly more apoptotic cell death in both the tumour and stroma tissue regions using the IHC VDS approach, compared to the mIF approach (Fig. 5E, upper panels). There were also significantly less % Ki67+ve cells in the stroma by mIF when compared to DAB slides (Fig. 5E, lower right panel). These differences are likely the result of spatial drift between sections using the IHC VDS approach.
Discussion
In this study we have compared different approaches for evaluating changes in biomarker expression in response to anti-cancer drugs in PDEs. PDEs do not involve tumour deconstruction and therefore can be utilised to study drug responses in both tumour and stroma, and within different regions of the same tumour using spatial profiling [1, 3]. They therefore offer many advantages over other preclinical models for monitoring responses to agents that target the tumour microenvironment such as immunotherapies as well as for examining the role of tumour heterogeneity in influencing drug response.
To take full advantage of PDEs as preclinical models, immunostaining approaches for spatial profiling need to be optimised. Previous approaches have utilised the laborious staining of sequential sections generated from FFPE-processed PDEs using IHC and manual alignment of the resulting staining patterns [2]. Here, we have introduced the use of digital pathology solutions as well as mIF for generating faster and more reproducible data.
We first compared results for the quantitation of IHC staining for Ki67 as a proliferation marker and cPARP as an apoptosis marker using three available computer-assisted digital pathology software packages (QuPath, Immunoratio and VisioPharm). When total cell counts were compared between QuPath and VisioPharm, we found that there were some differences between platforms. All three platforms generated highly comparable data that was not significantly different to the views of a histomorphometrist when expressed as % positivity (Fig. 3A). There was a range of significant differences between platforms, when tested against BA and PB statistical methods (Fig. 3B). However, although statistically significant, the differences were in fact small, highlighting the high degree of concordance between platforms. Our data show that, when comparing across software platforms, it is best to evaluate data as % positive cells and not direct cell counts, as this resulted in all three software packages generating data close to that of the histomorphometrist i.e. the ʻground truth’. For example, in untreated PDEs, the low level of difference in Ki67 positivity between the digital platforms and a histomorphometrist ranged from 3.09 to 4.09%, which is much lower than the range identified by 14 pathologists for Ki67 positivity in breast cancer, where the median positivity ranged from 20 to 40% [19]. There was also strong concordance between digital platforms when evaluating cPARP in PDEs, although the histomorphometrist consistently scored for higher levels of cell death (5.02–6.55% higher) in cisplatin-treated PDEs than the digital pathology solutions. The explanation for this may be that the observer included areas of late-apoptosis in the cell death score, where the cell nuclei had broken down to apoptotic bodies, and areas of necrosis, whereas the software packages were optimised to not segment small apoptotic bodies as this would over count apoptotic cells. However, overall, there is a strong level of consistency in the approaches and therefore we conclude that any of the three packages are suitable for quantitating Ki67 and cPARP expression changes in PDEs. It should also be borne in mind that the need for user input at the algorithm tuning stage may mean that results could vary between users as well as the version of the software packages used.
A critical advantage of spatial profiling with PDEs is the ability to separate drug responses in tumour and stromal areas. We therefore next assessed the robustness of digital approaches for doing this based on pan-cytokeratin staining for tumour areas. VDS, using digital software packages, allows for the separation of tumour and stromal regions in IHC-stained sequential sections, whereby tumour areas identified by a pan-cytokeratin antibody are overlaid onto the sequentially stained slides [20]. We have found that there are some noteworthy caveats to this approach, which do not fully remove the requirement for histomorphometrist analysis. Specifically, as expected [21], we found that pan-cytokeratin staining in NSCLC PDEs also stains normal parenchyma and airway. In an automated system, this staining results in over-identification of tumour areas (Fig. 4A) and, therefore, manual curation and/or additional tumour-specific markers are needed for full accuracy. In addition, there was imperfect section alignment that could only be overcome by including an expansion constant so that tumour areas on neighbouring slides were over-estimated. Consistent with our previous studies of NSCLC PDEs treated with cisplatin, we observed cPARP staining in some non-tumour regions [2], indicating cell death induction in stromal and immune cell compartments.
To overcome the aforementioned issues with VDS we transitioned to mIF, which allowed for the monitoring of multiple biomarkers simultaneously on a single slide. This resulted in significantly fewer cPARP positive cells being identified in the tumour and stroma, and fewer Ki67 positive cells in the stroma (Fig. 5E). There are a multitude of technical explanations as to why concordance between VDS and mIF was not achieved in our study. For example, a significant benefit of mIF over VDS is that fluorescent signals can be viewed separately, so it is easy to see signal co-localisation with a fluorescent counter stain. This is of particular use when weak signals from primary antibodies can be observed in isolation without interference. However, if fluorochromes are in close proximity, resonance energy transfer can also take place, quenching the signal of one of the fluorochromes. Despite the ability to visualise weak signals in isolation when using mIF, these are often masked by natural tissue auto-fluorescence, which can be significantly exacerbated by formaldehyde [22]. In the absence of multispectral imaging systems, this auto-fluorescent masking of weak positive signal can translate into lower positive identities as a result of having to set positive cell thresholding above background/weak positives [23]. A poorly optimised mIF experiment, therefore, has the potential to generate a lower signal-to-noise ratio compared to chromogenic IHC. Lower signal-to-noise ratios can also occur as a result of photo-bleaching of the reporter and the specific properties of instrument-specific excitation/emission filters [24]. Additionally, fluorescent imaging provides higher resolution of antigen localisation compared to chromogenic substrate deposits in IHC [25].
In summary, we have shown that digital pathology software packages facilitate spatial profiling of PDEs at endpoint, particularly when mIF is implemented. Thus, they provide a significant boost for the automation and increased throughput of the PDE-drug discovery pipeline.
References
Powley IR, Patel M, Miles G, Pringle H, Howells L, Thomas A, et al. Patient-derived explants (PDEs) as a powerful preclinical platform for anti-cancer drug and biomarker discovery. Br J Cancer. 2020;122:735–44.
Karekla E, Liao WJ, Sharp B, Pugh J, Reid H, Quesne JL, et al. Ex vivo explant cultures of non-small cell lung carcinoma enable evaluation of primary tumor responses to anticancer therapy. Cancer Res. 2017;77:2029–39.
Collins A, Miles GJ, Wood J, MacFarlane M, Pritchard C, Moss E. Patient-derived explants, xenografts and organoids: 3-dimensional patient-relevant pre-clinical models in endometrial cancer. Gynecol Oncol. 2020;156:251–9.
Majumder B, Baraneedharan U, Thiyagarajan S, Radhakrishnan P, Narasimhan H, Dhandapani M, et al. Predicting clinical response to anticancer drugs using an ex vivo platform that captures tumour heterogeneity. Nat Commun. 2015;6:6169.
Furukawa T, Kubota T, Hoffman RM. Clinical applications of the histoculture drug response assay. Clin Cancer Res. 1995;1:305–11.
Pirnia F, Frese S, Gloor B, Hotz MA, Luethi A, Gugger M, et al. Ex vivo assessment of chemotherapy-induced apoptosis and associated molecular changes in patient tumor samples. Anticancer Res. 2006;26:1765–72.
Garcia-Chagollan M, Carranza-Torres IE, Carranza-Rosales P, Guzman-Delgado NE, Ramirez-Montoya H, Martinez-Silva MG, et al. Expression of NK cell surface receptors in breast cancer tissue as predictors of resistance to antineoplastic treatment. Technol Cancer Res Treat. 2018;17:1533033818764499.
Wei B, Wang J, Zhang X, Qian Z, Wu J, Sun Y, et al. Combination of histoculture drug response assay and qPCR as an effective method to screen biomarkers for personalized chemotherapy in esophageal cancer. Oncol Lett. 2017;14:6915–22.
Stack EC, Wang C, Roman KA, Hoyt CC. Multiplexed immunohistochemistry, imaging, and quantitation: a review, with an assessment of Tyramide signal amplification, multispectral imaging and multiplex analysis. Methods. 2014;70:46–58.
Hofman P, Badoual C, Henderson F, Berland L, Hamila M, Long-Mira E, et al. Multiplexed immunohistochemistry for molecular and immune profiling in lung cancer-just about ready for prime-time? Cancers. 2019;11. https://doi.org/10.3390/cancers11030283.
Stalhammar G, Fuentes Martinez N, Lippert M, Tobin NP, Molholm I, Kis L, et al. Digital image analysis outperforms manual biomarker assessment in breast cancer. Mod Pathol. 2016;29:318–29.
Tuominen VJ, Ruotoistenmaki S, Viitanen A, Jumppanen M, Isola J. ImmunoRatio: a publicly available web application for quantitative image analysis of estrogen receptor (ER), progesterone receptor (PR), and Ki-67. Breast Cancer Res. 2010;12:R56.
Bankhead P, Loughrey MB, Fernandez JA, Dombrowski Y, McArt DG, Dunne PD, et al. QuPath: open source software for digital pathology image analysis. Sci Rep. 2017;7:16878.
Tang LH, Gonen M, Hedvat C, Modlin IM, Klimstra DS. Objective quantification of the Ki67 proliferative index in neuroendocrine tumors of the gastroenteropancreatic system: a comparison of digital image analysis with manual methods. Am J Surg Pathol. 2012;36:1761–70.
Bland JM, Altman DG. Measuring agreement in method comparison studies. Stat Methods Med Res. 1999;8:135–60.
Passing H, Bablok W. A new biometrical procedure for testing the equality of measurements from two different analytical methods. Application of linear regression procedures for method comparison studies in clinical chemistry, Part I. J Clin Chem Clin Biochem. 1983;21:709–20.
Tsujikawa T, Kumar S, Borkar RN, Azimi V, Thibault G, Chang YH, et al. Quantitative multiplex immunohistochemistry reveals myeloid-inflamed tumor-immune complexity associated with poor prognosis. Cell Rep. 2017;19:203–17.
Remark R, Merghoub T, Grabe N, Litjens G, Damotte D, Wolchok JD, et al. In-depth tissue profiling using multiplexed immunohistochemical consecutive staining on single slide. Sci Immunol. 2016;1:aaf6925.
Laenkholm AV, Grabau D, Moller Talman ML, Balslev E, Bak Jylling AM, Tabor TP, et al. An inter-observer Ki67 reproducibility study applying two different assessment methods: on behalf of the Danish Scientific Committee of Pathology, Danish breast cancer cooperative group (DBCG). Acta Oncol. 2018;57:83–9.
Roge R, Riber-Hansen R, Nielsen S, Vyberg M. Proliferation assessment in breast carcinomas using digital image analysis based on virtual Ki67/cytokeratin double staining. Breast Cancer Res Treat. 2016;158:11–9.
Blobel GA, Moll R, Franke WW, Vogt-Moykopf I. Cytokeratins in normal lung and lung carcinomas. Virchows Archiv B. 1984;45:407–29.
Baschong W, Suetterlin R, Laeng RH. Control of autofluorescence of archival formaldehyde-fixed, paraffin-embedded tissue in confocal laser scanning microscopy (CLSM). J Histochem Cytochem. 2001;49:1565–72.
Pivetta E, Spessotto P. Multispectral imaging technology: Visualize, analyze, phenotyping, and quantify immune cells in situ. Int J Biol Markers. 2020;35 1_Suppl:26–30.
Lin JR, Fallahi-Sichani M, Chen JY, Sorger PK. Cyclic Immunofluorescence (CycIF), a highly multiplexed method for single-cell imaging. Curr Protoc Chem Biol. 2016;8:251–64.
Robertson D, Savage K, Reis-Filho JS, Isacke CM. Multiple immunofluorescence labelling of formalin-fixed paraffin-embedded (FFPE) tissue. BMC Cell Biol. 2008;9:13.
Acknowledgements
We thank Kees Straatman and the Leicester Core Biotechnology Services Imaging Facility for support with image analysis. We also thank Angie Gilles, Linda Potter and Janine Morton of the Leicester Core Biotechnology Services Histology Facility. We also thank the MRC Toxicology Unit Histopathology Facility for support with digital scanning and image analysis. This research was supported and funded by the Explant Consortium comprising four partners: The University of Leicester, The MRC Toxicology Unit, Cancer Research UK Therapeutic Discovery Laboratories and LifeArc. Additional support was provided by the CRUK-NIHR Leicester Experimental Cancer Medicine Centre (C10604/A25151). Funding for GM was provided by Breast Cancer Now’s Catalyst Programme (2017NOVPCC1066), which is supported by funding from Pfizer.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Miles, G.J., Powley, I., Mohammed, S. et al. Evaluating and comparing immunostaining and computational methods for spatial profiling of drug response in patient-derived explants. Lab Invest 101, 396–407 (2021). https://doi.org/10.1038/s41374-020-00511-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-020-00511-3