Abstract
Dilated cardiomyopathy (DCM) remains a major cause of heart failure and carries a poor prognosis despite important advances in recent years. Better disease characterization using novel molecular techniques is needed to refine its progression. This study explored the proteomic signature of plasma-derived extracellular vesicles (EVs) obtained from DCM patients and healthy controls using size-exclusion chromatography (SEC). EV-enriched fractions were analyzed by liquid chromatography-mass spectrometry (LC-MS/MS). Raw data obtained from LC-MS/MS were analyzed against the Uniprot human database using MaxQuant software. Additional analyses using Perseus software were based on the Intensity-Based Absolute Quantification (iBAQ) values from MaxQuant analyses. A total of 90.07 ± 21 proteins (227 different proteins) in the DCM group and 96.52 ± 17.91 proteins (183 different proteins) in the control group were identified. A total of 176 proteins (74.6%) were shared by controls and DCM patients, whereas 51 proteins were exclusive for the DCM group and 7 proteins were exclusive for the control group. Fibrinogen (α, β and γ chain), serotransferrin, α-1-antitrypsin, and a variety of apolipoprotein family members (C-I, C-III, D, H or β-2-glycoprotein, and J or clusterin) were clustered in SEC-EVs derived from DCM patients relative to controls (p < 0.05). Regarding Gene Ontology analysis, response to stress and protein activation-related proteins were enriched in DCM-EVs compared with controls. Thus, the present study reports the distinct proteomic signature of circulating DCM-EVs compared with control-EVs. Furthermore, we confirm that SEC obtains highly purified EV fractions from peripheral blood samples for subsequent use in determining disease-specific proteomic signatures.
Similar content being viewed by others
Introduction
Despite important progress over the past few decades in the management of patients with dilated cardiomyopathy (DCM), it still portends a remarkable reduction in life expectancy and quality of life [1]. DCM is characterized by patchy interstitial fibrosis, degenerated cardiomyocytes, marked cardiac endothelial deficits, ventricular chamber enlargement or dilatation, systolic dysfunction with normal left ventricular (LV) wall thickness, progressive LV contractile dysfunction, ventricular and supraventricular arrhythmias, thromboembolism, and sudden death [2,3,4,5]. In the majority of cases, the primary cause is unknown, giving rise to the term ‘idiopathic’ DCM.
Biomarkers are indicators of variation in normal and pathological processes that are objectively measurable in biological systems [6]. In DCM, and by extension heart failure (HF), a variety of biomarkers reflective of different pathobiological processes (e.g., myocyte stress and stretch, myocyte apoptosis, cardiac interstitium, inflammation, oxidative stress, cardiac energetics, neurohormones and renal biomarkers) have been explored, but only natriuretic peptides are routinely used in clinical practice [7,8,9]. Molecular cardiology is progressing quickly, incorporating the potential of extracellular vesicles (EVs) and other moieties of proteins, lipids and circulating genomic sequences to refine disease progression.
Circulating EVs are vesicles with lipid bilayers that are produced by most cells and range in size from 50 to 200 nm [10]. The composition of EVs varies depending on the physiological and functional state of the producing cells, and EVs have been reported as potential biomarkers in a variety of diseases, including those of the cardiovascular system [11]. Thus, it has been speculated that the composition of circulating EVs in DCM may have a distinct signature [12]. Accordingly, we explored the proteomic signature of plasma-derived EVs obtained by size-exclusion chromatography (SEC) in DCM patients and healthy controls.
Materials and methods
Study population
Twenty DCM patients and 15 age- and sex-matched healthy controls with no cardiovascular disorders were recruited for peripheral venous blood extraction. All laboratory measurements were performed by experienced staff blinded to the patients’ clinical characteristics. The demographic characteristics of the DCM patients are shown in Supplementary Table 1. Briefly, this was a relatively young (62.3 ± 11.5 years) male (90%) cohort with reduced ejection fraction treated according to contemporary guidelines. Coronary angiography was performed in all DCM patients to rule out atherothrombosis. The duration of HF was 68.3 ± 60 months (median 40.6, Q1–Q3 [20.3–113.6]). None of the patients underwent heart transplantation during follow-up. The study protocol was approved by the Clinical Research Ethics Committee and conformed to the principles outlined in the Declaration of Helsinki [13]. Written informed consent was obtained from each donor.
Blood processing, and EV isolation and characterization
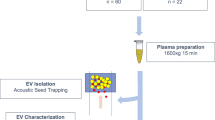
Peripheral blood was collected in BD Vacutainer™ Plastic Blood Collection Tubes with spray-coated K2EDTA (BD Bioscience, San Diego, CA) and processed within 4 h according to standard procedures [14]. In particular, all whole blood samples were first diluted 1:1 with calcium- and magnesium-free phosphate-buffered saline (PBS). Diluted blood samples were then layered carefully over 10–15 mL of Axis-Shield LymphoprepTM solution (1.077 g/mL; Alere Technologies AS, Oslo, Norway) and centrifuged at the same speed (i.e., 800 g) and with the same rotor type (SX4750A; Allegra X-12R centrifuge from Beckman Coulter) for 20 min. After centrifugation, the upper layer containing plasma from each sample was carefully removed from mononuclear cells (at the interface) and granulocytes/erythrocytes (bottom layers). Finally, collected plasmas were divided into single usage 1 mL aliquots to avoid later repeated freeze–thaw cycles and kept frozen at −80 °C until used for purification of EVs.
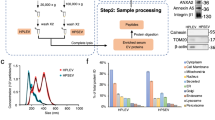
Two aliquots of plasma (~ 2 mL) from each donor stored at −80 °C were slowly defrosted on ice just once, loaded onto no-commercial 15 mL columns and separated by SEC using filtered PBS as the elution buffer [15]. SEC columns were prepared using Sepharose-CL2B with 75 nm pore size (Sigma-Aldrich, St. Louis, MO, USA) and a syringe 7 cm high and 1.9 cm width. After sepharose was well stacked for hours and stored at 4 °C, samples were loaded. In general, SEC fractions were both frozen and thawed rapidly for maximal preservation of morphology and function. The collected fractions (20 fractions of 0.5 mL each) were analyzed for the expression of specific EV markers (CD9, CD63 and CD81) by bead-based flow cytometry [12, 16, 17]. Briefly, a volume of each fraction was incubated with 0.5 µL aldehyde/sulfate–latex beads (4 µm; Invitrogen, Carlsbad, CA) for 15 min at room temperature (RT), resuspended in Bead Coupling Buffer (BCB) [PBS supplemented with 0.1% bovine serum albumin (BSA) and 0.01% NaN3 (Sigma-Aldrich)] and incubated overnight at RT. Thus, beads increase “vesicular size” and their detection capability by the cytometer. EV-coated beads were washed with the same buffer BCB and labeled with anti-CD9 (Clone VJ1/20), anti-CD63 (Clone TEA 3/18) and anti-CD81 (clone #G0709, from Santa Cruz Biotech) or polyclonal IgG isotype (Abcam, Cambridge, UK) antibodies for 30 min. After 30-min incubation with fluorescein isothiocyanate-conjugated goat anti-mouse secondary antibodies (SouthernBiotech, Birmingham, AL), and two washing steps with BCB, samples were analyzed by flow cytometry (FacsVerse; BD Biosciences, San Jose, CA) using the FlowJo software (Tree Star, Ashland, OR). Characterization was also performed by nanoparticle tracking analysis (NTA) and cryo-electron microscopy (cryo-EM) as previously described [12]. In particular, in NTA experiments, a total of 8660 ± 3051 completed tracks with a concentration of 50.2 ± 21.7 particles per frame were measured. Protein concentration of each fraction was also measured by absorbance 280 nm in Thermo Scientific Nanodrop® ND-1000 (Thermo Fisher Scientific, Waltham, MA). EV markers enriched fractions were also analyzed by Bradford assay (10 μL of sample; Bio-Rad Laboratories, USA) using a standard linear curve based on BSA (Sigma-Aldrich) before submitting to mass spectrometry.
Mass spectrometry
EV-enriched fractions were pooled and analyzed by liquid chromatography (nanoLCULTRA-EKSIGENT), followed by a 120-min gradient mass spectrometry (liquid chromatography-mass spectrometry (LC-MS/MS)) on an LTQ Orbitrap XL (Thermo Fisher, Carlsbad, CA). Briefly, previously lyophilized samples were dissolved in 100 µL of 6 M urea and 200 mM NH4HCO3 and sonicated for 15 min. The samples were finally digested with trypsin and LysC. As a quality control, BSA was digested in parallel and ran between each of the samples analyzed to avoid carryover and assess instrument performance. The samples were searched against the SwissProt Human database (April 2016) using the Mascot v2.5.1 algorithm (http://www.matrixscience.com/). Peptides were filtered based on false discovery rate (FDR); only peptides with FDR < 5% were retained.
Proteomic data processing and analysis
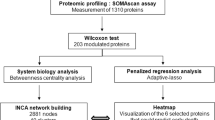
Raw data obtained from LC-MS/MS were analyzed using MaxQuant software (version 1.6.0.1) against the Uniprot human database (downloaded on 2 June 2017). Parameters set for single protein identification include: minimum peptide length of 7; maximum FDR for peptides and proteins of 1%; minimum peptides per protein of 1 and minimum unique peptides per protein of 0; minimum score for modified peptides of 40; and main search error of 4 ppm. In all searches, cysteine carbamidomethylation was established as a fixed modification, and methionine oxidation and acetylation of the N-terminus were established as variable modifications, with a maximum number of modifications per peptide of 5. Proteins identified as potential contaminants, those only identified by site or by a reverse sequence, and proteins with fewer than two unique peptides were not considered further. Additional analyses using Perseus software (version 1.6.0.2) were based on the Intensity-Based Absolute Quantification (iBAQ) values from MaxQuant analyses [18]. To that end, iBAQ values were logarithmized before performing the correlation plots, hierarchical clustering analysis, principal component analysis (PCA) and volcano plot. In the case of hierarchical clustering, z-score was calculated for the plot. Gene ontology (GO) terms for biological processes and cellular components were assessed using Perseus or the protein functional association network String (https://string-db.org). Data imputation to substitute non-quantified values with low valid intensities based on normal distribution (down-shift of 1.8 and distribution width of 0.3) were performed before PCA. The most significant proteins were identified by a volcano plot, plotting the fold-change difference of log2 iBAQ against -log2 (p-value). We have submitted all relevant data of our experiments to the EV-TRACK knowledgebase (EV-TRACK ID: EV170033) [19].
Statistical analysis
Data are presented as mean ± standard deviation (SD). Statistical analyses were performed using the Student’s t-test. Differences were considered significant when p < 0.05. For the volcano plot, s0 was set to 0.1.
Results
SEC-based isolation of plasma-derived EVs
Up to 20 chromatographic fractions were recovered by SEC per plasma sample. Fractions containing higher amounts of proteins (fraction 13 onward) were collected separately from those enriched by vesicular markers (CD9, CD63 and CD81), as determined by flow cytometry, and with undetectable protein content according to Abs 280 nm (Fig. 1a). By BCA, pooled fractions enriched in EVs, mainly fractions 8–10, showed a concentration among 50–100 µg. The CD9+, CD63+ and CD81+ fractions were then pooled and further recognized to contain typical EVs with the expected modal distribution (80–200 nm) and morphology (discernible membrane bilayer, round in shape) using NTA and cryo-EM methodologies (Figs. 1b, c).
SEC-based isolation and characterization of plasma EVs. a Representative analysis of EV-specific markers (CD9 in red, CD81 in green and CD63 in purple) in SEC fractions recovered from a DCM patient by flow cytometry. The amount of protein in the different EV-enriched fractions was also measured by the absorbance at 280 nm (blue line). Further analysis of pooled EV-enriched fractions by NTA b and cryo-EM c is also shown. Scale bar = 100 nm and ×25,000 magnification
Comparative cargo proteome analysis of SEC-EVs
The proteomic signatures of the EVs isolated from both groups were examined by mass spectrometry. After searching against Homo sapiens using MaxQuant software, a total of 344 proteins were identified, but only 234 fit the exclusion parameters used with Perseus software (identified by at least two unique peptides, not identified by reverse or by site). Bioinformatics analyses were applied, identifying a total of 90.07 ± 21 proteins (227 different proteins) in the DCM group and 96.52 ± 17.91 proteins (183 different proteins) in the control group. A total of 176 proteins (74.6%) were shared by controls and DCM patients, whereas 51 proteins were exclusive for the DCM group and 7 proteins were exclusive for the control group (Fig. 2). Intragroup similarities shown by the multi-scatter plots revealed higher correlation for DCM patients relative to controls (R = 0.77 ± 0.084 vs. R = 0.73 ± 0.10; Fig. 3). Of note, most of the proteins found in our samples are already described in the Proteomic Database Vesiclepedia (http://www.microvesicles.org) (Supplementary Figure 1).
Correlation analysis of plasma EVs. Multi-scatter plots analyzing intragroup (top plots) and inter-group (bottom plot) linear association and correlation within control and DCM samples analyzed. Pearson correlation values are labeled on each plot. The mean values are 0.73 ± 0.10, 0.77 ± 0.08 and 0.74 ± 0.08 for control, DCM and control vs. DCM samples, respectively
Data obtained from MaxQuant were submitted to PCA to assess differences in protein distribution between groups. Based on component 1, which accounts for 50.8% of sample variability, both EV groups exhibited a tendency to be differentially segregated (Fig. 4a). A volcano plot was generated to compare the significant differences in protein expression levels between both groups, revealing that 60 proteins were differentially expressed between EV groups (Fig. 4b). On the other hand, we found a set of common proteins that included the EV-specific markers galectin-3-binding protein, CD5 antigen-like protein, glyceraldehyde-3-phosphate dehydrogenase (Table 1), and others related to membrane-bound vesicles and extracellular exosomes and space as described after GO analyses (Table 1). Hierarchical clustering using the z-score of the samples more differentially clustered in the PCA was then obtained. Higher expression of the general proteome of DCM samples was observed compared with controls (Fig. 4c). Notably, fibrinogen (α, β and γ chain), serotransferrin, α-1-antitrypsin, and a variety of apolipoprotein family members (C-I, C-III, D, H or β-2-glycoprotein, and J or clusterin) were clustered in SEC-EVs derived from DCM patients relative to controls (p < 0.05; Table 2). Regarding GO analysis, stress response and protein activation-related proteins were enriched in DCM-EVs compared with controls (Fig. 5).
Comparative protein content analysis of plasma EVs. a Two-dimensional scatter plot of principal component analysis (PCA) showing components 1 and 2, which account for 50.8% and 7%, respectively, of the variability of all 236 proteins. A tendency to be clustered separately is circled for groups (DCM samples, green little circles; control samples, red little circles). b Representative volcano plot to calculate significantly differentially expressed proteins between groups. Each circle represents a protein. More than 60 proteins had p < 0.05. The first 20 proteins showing more significant differences are drawn as larger filled-in blue circles. c Higher expression was observed in general in DCM samples (red) compared with those from controls (green) in the hierarchical clustering obtained with the z-score for most differentially grouped samples
Discussion
DCM is a frequent cause of HF and often requires long and expensive therapies, including heart transplant or LV assist device implantation, resulting in an enormous burden on health care resources [1]. Because the value of a biomarker resides primarily in its capacity to reflect an association to diagnosis, prognosis or change in disease progression, there is currently intense research in the field of molecular cardiology to better identify and characterize disease-specific biomarkers. Companion biomarkers obtained from simple venipuncture may emerge in this new era as the liquid biopsy of the heart. However, a single biomarker is unlikely to reflect all facets of the HF syndrome, and a multi-marker strategy may better characterize the complexity of HF [20]. Better knowledge of the protein signature of the studied disease process of interest would aid in identifying such a multi-marker strategy. Here, we did not examine free circulating DCM-derived biomarkers, but those residing within circulating EVs. To the best of our knowledge, this study is the first to explore the proteomic signature of circulating DCM-EVs by SEC. Our data identified several proteins within circulating DCM-EVs that are not present in control-EVs, which set the pace for further analyzing whether they may be valid biomarkers of disease progression.
Shed under both normal and pathological conditions, EVs have been found in different biological fluids (e.g., blood, urine and saliva), and increasing evidence suggests that the information contained in these membrane vesicles are relevant diagnostic tools and a key source of biomarkers [21, 22]. Relative to currently available EV isolation methods, the protein content was undetectable in SEC fractions positive for tetraspanin markers, as most of the soluble proteins eluted well after fraction 10. According to the recommendations of the International Society for Extracelullar Vesicles, SEC is the cleanest method to efficiently isolate membrane bilayer-limited round-shaped EVs in a given sample [21]. As shown by NTA and cryo-EM analyses, SEC removes most of the over-represented plasma proteins for subsequent sophisticated analysis, such as proteomics [23].
Together with some robustly recognized specific membrane vesicle markers, a number of proteins, including fibrinogen, serotransferrin, α-1-antitrypsin, and several apolipoproteins, such as clusterin, were generally over-represented in the cargo proteome of circulating DCM-EVs compared with control-EVs. Some of these proteins have been reported to be free circulating biomarkers. Prospective studies support that increased levels of fibrinogen, which is crucial for platelet aggregation and contributes to endothelial injury, plasma viscosity and thrombus formation, are associated with a high risk of cardiovascular disease [24,25,26,27,28]. DCM patients often present with LV thrombi and exhibit a prothrombotic state mirroring that of high circulating fibrinogen levels [29]. Anemia, which is partially related to elevated serotransferrin levels, is a prevalent comorbidity in patients with HF and a marker of poor outcomes, with increased rehospitalization and mortality [30,31,32,33,34]. Also, proteomic analysis previously identified the broad-spectrum protease inhibitor α-1-antitrypsin as a putative biomarker for the evaluation of disease status [35, 36]. Clusterin is a ubiquitously expressed and versatile glycoprotein with cytoprotective properties that is involved in many physiological processes, including cell differentiation, lipid transportation, and DNA repair and cell cycle regulation [37]. This heat shock protein-like chaperone appears to be upregulated in diverse physiopathological settings of cell stress and tissue injury, such as myocardial infarction, ischemia, inflammation, apoptosis and oxidative stress, and is capable of preventing myocardial injury in experimental studies [38,39,40]. More recently, the prognostic value of plasma clusterin was demonstrated in patients with chronic HF. Koller et al. found an inverse correlation between clusterin levels and parameters of HF severity as indicated by NT-proBNP, LV ejection fraction and New York Heart Association class [41]. The authors speculated that the persistent and excessive tissue injury caused by inflammation and oxidative stress may lead to progressive depletion of circulating clusterin in these patients. However, our findings show that the amount of clusterin in the circulation of DCM patients is upregulated, but the majority of the protein appears to be profusely masked for conventional measurements (e.g., enzyme-linked immunosorbent assays), over-accumulating in circulating EVs. Thus, clusterin secretion is conversely promoted in the circulatory system of DCM patients, and we further hypothesize that its mechanism of action in the course of DCM worsening is closely regulated through the vesicular pathway.
Several limitations must be acknowledged with regard to the present study. Although the methodology of plasma preparation includes density gradient separation, which efficiently diminishes the presence of residual blood cells [14], other slowly sedimenting particles (i.e., platelets) can be detected in some extent. This should be taken into account in the validation of proteomics findings. Remarkably, the DCM-EVs identified from peripheral blood may have originated in any cell of the organism; we do not have evidence that they are provided by cardiomyocytes. Nevertheless, DCM is a cardiac derangement with implications in multiple organs once HF syndrome is established, as was the case in our patients. However, regardless of the origin, the EVs are reflective of the proteomic signature of the disease. Furthermore, none of the proteins reported in the DCM-EV signature are pathognomonic of DCM. Thus, if they are to become future biomarkers of DCM progression, it will have to be within multi-biomarker strategies that approach different disease pathways. Finally, another potential limitation of this study is selection bias; whether our findings apply to other HF cohorts of different etiologies needs to be investigated further.
In summary, the present study shows how the proteomic signature of circulating DCM-EVs compares with that of control-EVs. Among the identified proteins, our data indicate high clusterin levels within DCM-EVs, in contrast to the low circulating levels of clusterin previously reported in HF. Further research is needed to better understand its precise role in DCM progression. In addition, the present results confirm that SEC allows highly purified EV fractions derived from peripheral blood samples to be obtained for the subsequent search for disease-specific proteomic signatures. Larger well-designed studies and more standardized, cost-effective sample preparation and cargo analysis/validation protocols, including those increasing EV extraction yield and solubilization, are warranted if this technology aims to be center stage in the clinic in the near future.
References
Towbin JA, Bowles NE. The failing heart. Nature. 2002;415:227–33.
Jefferies JL, Towbin JA. Dilated cardiomyopathy. Lancet. 2010;375:752–62.
Towbin JA, Lorts A. Arrhythmias and dilated cardiomyopathy common pathogenetic pathways? J Am Coll Cardiol. 2011;57:2169–71.
Roura S, Planas F, Prat-Vidal C, et al. Idiopathic dilated cardiomyopathy exhibits defective vascularization and vessel formation. Eur J Heart Fail. 2007;9:995–1002.
Roura S, Bayes-Genis A. Vascular dysfunction in idiopathic dilated cardiomyopathy. Nat Rev Cardiol. 2009;6:590–8.
Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin Pharmacol Ther. 2001;69:89–95.
Roura S, Gálvez-Montón C, Fernández MA, et al. Circulating endothelial progenitor cells: potential biomarkers for idiopathic dilated cardiomyopathy. J Cardiovasc Transl Res. 2016;9:80–84.
Januzzi JL Jr.. Natriuretic peptides as biomarkers in heart failure. J Investig Med. 2013;61:950–5.
Gopal DM, Sam F. New and emerging biomarkers in left ventricular systolic dysfunction--insight into dilated cardiomyopathy. J Cardiovasc Transl Res. 2013;6:516–27.
Yáñez-Mó M, Siljander PR-M, Andreu Z, et al. Biological properties of extracellular vesicles and their physiological functions. J Extracell vesicles [Internet]. 2015;4:27066.
Lawson C, Vicencio JM, Yellon DM, et al. Microvesicles and exosomes: new players in metabolic and cardiovascular disease. J Endocrinol. 2016;228:R57–71.
Roura S, Gálvez-Montón C, de Gonzalo-Calvo D, et al. Extracellular vesicles do not contribute to higher circulating levels of soluble LRP1 in idiopathic dilated cardiomyopathy. J Cell Mol Med. 2017;21:3000–9. https://doi.org/10.1111/jcmm.13211.
Lynöe N, Sandlund M, Dahlqvist G, et al. Informed consent: study of quality of information given to participants in a clinical trial. BMJ. 1991;303:610–3.
Dagur PK, McCoy JP Jr. Collection, storage, and preparation of human blood cells. Curr Protoc Cytom. 2015;73:1–16.
Boing AN, Van Der Pol E, Grootemaat AE, et al. Single-step isolation of extracellular vesicles by size-exclusion chromatography. J Extracell Vesicles. 2014;3:23430.
Thery C, Amigorena S, Raposo G, et al. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol. 2006;3:22.
Suárez H, Gámez-Valero A, Reyes R, et al. A bead-assisted flow cytometry method for the semi-quantitative analysis of extracellular vesicles. Sci Rep. 2017;7:11271.
Tyanova S, Temu T, Sinitcyn P, et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods. 2016;13:731–40.
EV-TRACK Consortium, Van Deun J, Mestdagh P, et al. EV-TRACK: transparent reporting and centralizing knowledge in extracellular vesicle research. Nat Methods. 2017;14:228–32.
Bayes-Genis A, Núñez J, Núñez E, et al. Multi-biomarker profiling and recurrent hospitalizations in heart failure. Front Cardiovasc Med. 2016;3:37. (eCollection 2016).
Sáenz-Cuesta M, Mittelbrunn M, Otaegui D. Editorial: novel clinical applications of extracellular vesicles. Front Immunol. 2015;6:381.
Lener T, Gimona M, Aigner L, et al. Applying extracellular vesicles based therapeutics in clinical trials - an ISEV position paper. J Extracell Vesicles. 2015;4:30087.
Gámez-Valero A, Monguió-Tortajada M, Carreras-Planella L, et al. Size-exclusion chromatography-based isolation minimally alters extracellular vesicles’ characteristics compared to precipitating agents. Sci Rep. 2016;6:33641.
Lowe GD. Fibrinogen assays for cardiovascular risk assessment. Clin Chem. 2010;56:693–5.
Danesh J, Lewington S, Thompson SG, et al. Plasma fibrinogen level and the risk of major cardiovascular diseases and nonvascular mortality: an individual participant metaanalysis. JAMA. 2005;294:1799–809.
Kaptoge S, Di Angelantonio E, Pennells L, et al. C-reactive protein, fibrinogen, and cardiovascular disease prediction. N Engl J Med. 2012;367:1310–20.
Appiah D, Schreiner PJ, MacLehose RF, et al. Association of plasma gamma’ fibrinogen with incident cardiovascular disease: the Atherosclerosis Risk in Communities (ARIC) study. Arterioscler Thromb Vasc Biol. 2015;35:2700–6.
Becatti M, Marcucci R, Bruschi G, et al. Oxidative modification of fibrinogen is associated with altered function and structure in the subacute phase of myocardial infarction. Arterioscler Thromb Vasc Biol. 2014;34:1355–61.
Abdo AS, Kemp R, Barham J, et al. Dilated cardiomyopathy and role of antithrombotic therapy. Am J Med Sci. 2010;339:557–60.
Klip IT, Comin-Colet J, Voors AA, et al. Iron deficiency in chronic heart failure: an international pooled analysis. Am Heart J. 2013;165:575–82.
Comin-Colet J, Ruiz S, Cladellas M, et al. A pilot evaluation of the long-term effect of combined therapy with intravenous iron sucrose and erythropoietin in elderly patients with advanced chronic heart failure and cardio-renal anemia syndrome: influence on neurohormonal activation and clinical outcomes. J Card Fail. 2009;15:727–35.
Jankowska EA, Rozentryt P, Witkowska P, et al. Iron deficiency predicts impaired exercise capacity in patients with systolic chronic heart failure. J Card Fail. 2011;17:899–906.
Enjuanes C, Klip IT, Bruguera J, et al. Iron deficiency and health-related quality of life in chronic heart failure: results from a multicenter European study. Int J Cardiol. 2014;174:268–75.
Van Veldhuisen DJ, Anker SD, Ponikowski P, et al. Anemia and iron deficiency in heart failure: mechanisms and therapeutic approaches. Nat Rev Cardiol. 2011;8:485–93.
Gettins PG. Serpin structure, mechanism, and function. Chem Rev. 2002;102:4751–804.
Izquierdo I, Rosa I, Bravo SB, et al. Proteomic identification of putative biomarkers for early detection of sudden cardiac death in a family with a LMNA gene mutation causing dilated cardiomyopathy. J Proteom. 2016;148:75–84.
Jones SE, Jomary C. Clusterin. Int J Biochem Cell Biol. 2002;34:427–31.
Wyatt A, Yerbury J, Poon S, et al. Chapter 6: the chaperone action of clusterin and its putative role in quality control of extracellular protein folding. Adv Cancer Res. 2009;104:89–114.
Park S, Mathis KW, Lee IK. The physiological roles of apolipoprotein J/clusterin in metabolic and cardiovascular diseases. Rev Endocr Metab Disord. 2014;15:45–53.
Trougakos IP, Poulakou M, Stathatos M, et al. Serum levels of the senescence biomarker clusterin/apolipoprotein J increase significantly in diabetes type II and during development of coronary heart disease or at myocardial infarction. Exp Gerontol. 2002;37:1175–87.
Koller L, Richter B, Winter MP, et al. Clusterin/apolipoprotein J is independently associated with survival in patients with chronic heart failure. J Clin Lipidol. 2017;11:178–84.
Acknowledgements
We specifically acknowledge Dr. Francisco Sánchez-Madrid (Servicio de Inmunología, Hospital Universitario de la Princesa, IIS-IP, UAM; Cell-Cell Communication Laboratory, CNIC) and Dr. María Yáñez-Mó (Unidad de Investigación, Hospital Sta Cristina, IIS-IP; Departamento Biología Molecular/CBM-SO, UAM) for kindly gifting antibodies against CD9, CD63 and CD81. We also thank Beatriz González, Roser Cabanes, Margarita Rodríguez, Carmen Rivas, Nuria Benito and Alba Ros for invaluable work in the Heart Failure Clinic; Eduard Sabido and Eva Borràs from the UPF/CRG Proteomics Unit for proteomic data processing and analysis; and Hernando del Portillo (ICREA Research Professor at ISGLOBAL-IGTP) for access to the NTA instrument.
Funding
This work was supported by grants from the Ministerio de Economía, Industria y Competitividad (SAF2014-59892-R), Fundació La MARATÓ de TV3 (201405/10, 201502, 201516), Societat Catalana de Cardiologia, Generalitat de Catalunya (SGR 2014, CERCA Programme) and the Fundació Bancària La Caixa. This work was also funded by the Red de Terapia Celular—TerCel (RD16/0011/0006), CIBER Cardiovascular—(CB16/11/00403), and Fondo de Investigación Sanitaria, Instituto de Salud Carlos III (FIS PI14/01682) as part of the Plan Nacional de I+D+I cofounded by ISCIII-Sudirección General de Evaluación y el Fondo Europeo de Desarrollo Regional (FEDER).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Roura, S., Gámez-Valero, A., Lupón, J. et al. Proteomic signature of circulating extracellular vesicles in dilated cardiomyopathy. Lab Invest 98, 1291–1299 (2018). https://doi.org/10.1038/s41374-018-0044-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41374-018-0044-5
This article is cited by
-
Circulating cardiomyocyte-derived extracellular vesicles reflect cardiac injury during systemic inflammatory response syndrome in mice
Cellular and Molecular Life Sciences (2022)
-
Platelet proteome changes in dogs with congestive heart failure
BMC Veterinary Research (2020)
-
Extracellular vesicle isolation methods: rising impact of size-exclusion chromatography
Cellular and Molecular Life Sciences (2019)
-
Quantitative proteomics study reveals differential proteomic signature in dilated, restrictive, and hypertrophic cardiomyopathies
Journal of Proteins and Proteomics (2019)