Abstract
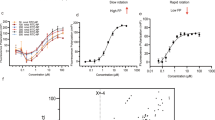
Enterovirus 71 (HEV71) epidemics in children and infants result mainly in mild symptoms; however, especially in the Asia-Pacific region, infection can be fatal. At present, no therapies are available. We have used structural analysis of the complete virus to guide the design of HEV71 inhibitors. Analysis of complexes with four 3-(4-pyridyl)-2-imidazolidinone derivatives with varying anti-HEV71 activities pinpointed key structure-activity correlates. We then identified additional potentially beneficial substitutions, developed methods to reliably triage compounds by quantum mechanics–enhanced ligand docking and synthesized two candidates. Structural analysis and in vitro assays confirmed the predicted binding modes and their ability to block viral infection. One ligand (with IC50 of 25 pM) is an order of magnitude more potent than the best previously reported inhibitor and is also more soluble. Our approach may be useful in the design of effective drugs for enterovirus infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
McMinn, P.C. Enterovirus 71 in the Asia-Pacific region: an emerging cause of acute neurological disease in young children. Neurol J. Southeast Asia 8, 57–63 (2003).
Rossmann, M.G. et al. Structure of a human common cold virus and functional relationship to other picornaviruses. Nature 317, 145–153 (1985).
Ren, J. et al. Picornavirus uncoating intermediate captured in atomic detail. Nat. Commun. 4, 1929 (2013).
Wang, X. et al. A sensor-adaptor mechanism for enterovirus uncoating from structures of EV71. Nat. Struct. Mol. Biol. 19, 424–429 (2012).
Rotbart, H.A. Treatment of picornavirus infections. Antiviral Res. 53, 83–98 (2002).
Tsang, S.K. et al. A structurally biased combinatorial approach for discovering new anti-picornaviral compounds. Chem. Biol. 8, 33–45 (2001).
Phelps, D.K. & Post, C.B. A novel basis of capsid stabilization by antiviral compounds. J. Mol. Biol. 254, 544–551 (1995).
Tsang, S.K., Danthi, P., Chow, M. & Hogle, J.M. Stabilization of poliovirus by capsid-binding antiviral drugs is due to entropic effects. J. Mol. Biol. 296, 335–340 (2000).
De Palma, A.M., Vliegen, I., De Clercq, E. & Neyts, J. Selective inhibitors of picornavirus replication. Med. Res. Rev. 28, 823–884 (2008).
Feil, S.C. et al. An orally available 3-ethoxybenzisoxazole capsid binder with clinical activity against human rhinovirus. Acs Med. Chem. Lett. 3, 303–307 (2012).
Pevear, D.C., Tull, T.M., Seipel, M.E. & Groarke, J.M. Activity of pleconaril against enteroviruses. Antimicrob. Agents Chemother. 43, 2109–2115 (1999).
Shia, K.S. et al. Design, synthesis, and structure-activity relationship of pyridyl imidazolidinones: a novel class of potent and selective human enterovi-rus 71 inhibitors. J. Med. Chem. 45, 1644–1655 (2002).
Ke, Y.Y. & Lin, T.H. Modeling the ligand-receptor interaction for a series of inhibitors of the capsid protein of enterovirus 71 using several three-dimensional quantitative structure-activity relationship techniques. J. Med. Chem. 49, 4517–4525 (2006).
Plevka, P., Perera, R., Cardosa, J., Kuhn, R.J. & Rossmann, M.G. Crystal structure of human enterovirus 71. Science 336, 1274 (2012).
Plevka, P. et al. Structure of human enterovirus 71 in complex with a capsid-binding inhibitor. Proc. Natl. Acad. Sci. USA 110, 5463–5467 (2013).
von Itzstein, M. et al. Rational design of potent sialidase-based inhibitors of influenza virus replication. Nature 363, 418–423 (1993).
Warren, G.L. et al. A critical assessment of docking programs and scoring functions. J. Med. Chem. 49, 5912–5931 (2006).
Axford, D. et al. In situ macromolecular crystallography using microbeams. Acta Crystallogr. D Biol. Crystallogr. 68, 592–600 (2012).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Chang, C.S. et al. Design, synthesis, and antipicornavirus activity of 1-[5-(4-arylphenoxy)alkyl]-3-pyridin-4-ylimidazolidin-2-one derivatives. J. Med. Chem. 48, 3522–3535 (2005).
Cho, A.E., Guallar, V., Berne, B.J. & Friesner, R. Importance of accurate charges in molecular docking: quantum mechanical/molecular mechanical (QM/MM) approach. J. Comput. Chem. 26, 915–931 (2005).
Walter, T.S. et al. A plate-based high-throughput assay for virus stability and vaccine formulation. J. Virol. Methods 185, 166–170 (2012).
Goodford, P.J. A computational procedure for determining energetically favorable binding sites on biologically important macromolecules. J. Med. Chem. 28, 849–857 (1985).
Schüttelkopf, A.W. & van Aalten, D.M. PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 60, 1355–1363 (2004).
Kenakin, T.P. Pharmacologic Analysis of Drug-Receptor Interaction. (Lippincott Raven, 1993).
Joseph-McCarthy, D., Hogle, J.M. & Karplus, M. Use of the multiple copy simultaneous search (MCSS) method to design a new class of picornavirus capsid binding drugs. Proteins 29, 32–58 (1997).
Joseph-McCarthy, D., Tsang, S.K., Filman, D.J., Hogle, J.M. & Karplus, M. Use of MCSS to design small targeted libraries: application to picornavirus ligands. J. Am. Chem. Soc. 123, 12758–12769 (2001).
Hiremath, C.N., Grant, R.A., Filman, D.J. & Hogle, J.M. Binding of the antiviral drug WIN51711 to the sabin strain of type 3 poliovirus: structural comparison with drug binding in rhinovirus 14. Acta Crystallogr. D Biol. Crystallogr. 51, 473–489 (1995).
Grant, R.A. et al. Structures of poliovirus complexes with anti-viral drugs: implications for viral stability and drug design. Curr. Biol. 4, 784–797 (1994).
Zhang, Y. et al. Structural and virological studies of the stages of virus replication that are affected by antirhinovirus compounds. J. Virol. 78, 11061–11069 (2004).
Lentz, K.N. et al. Structure of poliovirus type 2 Lansing complexed with antiviral agent SCH48973: comparison of the structural and biological properties of three poliovirus serotypes. Structure 5, 961–978 (1997).
Diana, G.D. et al. A model for compounds active against human rhinovirus-14 based on X-ray crystallography data. J. Med. Chem. 33, 1306–1311 (1990).
Lipinski, C.A., Lombardo, F., Dominy, B.W. & Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 46, 3–26 (2001).
Hadfield, A.T. et al. Structural studies on human rhinovirus 14 drug-resistant compensation mutants. J. Mol. Biol. 253, 61–73 (1995).
Hopkins, A.L. et al. Design of non-nucleoside inhibitors of HIV-1 reverse transcriptase with improved drug resistance properties. 1. J. Med. Chem. 47, 5912–5922 (2004).
Walter, T.S. et al. A procedure for setting up high-throughput nanolitre crystallization experiments. I. Protocol design and validation. J. Appl. Crystallogr. 36, 308–314 (2003).
Walter, T.S. et al. A procedure for setting up high-throughput nanolitre crystallization experiments: crystallization workflow for initial screening, automated storage, imaging and optimization. Acta Crystallogr. D Biol. Crystallogr. 61, 651–657 (2005).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
French, S. & Wilson, K. On the treatment of negative intensity observations. Acta Crystallogr. A 34, 517–525 (1978).
Brünger, A.T. et al. Crystallography & NMR system (CNS), a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Nicholls, R.A., Long, F. & Murshudov, G.N. Low-resolution refinement tools in REFMAC5. Acta Crystallogr. D Biol. Crystallogr. 68, 404–417 (2012).
Cowtan, K. Recent developments in classical density modification. Acta Crystallogr. D Biol. Crystallogr. 66, 470–478 (2010).
Kleywegt, G.J. Dictionaries for Heteros. CCP4/ESF-EACBM Newsletter Prot. Crystallogr. 31, 45–50 (1995).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Pettersen, E.F. et al. UCSF Chimera: a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Banks, J.L. et al. Integrated modeling program, applied chemical theory (IMPACT). J. Comput. Chem. 26, 1752–1780 (2005).
Reed, L.J.M.H. A simple method of estimating fifty percentage endpoints. Am. J. Hyg. 27, 493–497 (1938).
Tetko, I.V. et al. Virtual computational chemistry laboratory: design and description. J. Comput. Aided Mol. Des. 19, 453–463 (2005).
Acknowledgements
We thank A. Kotecha for assistance with Diamond data collection and the beamline staff at Diamond light source beamlines I03 and I24 for expert assistance and advice. GPP and GPV compounds were made by F. Prauchart (Department of Pharmaceutical Chemistry, University of Innsbruck). MS analyses were carried out by C. Schofield, and P. Abrusci helped with the Sigma Plot program. Administrative and high-performance computing was supported by the Wellcome Trust Core Award, grant no. 090532/Z, and particular help was provided by R. Esnouf. Work was supported by the Chinese National Major Project of Infectious Disease, Ministry of Science and Technology 973 Project (grant nos. 2011CB910300 and 2014CB542800) and Major National Science and Technology Programs (grant no. 2012ZX10004701). D.I.S., E.E.F. and T.S.W. are supported by the UK Medical Research Council (G110525 and G100099), J.R. by the Wellcome Trust, J.K. by Sanofi Pasteur and L.D.C. by the World Health Organization. Research leading to these results received funding from the European Union FP7, SILVER grant no. 260644.
Author information
Authors and Affiliations
Contributions
D.I.S. and Z.R. supervised and coordinated the project; J.W., Z.H., X.L., G.P. and W.P. made samples available; X.W. purified and crystallized the samples and performed thermofluor experiments; G.P. and J.G. provided the GPP ligands; L.D.C. designed ALD and NLD; L.D.C. and J.A.B.S. ran the in silico docking and analyzed data under supervision by D.I.S.; L.D.C. and T.S.W. soaked crystals for data collection, which was performed by L.D.C., J.A.B.S., J.R. and E.E.F.; L.D.C., J.A.B.S., J.R. and D.I.S. contributed to data processing, structure determination and model building; J.K. performed the in vitro TCID50 assay and together with N.S. and D.J.R. analyzed the data. L.D.C., E.E.F. and D.I.S., in discussion with J.R., D.J.R. and Z.R., wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Comparison of complexes of EV71 with sphingosine and GPP2.
Sphingosine (blue) - EV71 VP1 (blue) (PDBID: 3VBF), superimposed on GPP2 (magenta) - EV71 VP1 (cyan). The RMS difference between all Cα atoms of the icosahedral protomer is 0.4 Å.
Supplementary Figure 2 Predicted binding affinity (GOLD score or ΔG) versus experimental –log(IC50 (pIC50) values.
The compounds are the 3-(4-Pyridyl)-2-imidazolidinone derivatives listed in Supplementary Table 1. The calculated points for ALD and NLD are in red whilst the experimental values are reported in yellow on the plot. Plot (a) shows results for GOLD (with GOLD fitness scores), (b) for Glide (energy of interaction expressed as van der Waals plus electrostatic interactions) (c) for Rosetta (interface_delta), (d) GOLD/AMBER (ΔG of binding). For details of the protocols see the methods section of the main text. The ligand 42-8e reported in the Supplementary Table 1 could not be docked into the crystal structure using Glide. The following ligands: 2-d-21, 10-26, 4-24 reported in the Supplementary Table 1 were excluded from the Rosetta plot. They represent outlier points, because these molecules were partially docked in the binding.
Supplementary Figure 3 Stereo diagrams comparing the results of the various docking protocols for GPP2.
The side chain of Ile113 is not shown, to reveal the hydrogen bond, drawn as dotted lines, between the main chain nitrogen of this residue and the ligand oxygen. Residues from VP1 are shown in blue and Ile24 from VP3 in orange. (a) Overlay of the docked ligand GPP2 (light green) by QMPLD method and the structurally determined conformation of GPP2 (magenta) (RMSD for all inhibitor atoms 1.3 Å). (b) Overlay of the docked ligand GPP2 by GOLD (violet) and the structurally determined conformation of GPP2 (magenta) (RMSD for all inhibitor atoms 0.9 Å). (c) Overlay of the docked ligand GPP2 by Rosetta (gray) and the structurally determined conformation of GPP2 (magenta) (note GPP2 is docked upside down).
Supplementary Figure 4 Docking of GPP3 and NLD molecules in HEV71 crystal structures.
(a) Docking of GPP3 by QMPLD into the 2.7 Å crystal structure reported by Plevka et al. 2013 (reference 15 in main text, PDBID:3ZFE). In cyan the sphingosine molecule found in the 3ZFE crystal structure and in grey GPP3 docked by QMPLD. The dashed line represents the hydrogen bond between GPP3 and Ile113 of VP1. (b) In magenta the GPP3 molecule found in the crystal structure and in grey GPP3 molecule docked by QMPLD. The dashed line represents the hydrogen bond between GPP3 and Ile113 of VP1. The residue Phe135 is at the front whereas residue Phe155 is towards the back of the figure. Residue Ile24 of VP3 is coloured orange. The RMSD between the two virus structures (all atoms) is 0.8 Å, whilst the RMSD between the two GPP3 positions is 1.6 Å. (c) Docking of GPP3 by QMPLD into the 3.7 Å crystal structure reported by Plevka et al. 2013 (reference 14 in main text, PDBID:4AED) with the experimentally observed conformation. In gray GPP3 molecule docked by QMPLD into the 4AED crystal structure and in magenta the experimentally observed conformation of GPP3 (obtained by superposition of our complex onto the 4AED structure). (d) In grey the protonated form of the GPP3 molecule docked by QMPLD into the 4AED crystal structure and in magenta the experimentally observed conformation of GPP3. The dashed line represents the hydrogen bond established by GPP3 and residue Ile113 of VP1. Residue Phe135 is at the front whereas residue Phe155 is towards the back of the figure. The RMSD between the two virus structures (4AED and our complex, all atoms) is 1.1 Å, whilst the RMSD between the two GPP3 positions shown in (c) is 13.3 Å (GPP3 is docked upside down) and that between the two GPP3 positions in shown in (d) is 1.8 Å. (e) Docking of NLD by QMPLD into the 2.7 Å crystal structure reported by Plevka et al. 2013 (reference 15 in main text, PDBID:3ZFE) with the experimentally observed conformation. In cyan the sphingosine molecule found in the 3ZFE crystal structure and in grey NLD molecule docked by QMPLD. The dashed line represents the hydrogen bond established by GPP3 and residue Ile113 of VP1. (f) In magenta the NLD molecule found in the crystal structure and in grey NLD docked by QMPLD. The dashed line is the hydrogen bond between NLD and Ile113 and Gln202 of VP1. Phe135 is towards the front and Phe155 towards the back of the figure. Residue Ile24 of VP3 is coloured orange. The RMSD between the two virus structures (all atoms) is 0.8 Å, whilst the RMSD between the two NLD positions is 1.6 Å.
Supplementary Figure 5 Mass spectrum of ALD and NLD compounds.
(a) Upper panel, experimental peaks with the associated molecular mass values. The main species in the sample has a measured molecular mass of 476.2266 m/z corresponding to the introduction of an amide group on the pyridine moiety of 3-(4-pyridyl)-2-imidazolidinone. The lower panel shows the theoretical molecular mass for this compound. (b) Upper panel shows the experimental peaks with the associated molecular mass values. The main specie in the sample has a measured molecular mass of 448.2311 m/z corresponding to the introduction of an amine group on the pyridine moiety of 3-(4-pyridyl)-2-imidazolidinone. The lower panel shows the theoretical molecular mass for this compound.
Supplementary Figure 6 Structural comparison of key residues involved in forming a hydrophobic trap for pocket binders.
Structurally equivalent residues within VP1 pocket of EV71 (blue), Poliovirus type 2 (yellow, PDBID:1EAH) and rhinovirus 14 (orange, PDBID:1NA1). These residues are critical for positioning the compounds within the pocket.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3 and Supplementary Note (PDF 6136 kb)
Rights and permissions
About this article
Cite this article
De Colibus, L., Wang, X., Spyrou, J. et al. More-powerful virus inhibitors from structure-based analysis of HEV71 capsid-binding molecules. Nat Struct Mol Biol 21, 282–288 (2014). https://doi.org/10.1038/nsmb.2769
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2769
This article is cited by
-
Insights into enterovirus a-71 antiviral development: from natural sources to synthetic nanoparticles
Archives of Microbiology (2023)
-
Antivirals blocking entry of enteroviruses and therapeutic potential
Journal of Biomedical Science (2021)
-
Glutathione facilitates enterovirus assembly by binding at a druggable pocket
Communications Biology (2020)
-
Genetic characterization of VP1 of coxsackieviruses A2, A4, and A10 associated with hand, foot, and mouth disease in Vietnam in 2012–2017: endemic circulation and emergence of new HFMD-causing lineages
Archives of Virology (2020)
-
Evaluation of the virucidal effects of rosmarinic acid against enterovirus 71 infection via in vitro and in vivo study
Virology Journal (2019)