Abstract
As part of routine histological grading, for every invasive breast cancer the mitotic count is assessed by counting mitoses in the (visually selected) region with the highest proliferative activity. Because this procedure is prone to subjectivity, the present study compares visual mitotic counting with deep learning based automated mitotic counting and fully automated hotspot selection. Two cohorts were used in this study. Cohort A comprised 90 prospectively included tumors which were selected based on the mitotic frequency scores given during routine glass slide diagnostics. This pathologist additionally assessed the mitotic count in these tumors in whole slide images (WSI) within a preselected hotspot. A second observer performed the same procedures on this cohort. The preselected hotspot was generated by a convolutional neural network (CNN) trained to detect all mitotic figures in digitized hematoxylin and eosin (H&E) sections. The second cohort comprised a multicenter, retrospective TNBC cohort (n = 298), of which the mitotic count was assessed by three independent observers on glass slides. The same CNN was applied on this cohort and the absolute number of mitotic figures in the hotspot was compared to the averaged mitotic count of the observers. Baseline interobserver agreement for glass slide assessment in cohort A was good (kappa 0.689; 95% CI 0.580–0.799). Using the CNN generated hotspot in WSI, the agreement score increased to 0.814 (95% CI 0.719–0.909). Automated counting by the CNN in comparison with observers counting in the predefined hotspot region yielded an average kappa of 0.724. We conclude that manual mitotic counting is not affected by assessment modality (glass slides, WSI) and that counting mitotic figures in WSI is feasible. Using a predefined hotspot area considerably improves reproducibility. Also, fully automated assessment of mitotic score appears to be feasible without introducing additional bias or variability.
Similar content being viewed by others
Introduction
Breast cancer is among the most prevalent cancer types in the world. It is estimated that the incidence of invasive breast cancer in the United Stated of America will increase by more than 50% in the year 2030 compared to the year 2011 [1]. One of the most powerful prognostic tools for early stage invasive breast cancer is histological grading by means of the Nottingham grading system [2,3,4]. Grading is therefore part of the routine pathological workup of every newly diagnosed invasive breast cancer. The histological grade of invasive breast cancer is determined by microscopical assessment of three distinct tumor features: the extent of tubule formation, nuclear pleomorphism, and the mitotic density. Mitotic density, which has been shown to provide the strongest prognostic value of these three elements [5], is assessed in two steps. First, the pathologist examines the slides containing tumor to visually select the region with the highest proliferative activity, which is mostly found at the tumor periphery. Next, within this region all mitotic figures are counted in an area of 2 mm2 (consisting of 10 consecutive high-power fields (HPF). Both the area selection and the evaluation of candidate mitotic figures have been shown to be prone to interobserver and intraobserver variability [6, 7].
Recent advances in machine learning, combined with the possibility to create digital scans of tissue sections (so-called whole slide images; WSI) have paved the way for fully automated analysis of histological slides, even approaching the accuracy of pathologists for certain well-defined tasks [8]. Current state of the art machine learning algorithms consist of deep convolutional neural networks (CNN). These models consist of interconnected artificial neurons that exchange information in order to solve a particular vision task. They are trained to recognize visual patterns using labeled data in order to classify whole images or detect objects within images. In a previous study, we trained a CNN to detect individual mitotic figures in breast cancer WSI with high accuracy [9]. However, translating these results into real benefit for routine clinical practice requires careful evaluation and comparison with existing methods.
Previous research has mostly focused on the accuracy of deep learning methods in detecting individual mitotic figures. The current study aimed to validate automated assessment of mitotic density in WSI, including detection of mitotic figures as well as objective hotspot determination. Automatically assessed mitotic densities were compared with data from pathologists, both using the absolute number of mitotic figures as well as translating the mitotic count into a mitotic score as defined by the Nottingham grading system.
Materials and methods
Study design
In this study, we used a cohort of prospectively included breast cancer cases (Cohort A) and a previously described TNBC cohort [10, 11]. All H&E tissue slides were digitally scanned, producing WSI on which our previously described automated mitosis detection algorithm was applied [9].
For cohort A, the mitotic count on glass slides was assessed by a pathologist specialized in breast cancer (PB) as part of routine diagnostics and additionally by a pathology resident (MCAB). Also, both observers performed mitotic counting for the same cases in an automatically generated 2 mm2 hotspot area on WSI. The mitotic counts on glass slides of the TNBC cohort were obtained as part of a central histopathological revision, after which two additional pathologists independently assessed the mitotic count.
Patient and tissue selection
Cohort A
From November 2015 until April 2017, all primary operated breast cancers which were pathologically examined in the Radboud University Medical Center (Radboudumc, Nijmegen, the Netherlands) were prospectively included (n = 221). Of this cohort, 90 cases were randomly selected while stratifying for the mitotic frequency score given during routine clinical practice by a specialized breast pathologist (PB) on glass slides: 30 cases for mitotic score 1 (≤7 mitotic figures per 2 mm2), 29 cases for mitotic score 2 (8–12 mitotic figures per 2 mm2) and 31 cases for mitotic score 3 (≥13 mitotic figures per 2 mm2). Patient and tumor characteristics were retrieved from the pathology reports (Table 1), but were not used as selection criteria. All breast tumor characteristics were routinely classified by the prevailing guidelines for breast cancer reporting in the Netherlands [12]. A pathology resident (MCAB) independently assessed the mitotic count for these 90 cases on glass slides, blinded for the initial mitotic score. Both observers reported the absolute number of their mitotic count as well as the resulting mitotic frequency scores [2].
Cohort B, TNBC cohort
Cases were included from a previously described multicenter, retrospective TNBC cohort [10]. In this study, three independent observers (MCAB: pathology resident; WV and PCC: pathologists with special interest in breast cancer) assessed the mitotic count for every tumor by conventional microscopic glass slide assessment according to the Nottingham grading system [2,3,4] as outlined under Cohort A. For the experiments in this study, only the mitotic count averaged over the three observers was used (n = 298). Patient and tumor characteristics of the TNBC cohort are summarized in Table 1.
Automatic mitotic counting
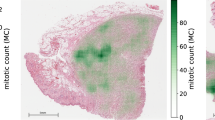
H&E tissue sections of cohort A and cohort B were scanned using a Pannoramic 250 Flash II slide scanner (3DHistech, Hungary) using a 40X objective (numerical aperture: 0.95; resulting in a specimen level pixel size of 0.12 × 0.12 µm). A previously described CNN for mitosis detection in H&E was applied to all WSI [9]. To allow comparison with routine manual mitotic counting, the algorithm was expanded to automatically determine a 2 mm2 circular area in the WSI having the highest density of mitotic figures. Apart from being restricted to a circular shape, this resembles clinical practice [2] (Fig. 1a, b). A mitotic figure was counted if its center pixel (being defined as the pixel with the highest detection probability in the cell neighborhood) was contained within the circle boundary. Because this CNN was not trained to discriminate mitotic figures in different tissue types (benign versus malignant), a number of cases with a very low mitotic count showed a hotspot outside the tumor. For these cases, the tumor was manually delineated and the CNN was applied again, now forced to select the hotspot in the delineated invasive tumor area. The absolute number of automatically detected mitotic figures was reported and additionally translated into the mitotic frequency scores.
Example of results of the CNN applied on the digitized H&E section of one of the tumors in cohort A. a Overview on low magnification of the CNN output. Every detected mitotic figure is pointed out with a green dot. The yellow circle indicates the 2 mm2 area with the highest number of mitotic figures. b Hotspot area as found by the CNN on higher magnification with the detected mitotic figures by the CNN labeled with green circles. c Identical hotspot area as in B in which the mitotic figures which were detected by observer 1 are labeled with red circles. Green arrow: mitotic figure detected by observer 1 and not labeled by the CNN. Green arrow head: mitotic figure detected by the CNN and not labeled by observer 1. d Identical hotspot area as in B in which the mitotic figures which were detected by observer 2 are labeled with blue circles. Green arrows: mitotic figures detected by observer 2 and not labeled by the CNN. Green arrow heads: mitotic figures detected by the CNN and not labeled by observer 2
Manual mitotic counting in CNN generated hotspot area in cohort A
To study the effect of hotspot selection on the mitotic count, the CNN generated hotspot was visually marked in all 90 WSI of cohort A. The two observers who assessed the mitotic count on glass slides (PB, MCAB) independently assessed the mitotic count in the designated hotspot by clicking each mitotic figure in the predefined circle on a computer screen using in-house built software [13] (Figs. 1c, d). If a mitotic figure was situated in the border of the circle, it was counted when minimal half of the mitotic figure was contained within the circle boundary, regardless of the rest of the cell body. The washout period between glass assessment and digital assessment ranged from two months to more than 1 year.
Ethical approval
The requirement for ethical approval was waived by the institutional review board (cohort A: case number 2015-1637; cohort B: case number 2015-1711) of the Radboudumc. All patient material and data were treated according to the code of conduct for the use of data in health research [14] and the code of conduct for dealing responsibly with human tissue in the context of health research [15].
Statistical analysis
To test the agreement of the mitotic scores, linearly weighted kappa scores were calculated. To assess the observer agreement for the absolute numbers of detected mitotic figures, the intraclass correlation coefficient (ICC) was used. A 2-way random-effects model tested for absolute agreement with reliability calculated from a single measure (corresponding to ICC [1, 2] according to the Shrout and Fleiss convention[16]) was used. The widely applied guidelines by Cicchetti [17] were used for the interpretation of the kappa and ICC values (>0.40: poor; 0.40–0.59: fair; 0.60–0.74: good; >0.75: excellent). To visualize observer variability for the mitotic counts, Bland-Altman plots were generated with corresponding 95% limits of agreement (LA) which are equivalent to a distance of 1.96 SD of the mean of the observations [18]. Outliers were defined as detections lying outside these 95% limits of agreement. In addition, mean absolute differences were calculated.
Per observer, the repeated measures ANOVA analysis was performed to test if the mean difference between glass assessment and assessment with predefined hotspot selection was statistically different.
Because triple negative breast cancers display wide ranges of mitotic counts, cohort B was additionally used to study the relationship between the absolute numbers of detected mitotic figures by observers and the CNN. A scatterplot was made to visualize the relation between the averaged and automatic mitotic counts in cohort B. Linear regression analysis without a constant was used to calculate the equation which described the relationship between manual and automatic mitotic counts most accurately.
For all analyses, confidence intervals were set at the 95% level and a minimal p value of <0.05 was considered statistically significant. All analyses were performed using statistical software SPSS (version 25.0; IBM, Chicago, USA) and R (version 3.5.1).
Results
Automatic mitotic counting
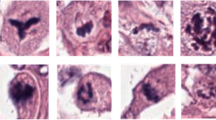
All CNN detected mitotic figures in cohort A and B were visualized on the WSI with the corresponding 2 mm2 circle in which the highest density of mitotic figures was detected (Figs. 1a, b, cohort A). For 52 of the 90 cases in cohort A, a tumor annotation was made to force the CNN to count within the tumor area. Figure 2a shows an example of correctly identified mitotic figures by the CNN in a tumor from cohort A. The CNN was able to handle tumors with extremely aberrant nuclear morphology (Fig. 2b, cohort B). The CNN was not completely robust against several phenomena (Fig. 2c: ink; Fig. 2d: lymphocytes and fibroblasts) which in some cases resulted in false positive detections.
Examples of true positive and false positive detections of the convolutional neural network (CNN). a Area with correctly detected mitotic figures, tumor from cohort A. b Area with detected mitotic figures in a high grade tumor with severely pleomorphic nuclear morphology, tumor from cohort B. c Area with false positive detections (ink), tumor from cohort A. d Area with false positive detections (left: lymphocyte; right: fibroblast), tumor from cohort A
Baseline agreement for mitotic counting without predefined hotspot selection using glass slides (cohort A)
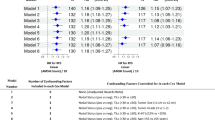
The upper section of Table 2 shows cross comparisons for the mitotic scores of the glass assessments of observer 1 versus observer 2 for the 90 tumors in cohort A. Using this assessment method, the two observers never differed more than 1 class from each other. The linear weighted kappa statistic of the mitotic score for observer 1 and 2 using conventional glass slide assessment was 0.689 (95% CI 0.580–0.799; p < 0.001). For the agreement on absolute numbers of detected mitotic figures in cohort A on glass slides, the intraclass coefficient (ICC) was 0.835 (95% CI 0.755–0.890; p < 0.001) between both observers. Figure 3a shows the Bland Altman plot of observer 1 and 2 with the corresponding 95% limits of agreement for mitotic counting on glass slides.
Bland Altman plots with mean difference (yellow dashed line) and corresponding 95% limits of agreements (LA; blue dashed lines) for comparison of mitotic counts between observer 1 and 2 in cohort A. Detections lying outside the upper and lower LA are colored red. For visualization purposes, for the x-axis a log scale was used. a Observer 1 versus observer 2; no predefined hotspot selection; conventional glass slide assessment. b Observer 1 versus observer 2; convolutional neural network (CNN) defined hotspot area; whole slide image (WSI) assessment
Interobserver variability for mitotic counting in the CNN generated hotspot area (cohort A)
For counting in a predefined region, the kappa statistic for agreement between observer 1 and 2 was 0.814 (95% CI 0.719–0.909; p < 0. 001). The lower section of Table 2 shows the corresponding cross comparisons for the mitotic scores between observer 1 and 2. Figure 3b shows the Bland Altman plot for the absolute mitotic counts of observer 1 and 2 in the CNN generated hotspot region. The mean absolute difference between observers for conventional glass slide assessment was 4.8 (SD 5.4) and for counting in the predefined hotspot 4.4 (SD 6.6).
Impact of hotspot selection on the mitotic count (cohort A)
For both observers, the intraobserver variation for detection of mitotic figures with and without the predefined counting area was calculated. Table 3 shows cross comparisons for the mitotic scores for observer 1 (upper part) and 2 (lower part). Kappa statistics on intraobserver agreement were 0.698 (95% CI 0.587–0.808) and 0.684 (95% CI 0.577–0.791) for observer 1 and 2, respectively. Intraobserver agreement on absolute numbers of detected mitotic figures are visualized in Fig. 4. Repeated measures ANOVA analysis showed no statistical difference between counting with and without predefined hotspot selection (observer 1: p = 0.468; observer 2: p = 0.969).
Bland Altman plots with mean difference (yellow dashed line) and corresponding 95% limits of agreements (LA; blue dashed lines) for comparison of mitotic counts between no predefined hotspot mitotic count assessment and assessment in a convolutional neural network (CNN) predefined area in cohort A. Detections lying outside the upper and lower LA are colored red. For visualization purposes, for the x-axis a log scale was used. a Observer 1; no predefined hotspot selection; conventional glass slide assessment versus CNN defined hotspot area; whole slide image (WSI) assessment. b Observer 2; no predefined hotspot selection; conventional glass slide assessment versus CNN defined hotspot area; WSI assessment
Automated versus manual mitotic counting (cohort A)
The results on agreement between the mitotic scores of the CNN and the scores of both assessment methods of observer 1 and 2 are outlined in Table 4. For glass slide assessment versus automatic mitotic counting, observer 1 yielded a kappa score of 0.604 (95% CI 0.477–0.731) and observer 2 of 0.609 (95% CI 0.484–0.734). The agreement on absolute numbers between observers using glass slide assessment and the CNN were 0.828 (95% CI 0.750–0.883; p < 0.001) for observer 1 and 0.757 (95% CI 0.638–0.839; p < 0.001) for observer 2. For counting in the predefined hotspot area, kappa scores were 0.654 (95% CI 0.530–0.777) and 0.794 (95% CI 0.691–0.896) for observer 1 and 2, respectively. The agreement on absolute numbers between observers counting in the predefined hotspot and the CNN was 0.895 (95% CI 0.845–0.930; p < 0.001) and 0.888 (95% CI 0.783–0.936; p < 0.001) for observer 1 and 2, respectively. Figure 5 visualizes the agreement between observers and automatic mitotic counting by the CNN Table 5.
Bland Altman plots with mean difference (yellow dashed line) and corresponding 95% limits of agreements (LA; blue dashed lines) for comparison of mitotic counts between observers and the convolutional neural network (CNN) in cohort A. Detections lying outside the upper and lower LA are colored red. For visualization purposes, for the x-axis a log scale was used. a Observer 1 versus CNN; CNN defined hotspot area; whole slide image (WSI) assessment. b Observer 2 versus CNN; CNN defined hotspot area; WSI assessment. c Observer 1 versus CNN; no predefined hotspot selection; conventional glass slide assessment. d Observer 2 versus CNN; no predefined hotspot selection; conventional glass slide assessment
Mathematical relationship between automatic and manual mitotic counting (cohort B, TNBC cohort)
In the selected cases of cohort B, the mean mitotic score of the averaged observer counts ranged from 1 to 187 mitoses per 2 mm2 (mean 37.6; SD 23.4) and of the CNN ranged between 1 and 269 mitotic figures per 2 mm2 (mean 57.6; SD 42.2). Linear regression analysis showed a Pearson’s correlation coefficient (r) of 0.810 (95% CI 0.762–0.855) between manual and automatic counts (Fig. 6). The line which approached the relationship between the averaged manual countings and the automatic countings most accurately was described by the equation:
Scatter plot of the mitotic counts of the averaged observers and the convolutional neural network (CNN) in cohort B. The blue line represents the linear regression function which described the relation between manual and automatic mitotic counts most accurately. For visualization purposes, for both axes a log scale was used. Therefore, the blue line does not pass the 0,0 coordinate in this graph. r: Pearson’s correlation coefficient
Mitotic counting of CNN = 1.512 * mitotic count averaged over observers
Discussion
In this study, we explored the possibility to use CNNs to (partly) automate mitotic counting for breast cancer. We studied the effect of automated hotpot selection on manual mitotic counting compared with conventional glass slide assessment. In addition, fully automatic mitotic counting by a CNN was studied. We showed that manual mitotic counting is not affected by assessment modality (glass slides, WSI). Counting mitoses in a computer-selected hotspot area considerably improved the interobserver agreement between observers. Mitotic counts independently assessed by the CNN were comparable to results of the observers.
Human visual mitotic counting is known to possess a certain degree of intra and interobserver variability. Both selection of the area to count as well as the actual counting introduce variability, leading to interobserver kappa values as low as 0.34 [19] for mitotic counting on glass slides. Preselection of the counting area was shown to increase reproducibility, having an interobserver kappa of 0.642 on glass slides [20]. In the same study they found an interobserver variability on WSI which is comparable to glass slides, having a kappa of 0.635 (ICC of 0.924) for the same cases and in the identical preselected hotspot. This ICC was somewhat higher than that found in the present study (0.852), whereas the kappa value in the present study of 0.814 is higher. The latter may be attributed to the fact that we calculated a linearly weighted kappa rather than an unweighted kappa. This shows that observer reproducibility can be enhanced by pre-selecting the area to count and also that transitioning to WSI does not impact reproducibility of mitotic counting.
After the United States Food and Drug Administration allowed the use of WSI for routine clinical practice [21], professional associations such as the College of American Pathologists and The Royal College of Pathologists (UK) have made recommendations to validate the use of WSI in the diagnostic setting [22, 23]. The use of WSI in primary diagnostics should be validated for a specific task of interest. With respect to the use of WSI for breast cancer grading, multiple comparison studies have been performed which show that using WSI is feasible for this particular task [22, 23]. The mitotic count is potentially the most challenging element of the three-tier breast cancer grading system to assess on WSI. To be able to detect mitotic figures, WSI should be of high resolution to appreciate the subtle features that identify a mitotic figure. In addition, the inability to inspect different focal planes can influence the number of detected mitotic figures when using WSI.
In a recent publication of Rakha et al [24], the histological grade scored on glass slides was reassessed by an expert pathologist on WSI after a washout period in more than 1600 retrospective breast cancer cases. Comparison between the original mitotic score on glass slides versus WSI resulted in an interobserver kappa of 0.47 which is markedly higher than the kappa of 0.34 found for interobserver glass slide mitotic score assessment [19]. In the present study, we found an average intraobserver kappa between glass slide and WSI mitotic scores of 0.691 (range: 0.684–0.698; ICC = 0.786), applying a preselected hotspot in WSI assessment. Counting in identical hotspot regions in both glass slide and WSI assessment even resulted in a kappa of 0.779 [20]. The transition from glass slides to WSI appears to introduce less variability than hotspot selection or the variability caused by having different observers.
Transitioning to fully automated assessment of mitotic density on WSI potentially can reduce both observer variability and variability caused by hotspot selection. Fully automatic generation of the mitotic counts by a CNN was compared with both conventional glass slide assessment and WSI with hotspot assessment of the mitotic count by the observers. The agreement based on the linear weighted kappa was good and the ICC values were excellent, which is in line with the results of Veta et al [7]. The agreement between the observers and the CNN improved when the predefined hotspot area was used by the observers. Our experiment with the TNBC cohort, in which mitotic figures usually are abundantly present, shows that for higher numbers of mitoses the CNN has an increasingly higher mitotic count as compared to the observers. It is conceivable that humans are less critical in their counting when the mitotic density is much higher than the highest Nottingham grading system cut off value, a phenomenon that will not hamper CNN based counting.
This study has several strengths. We used two different breast cancer cohorts to study the effect of automating mitotic count for invasive breast cancer. The TNBC cohort comprised of cases from 5 different hospitals (including both academic and general hospitals), introducing essential variation in tissue morphology which is needed to adequately compare and evaluate assessment methods for obtaining the mitotic count. To the best of our knowledge, this is the first study which performed a stepwise evaluation of mitotic count assessment from the conventional glass slide assessment method which served as baseline, to CNN assisted assessment of the mitotic count by generating a predefined hotspot area, and fully automatic mitotic counting by a CNN. Our research is limited by the selection criteria of tumors in cohort A, which were selected based on the initial mitotic count of one of the involved observers during routine clinical work up. For tumors in which sparse mitotic figures were present, the CNN defined the hotspot area on false positive detections. Therefore, for 52 of the 90 cases in cohort A, an outline of the tumor area was made to force the CNN to count in this area. In the future we aim to extend the training of the CNN to overcome some straightforward false positive detections. In addition, to make automatic mitotic counting more robust, the current CNN for mitoses detection can be combined with a CNN which is trained to detect invasive breast cancer.
In conclusion, by using a stepwise approach we studied the effect of using WSI and introducing CNN based features to improve the accuracy and reproducibility of the mitotic count in breast cancer. We showed that observer agreement was not affected by assessment modality (glass slides, WSI), which suggests that using WSI to count mitoses in breast cancer can be done reliably. Counting mitotic figures in a hotspot area which was generated by a CNN remarkably improved interobserver agreement. Comparison of fully automatic assessment of the mitotic count by a CNN with observers yielded results which were comparable with agreement results between observers.
References
Rosenberg PS, Barker KA, Anderson WF. Estrogen receptor status and the future burden of invasive and in situ breast cancers in the United States. J Natl Cancer Inst. 2015;107.
Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 1991;19:403–10.
Rakha EA, El-Sayed ME, Lee AH, Elston CW, Grainge MJ, Hodi Z, et al. Prognostic significance of Nottingham histologic grade in invasive breast carcinoma. J Clin Oncol. 2008;26:3153–8.
Rakha EA, Aleskandarany MA, Toss MS, Mongan NP, El-Sayed ME, Green AR, et al. Impact of breast cancer grade discordance on prediction of outcome. Histopathology. 2018:73:904–15.
Chang JM, McCullough AE, Dueck AC, Kosiorek MS, Ocal IT, Lidner TK, et al. Back to basics: Traditional Nottingham grade mitotic counts alone are significant in predicting survival in invasive breast carcinoma. Ann Surg Oncol. 2015;22(Suppl 3):S509–15.
Bonert M, Tate AJ. Mitotic counts in breast cancer should be standardized with a uniform sample area. Biomed Eng Online. 2017;16:28.
Veta M, van Diest PJ, Jiwa M, Al-Janabi S, Pluim JPW. Mitosis counting in breast cancer: object-level interobserver agreement and comparison to an automatic method. PLoS One. 2016;11:e0161286.
Ehteshami Bejnordi B, Veta M, van Diest PJ, van Ginneken B, Karssemeijer N, Litjens G, et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. J Am Med Assoc. 2017;318:2199–210.
Tellez D, Balkenhol M, Otte-Holler I, van de Loo R, Vogels R, Bult P, et al. Whole-slide mitosis detection in H&E breast histology using PHH3 as a reference to train distilled stain-invariant convolutional networks. Trans Med Imaging. 2018;37:28.
Balkenhol MCA, Vreuls W, Wauters CAP, Mol SJ, van der Laak JAWM, Bult P. Histological subtype is an independent prognostic feature of triple negative breast cancer.
Balkenhol MCA, Bult P, Tellez D, Vreuls W, Clahsen PC, Ciompi F, et al. Deep learning and manual assessment show that the absolute mitotic count does not contain prognostic information in triple negative breast cancer. Cell Oncol. 2019. https://doi.org/10.1007/s13402-019-00445-z.
Netherlands Comprehensive Cancer Registry. Guideline Breast Cancer. 2018. http://www.oncoline.nl. Accessed 25 May 2018.
Litjens G, Bandi P, Ehteshami Bejnordi B, Geessink O, Balkenhol M, Bult P, et al. 1399 H&E-stained sentinel lymph node sections of breast cancer patients: the CAMELYON dataset. Gigascience. 2018;7.
Foundation Federation of Dutch Medical Scientific Societies (Federa). Code of conduct for medical research. 2004. https://www.federa.org/codes-conduct. Accessed 2 May 2018.
Foundation Federation of Dutch Medical Scientific Societies (Federa). Human tissue and medical research: Code of conduct for responsible use. 2011. https://www.federa.org/codes-conduct. Accessed 2 May 2018.
Koo TK, Li MY. A guideline of selecting and reporting intraclass correlation coefficients for reliability research. J Chiropr Med. 2016;15:155–63.
Cicchetti DV. Guidelines, criteria, and rules of thumb for evaluating normed and standardized assessment instruments in psychology. Psychological Assessment. 1994;6:284–90.
Bland JM, Altman DG. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet. 1986;8476:307–10.
Rakha EA, Bennett RL, Coleman D, Pinder SE, Ellis IO. UK National Coordinating Committee for Breast Pathology (EQA Scheme Steering Committee). Review of the national external quality assessment (EQA) scheme for breast pathology in the UK. J Clin Pathol. 2017;70:51–57.
Al-Janabi S, van Slooten HJ, Visser M, van der Ploeg T, van Diest PJ, Jiwa M. Evaluation of mitotic activity index in breast cancer using whole slide digital images. PLoS One. 2013;8:e82576.
U.S. Food and Drug Administration, U.S. Department of Health and Human Services. 2018. https://www.fda.gov/newsevents/newsroom/pressannouncements/ucm552742.htm. Accessed 7 November 2018.
Pantanowitz L, Sinard JH, Henricks WH, Fatheree LA, Carter AB, Contis L, et al. Validating whole slide imaging for diagnostic purposes in pathology: guideline from the College of American Pathologists Pathology and Laboratory Quality Center. Arch Pathol Lab Med. 2013;137:1710–22.
Cross S, Furness P, Igali L, Snead D, Treanor D The Royal College of Pathologists guidelines: Best practice recommendations for implementing digital pathology. Version number 1. Protocol active since: January 2018.
Rakha EA, Aleskandarani M, Toss MS, Green AR, Ball G, Ellis IO, et al. Breast cancer histologic grading using digital microscopy: concordance and outcome association. J Clin Pathol. 2018;71:680–6.
Lakhani SR, Ellis IO, Schnitt SJ, Tan PH, vande Vijver MJ. WHO classification of tumours of the breast. 4th ed. Lyon: IARC Press; 2012.
Sobin LH, Wittekind C. TNM classification of malignant tumours (UICC). 6th ed. Hoboken: Wiley-Blackwell; 2002.
Sobin LH, Gospodarowicz MK, Wittekind C. TNM classification of malignant tumours (UICC). 7th ed. Hoboken: Wiley-Blackwell; 2011.
Acknowledgements
The authors thank the registration team of the Netherlands Comprehensive Cancer Registry (IKNL) for the collection of data. This study was funded by a Junior Researcher grant from the Radboud University Medical Center Institute for Health Sciences (RIHS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Jeroen van der Laak is member of the scientific advisory boards of Philips, the Netherlands and ContextVision, Sweden and receives research funding from Philips, the Netherlands and Sectra, Sweden. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Balkenhol, M.C.A., Tellez, D., Vreuls, W. et al. Deep learning assisted mitotic counting for breast cancer. Lab Invest 99, 1596–1606 (2019). https://doi.org/10.1038/s41374-019-0275-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-019-0275-0
This article is cited by
-
Enhancing mitosis quantification and detection in meningiomas with computational digital pathology
Acta Neuropathologica Communications (2024)
-
Artificial intelligence applications in histopathology
Nature Reviews Electrical Engineering (2024)
-
Automated mitotic spindle hotspot counts are highly associated with clinical outcomes in systemically untreated early-stage triple-negative breast cancer
npj Breast Cancer (2024)
-
OralEpitheliumDB: A Dataset for Oral Epithelial Dysplasia Image Segmentation and Classification
Journal of Imaging Informatics in Medicine (2024)
-
Current status and prospects of artificial intelligence in breast cancer pathology: convolutional neural networks to prospective Vision Transformers
International Journal of Clinical Oncology (2024)