Abstract
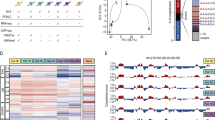
The dynamics of transcription factors play important roles in a variety of biological systems. However, the mechanisms by which these dynamics are decoded into different transcriptional responses are not well understood. Here we focus on the dynamics of the tumor-suppressor protein p53, which exhibits a series of pulses in response to DNA damage. We performed time course RNA sequencing (RNA-seq) and chromatin immunoprecipitation sequencing (ChIP-seq) measurements to determine how p53 oscillations are linked with gene expression genome wide. We discovered multiple distinct patterns of gene expression in response to p53 pulses. Surprisingly, p53-binding dynamics were uniform across all genomic loci, even for genes that exhibited distinct mRNA dynamics. Using a mathematical model, supported by additional experimental measurements in response to sustained p53 input, we determined that p53 binds to and activates transcription of its target genes uniformly, whereas post-transcriptional mechanisms are responsible for the differences in gene expression dynamics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Cai, L., Dalal, C.K. & Elowitz, M.B. Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455, 485–490 (2008).
Hoffmann, A., Levchenko, A., Scott, M.L. & Baltimore, D. The IkappaB-NF-kappaB signaling module: temporal control and selective gene activation. Science 298, 1241–1245 (2002).
Purvis, J.E. et al. p53 dynamics control cell fate. Science 336, 1440–1444 (2012).
Paek, A.L., Liu, J.C., Loewer, A., Forrester, W.C. & Lahav, G. Cell-to-cell variation in p53 dynamics leads to fractional killing. Cell 165, 631–642 (2016).
Hao, N. & O'Shea, E.K. Signal-dependent dynamics of transcription factor translocation controls gene expression. Nat. Struct. Mol. Biol. 19, 31–39 (2011).
Covert, M.W., Leung, T.H., Gaston, J.E. & Baltimore, D. Achieving stability of lipopolysaccharide-induced NF-kappaB activation. Science 309, 1854–1857 (2005).
Batchelor, E., Loewer, A., Mock, C. & Lahav, G. Stimulus-dependent dynamics of p53 in single cells. Mol. Syst. Biol. 7, 488 (2011).
Hansen, A.S. & O'Shea, E.K. Encoding four gene expression programs in the activation dynamics of a single transcription factor. Curr. Biol. 26, R269–R271 (2016).
Werner, S.L., Barken, D. & Hoffmann, A. Stimulus specificity of gene expression programs determined by temporal control of IKK activity. Science 309, 1857–1861 (2005).
Lahav, G. et al. Dynamics of the p53-Mdm2 feedback loop in individual cells. Nat. Genet. 36, 147–150 (2004).
Lev Bar-Or, R. et al. Generation of oscillations by the p53-Mdm2 feedback loop: a theoretical and experimental study. Proc. Natl. Acad. Sci. USA 97, 11250–11255 (2000).
Weinberg, R.L., Veprintsev, D.B., Bycroft, M. & Fersht, A.R. Comparative binding of p53 to its promoter and DNA recognition elements. J. Mol. Biol. 348, 589–596 (2005).
Qian, H., Wang, T., Naumovski, L., Lopez, C.D. & Brachmann, R.K. Groups of p53 target genes involved in specific p53 downstream effects cluster into different classes of DNA binding sites. Oncogene 21, 7901–7911 (2002).
Kracikova, M., Akiri, G., George, A., Sachidanandam, R. & Aaronson, S.A. A threshold mechanism mediates p53 cell fate decision between growth arrest and apoptosis. Cell Death Differ. 20, 576–588 (2013).
Murray-Zmijewski, F., Slee, E.A. & Lu, X. A complex barcode underlies the heterogeneous response of p53 to stress. Nat. Rev. Mol. Cell Biol. 9, 702–712 (2008).
Flores, E.R. et al. p63 and p73 are required for p53-dependent apoptosis in response to DNA damage. Nature 416, 560–564 (2002).
Samuels-Lev, Y. et al. ASPP proteins specifically stimulate the apoptotic function of p53. Mol. Cell 8, 781–794 (2001).
Oda, K. et al. p53AIP1, a potential mediator of p53-dependent apoptosis, and its regulation by Ser-46-phosphorylated p53. Cell 102, 849–862 (2000).
Smeenk, L. et al. Role of p53 serine 46 in p53 target gene regulation. PLoS One 6, e17574 (2011).
Sykes, S.M. et al. Acetylation of the p53 DNA-binding domain regulates apoptosis induction. Mol. Cell 24, 841–851 (2006).
Nikulenkov, F. et al. Insights into p53 transcriptional function via genome-wide chromatin occupancy and gene expression analysis. Cell Death Differ. 19, 1992–2002 (2012).
Verfaillie, A. et al. Multiplex enhancer-reporter assays uncover unsophisticated TP53 enhancer logic. Genome Res. 26, 882–895 (2016).
Porter, J.R., Fisher, B.E. & Batchelor, E. p53 pulses diversify target gene expression dynamics in an mRNA half-life-dependent manner and delineate co-regulated target gene subnetworks. Cell Syst. 2, 272–282 (2016).
Melanson, B.D. et al. The role of mRNA decay in p53-induced gene expression. RNA 17, 2222–2234 (2011).
Batchelor, E., Mock, C.S., Bhan, I., Loewer, A. & Lahav, G. Recurrent initiation: a mechanism for triggering p53 pulses in response to DNA damage. Mol. Cell 30, 277–289 (2008).
Wei, C.L. et al. A global map of p53 transcription-factor binding sites in the human genome. Cell 124, 207–219 (2006).
El-Deiry, W.S. et al. WAF1, a potential mediator of p53 tumor suppression. Cell 75, 817–825 (1993).
Botcheva, K., McCorkle, S.R., McCombie, W.R., Dunn, J.J. & Anderson, C.W. Distinct p53 genomic binding patterns in normal and cancer-derived human cells. Cell Cycle 10, 4237–4249 (2011).
El-Deiry, W.S., Kern, S.E., Pietenpol, J.A., Kinzler, K.W. & Vogelstein, B. Definition of a consensus binding site for p53. Nat. Genet. 1, 45–49 (1992).
Riley, T., Sontag, E., Chen, P. & Levine, A. Transcriptional control of human p53-regulated genes. Nat. Rev. Mol. Cell Biol. 9, 402–412 (2008).
Fischer, M., Steiner, L. & Engeland, K. The transcription factor p53: not a repressor, solely an activator. Cell Cycle 13, 3037–3058 (2014).
Schlereth, K. et al. Characterization of the p53 cistrome–DNA binding cooperativity dissects p53′s tumor suppressor functions. PLoS Genet. 9, e1003726 (2013).
Rashi-Elkeles, S. et al. Parallel profiling of the transcriptome, cistrome, and epigenome in the cellular response to ionizing radiation. Sci. Signal. 7, rs3 (2014).
Schueler, M. et al. Differential protein occupancy profiling of the mRNA transcriptome. Genome Biol. 15, R15 (2014).
Zambrano, S., De Toma, I., Piffer, A., Bianchi, M.E. & Agresti, A. NF-κB oscillations translate into functionally related patterns of gene expression. eLife 5, e09100 (2016).
Hao, S. & Baltimore, D. The stability of mRNA influences the temporal order of the induction of genes encoding inflammatory molecules. Nat. Immunol. 10, 281–288 (2009).
Brummelkamp, T.R., Bernards, R. & Agami, R. A system for stable expression of short interfering RNAs in mammalian cells. Science 296, 550–553 (2002).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Kuleshov, M.V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Chen, E.Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics 14, 128 (2013).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, Q., Brown, J.B., Huang, H. & Bickel, P.J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Landt, S.G. et al. ChIP-seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Crooks, G.E., Hon, G., Chandonia, J.M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
We thank S. Boswell for help with RNA-seq experiments and X. Zhang (Broad Institute) for help with p53 ChIP experiments, M. Springer for helpful discussions and advice on modeling, and members of the Lahav lab for comments and discussion. This research was supported by Novartis Institutes for Biomedical Research and NIH grant GM083303. A.H. was supported by Boehringer Ingelheim Fonds, PhD fellowship. J.S.-O. was supported by NIH grant CA207727. J.E.P. was supported by NIH grant GM102372. M.L.B. was supported by NIH grant HG003985.
Author information
Authors and Affiliations
Contributions
A.H., J.E.P., W.C.F. and G.L. conceived experiments. A.H., M.L.B. and G.L. conceived analyses. A.H. performed experiments and A.H. and J.S.-O. performed analyses. A.H. and G.L. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of gene expression clusters shows p53 dependence and enrichment of known p53 target genes among induced genes. (related to Figure 1)
A. Quantiles (median in black, shaded area represents the 25% and 75% quantiles) of RNA-Seq data by cluster for MCF7 p53 wild-type cell line (blue) and MCF7 p53-shRNA cell line (gray). B. Mapping of the genes in each cluster to known target genes, obtained from Riley et al. 200830. P-values were calculated using the Binomial statistic, * = p-value<0.01. C. Top five enriched GO Biological Process categories in each cluster (FDR<0.05). GO Enrichment analysis was done using Enrichr software39,40. Both p-value and FDR are shown.
Supplementary Figure 2 Properties of p53 ChIP peaks that map to the induced clusters (related to Figure 3).
Distribution of maximal reads of p53 peaks per cluster. T-test was done to evaluate significance; ns stands for not significant (pval >0.2 for all comparisons). Black lines indicate the median, and the boxes and whiskers extend to the 25%-75% and the 5%-95% quantiles, respectively.
Supplementary Figure 3 Evaluation of model fit and parameters. (Related to Figures 4 and 5).
A. Distribution of R2 values for the model fit. Comparison between induced genes with p53 ChIP peak and induced genes without a p53 ChIP peak. B. Correlation of model derived kp values with maximal p53 ChIP peak signal for each gene. The signal from the closest peak was used for genes with multiple p53 ChIP peaks. C. Correlation of model derived kd values with published mRNA half-life data in MCF7 cells34. D. Schematic of gene expression dynamical features that were computed for the model stimulation. E-H. Simulation of the model (equation 1) for the shown range of kp and kd values. Heat maps of the gene expression properties derived for each kp and kd combination. I. Distribution of R2 values for the model fit for sustained data. J. Model simulation with p53 sustained input and only varying the kd parameter values.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Table 1. (PDF 501 kb)
Supplementary Data Set 1
229 differentially expressed genes (Figure 1). (XLSX 67 kb)
Mean FPKM, log2(fold change) across 2 independent experiments as well as cluster assignments are provided.
Supplementary Data Set 2
p53 and H3K27ac ChIP peaks (Figure 2). (XLSX 4738 kb)
Genomic positions, normalized read counts per time-point for the ChIP and corresponding Input samples are provided. Closest gene assignments, with distance to TSS (bp) are provided for p53 ChIP peaks. Distance (bp) to the closest p53 ChIP peak is given for all H3K27ac peaks (used for Figure 2H).
Supplementary Data Set 3
p53 bound and differentially induced genes (Figure 3B) and results of model fit (Figure 4). (XLSX 11 kb)
Cluster number assignments and the kp and kd parameters resulting from the model fit (Figure 4) are provided.
Supplementary Data Set 4
Gene expression under the sustained p53 condition (Figure 5). (XLSX 16 kb)
Cluster assignments, mean FPKM values across 2 independent replicates and the resulting R2 values from the model prediction are provided.
Supplementary Data Set 5
Un-cropped western blot images for Figures 1A, 2A, 5A. (PDF 785 kb)
Rights and permissions
About this article
Cite this article
Hafner, A., Stewart-Ornstein, J., Purvis, J. et al. p53 pulses lead to distinct patterns of gene expression albeit similar DNA-binding dynamics. Nat Struct Mol Biol 24, 840–847 (2017). https://doi.org/10.1038/nsmb.3452
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3452
This article is cited by
-
p53 rapidly restructures 3D chromatin organization to trigger a transcriptional response
Nature Communications (2024)
-
Orthogonal inducible control of Cas13 circuits enables programmable RNA regulation in mammalian cells
Nature Communications (2024)
-
p53-dependent DNA repair during the DNA damage response requires actin nucleation by JMY
Cell Death & Differentiation (2023)
-
CRISPR screens reveal genetic determinants of PARP inhibitor sensitivity and resistance in prostate cancer
Nature Communications (2023)
-
Genome-wide CRISPR screening reveals ADCK3 as a key regulator in sensitizing endometrial carcinoma cells to MPA therapy
British Journal of Cancer (2023)