Key Points
-
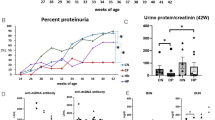
In systemic lupus erythematous (SLE), CD4+ T cells have a hypermetabolic state dominated by oxidation, mitochondrial abnormalities, activation of mTORC1 and increased glucose flux
-
Targeting T cell metabolism has therapeutic effects in mouse models of lupus and in the T cells of patients with SLE
-
Cell-specific metabolic imbalances probably also affect other immune cells in SLE, including neutrophils, plasma cells and macrophages, and specific metabolic targeting of these cells could have therapeutic benefit
-
A better understanding of the complexities of immunometabolism in SLE could lead to personalized therapeutic options
-
The metabolome, potentially intersecting with the microbiota, might provide biomarkers for SLE
Abstract
Systemic lupus erythematosus (SLE) is an autoimmune disease mediated by pathogenic autoantibodies directed against nucleoprotein complexes. Beyond the activation of autoreactive B cells, this process involves dysregulation in many other types of immune cells, including CD4+ T cells, dendritic cells, macrophages and neutrophils. Metabolic substrate utilization and integration of cues from energy sensors are critical checkpoints of effector functions in the immune system, with common as well as cell-specific programmes. Patients with SLE and lupus-prone mice present with activated metabolism of CD4+ T cells, and the use of metabolic inhibitors to normalize these features is associated with therapeutic effects. Far less is known about the metabolic requirements of B cells and myeloid cells in SLE. This article reviews current knowledge of the alterations in metabolism of immune cells in patients with SLE and mouse models of lupus in the context of what is known about the metabolic regulation of these cells during normal immune responses. How these alterations might contribute to lupus pathogenesis and how they can be targeted therapeutically are also discussed.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Liu, Z. & Davidson, A. Taming lupus — a new understanding of pathogenesis is leading to clinical advances. Nat. Med. 18, 871–882 (2012).
Sang, A., Yin, Y., Zheng, Y.-Y. & Morel, L. in Progress in Molecular Biology and Translational Science Vol. 105 (ed. Conn, P. M.) 321–370 (Academic Press, 2012).
Gergely, P. et al. Persistent mitochondrial hyperpolarization, increased reactive oxygen intermediate production, and cytoplasmic alkalinization characterize altered IL-10 signaling in patients with systemic lupus erythematosus. J. Immunol. 169, 1092–1101 (2002).
Fernandez, D., Bonilla, E., Mirza, N., Niland, B. & Perl, A. Rapamycin reduces disease activity and normalizes T cell activation-induced calcium fluxing in patients with systemic lupus erythematosus. Arthritis Rheum. 54, 2983–2988 (2006).
Lai, Z. W. et al. N-Acetylcysteine reduces disease activity by blocking mammalian target of rapamycin in T cells from systemic lupus erythematosus patients: a randomized, double-blind, placebo-controlled trial. Arthritis Rheum. 64, 2937–2946 (2012).
Frauwirth, K. A. et al. The CD28 signaling pathway regulates glucose metabolism. Immunity 16, 769–777 (2002).
Moulton, V. R. & Tsokos, G. C. T cell signaling abnormalities contribute to aberrant immune cell function and autoimmunity. J. Clin. Invest. 125, 2220–2227 (2015).
Fernandez, D. & Perl, A. Metabolic control of T cell activation and death in SLE. Autoimmun. Rev. 8, 184–189 (2009).
Choi, S. C., Titov, A. A., Sivakumar, R., Li, W. & Morel, L. Immune metabolism in systemic lupus erythematosus. Curr. Rheumatol. Rep. 18, 66 (2016).
Li, W., Sivakumar, R., Titov, A. A., Choi, S. C. & Morel, L. Metabolic factors that contribute to lupus pathogenesis. Crit. Rev. Immunol. 36, 75–98 (2016).
Gergely, P. et al. Mitochondrial hyperpolarization and ATP depletion in patients with systemic lupus erythematosus. Arthritis Rheum. 46, 175–190 (2002).
Perl, A., Gergely, P. Jr & Banki, K. Mitochondrial dysfunction in T cells of patients with systemic lupus erythematosus. Int. Rev. Immunol. 23, 293–313 (2004).
Caza, T. N., Talaber, G. & Perl, A. Metabolic regulation of organelle homeostasis in lupus T cells. Clin. Immunol. 144, 200–213 (2012).
Buck, M. D. et al. Mitochondrial dynamics controls T cell fate through metabolic programming. Cell 166, 63–76 (2016).
Doherty, E., Oaks, Z. & Perl, A. Increased mitochondrial electron transport chain activity at complex I is regulated by N-acetylcysteine in lymphocytes of patients with systemic lupus erythematosus. Antioxid. Redox Signal. 21, 56–65 (2014).
Perl, A., Hanczko, R., Telarico, T., Oaks, Z. & Landas, S. Oxidative stress, inflammation and carcinogenesis are controlled through the pentose phosphate pathway by transaldolase. Trends Mol. Med. 17, 395–403 (2011).
Perl, A. Oxidative stress in the pathology and treatment of systemic lupus erythematosus. Nat. Rev. Rheumatol. 9, 674–686 (2013).
Tsokos, G. C. Systemic lupus erythematosus. N. Engl. J. Med. 365, 2110–2121 (2011).
Perry, D. J. et al. Murine lupus susceptibility locus Sle1c2 mediates CD4+ T cell activation and maps to estrogen-related receptor gamma. J. Immunol. 189, 793–803 (2012).
Huss, J. M., Garbacz, W. G. & Xie, W. Constitutive activities of estrogen-related receptors: transcriptional regulation of metabolism by the ERR pathways in health and disease. Biochim. Biophys. Acta 1852, 1912–1927 (2015).
Vyshkina, T. et al. Association of common mitochondrial DNA variants with multiple sclerosis and systemic lupus erythematosus. Clin. Immunol. 129, 31–35 (2008).
Yu, X. et al. Association of UCP2 -866 G/A polymorphism with chronic inflammatory diseases. Genes Immun. 10, 601–605 (2009).
Yang, Z. et al. Restoring oxidant signaling suppresses proarthritogenic T cell effector functions in rheumatoid arthritis. Sci. Transl Med. 8, 331ra38 (2016).
Powell, J. D., Pollizzi, K. N., Heikamp, E. B. & Horton, M. R. Regulation of immune responses by mTOR. Annu. Rev. Immunol. 30, 39–68 (2012).
Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nat. Rev. Immunol. 12, 325–338 (2012).
Perl, A. Activation of mTOR (mechanistic target of rapamycin) in rheumatic diseases. Nat. Rev. Rheumatol. 12, 169–182 (2016).
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013).
Craft, J. E. Follicular helper T cells in immunity and systemic autoimmunity. Nat. Rev. Rheumatol. 8, 337–347 (2012).
Blanco, P., Ueno, H. & Schmitt, N. T follicular helper (Tfh) cells in lupus: activation and involvement in SLE pathogenesis. Eur. J. Immunol. 46, 281–290 (2016).
Ray, J. P. et al. The interleukin-2-mTORc1 kinase axis defines the signaling, differentiation, and metabolism of T helper 1 and follicular B helper T cells. Immunity 43, 690–702 (2015).
Ramiscal, R. R. et al. Attenuation of AMPK signaling by ROQUIN promotes T follicular helper cell formation. eLife 4, e08698 (2015).
Pratama, A. et al. MicroRNA-146a regulates ICOS-ICOSL signalling to limit accumulation of T follicular helper cells and germinal centres. Nat. Commun. 6, 6436 (2015).
Zeng, H. et al. mTORC1 and mTORC2 kinase signaling and glucose metabolism drive follicular helper t cell differentiation. Immunity 45, 540–554 (2016).
Fernandez, D. & Perl, A. mTOR signaling: a central pathway to pathogenesis in systemic lupus erythematosus? Discov. Med. 9, 173–178 (2010).
Yin, Y. et al. Normalization of CD4+ T cell metabolism reverses lupus. Sci. Transl Med. 7, 274ra18 (2015).
Yin, Y. et al. Glucose oxidation is critical for CD4+ T cell activation in a mouse model of systemic lupus erythematosus. J. Immunol. 196, 80–90 (2016).
Lui, S. L. et al. Rapamycin attenuates the severity of established nephritis in lupus-prone NZB/W F1 mice. Nephrol. Dial. Transplant. 23, 2768–2776 (2008).
Lai, Z. W. et al. Mechanistic target of rapamycin activation triggers IL-4 production and necrotic death of double-negative T cells in patients with systemic lupus erythematosus. J. Immunol. 191, 2236–2246 (2013).
Kato, H. & Perl, A. Mechanistic target of rapamycin complex 1 expands Th17 and IL-4+ CD4-CD8- double-negative T cells and contracts regulatory T cells in systemic lupus erythematosus. J. Immunol. 192, 4134–4144 (2014).
Fernandez, D. R. et al. Activation of mammalian target of rapamycin controls the loss of TCRζ in lupus T cells through HRES-1/Rab4-regulated lysosomal degradation. J. Immunol. 182, 2063–2073 (2009).
Perl, A. et al. Comprehensive metabolome analyses reveal N-acetylcysteine-responsive accumulation of kynurenine in systemic lupus erythematosus: implications for activation of the mechanistic target of rapamycin. Metabolomics 11, 1157–1174 (2015).
Psarelis, S. & Nikiphorou, E. Coexistence of SLE, tuberous sclerosis and aggressive natural killer-cell leukaemia: coincidence or correlated? Lupus 26, 107–108 (2017).
Olde Bekkink, M., Ahmed-Ousenkova, Y. M., Netea, M. G., van der Velden, W. J. & Berden, J. H. Coexistence of systemic lupus erythematosus, tuberous sclerosis and aggressive natural killer-cell leukaemia: coincidence or correlated? Lupus 25, 766–771 (2016).
Carrasco Cubero, C., Bejarano Moguel, V., Fernandez Gil, M. A. & Alvarez Vega, J. L. Coincidence of tuberous sclerosis and systemic lupus erythematosus-a case report. Reumatol. Clin. 12, 219–222 (2016).
Singh, N., Birkenbach, M., Caza, T., Perl, A. & Cohen, P. L. Tuberous sclerosis and fulminant lupus in a young woman. J. Clin. Rheumatol. 19, 134–137 (2013).
Wahl, D. R. et al. Characterization of the metabolic phenotype of chronically activated lymphocytes. Lupus 19, 1492–1501 (2010).
Dimeloe, S. et al. The immune-metabolic basis of effector memory CD4+ T cell function under hypoxic conditions. J. Immunol. 196, 106–114 (2016).
Sobel, E. S. et al. Defective response of CD4+ T cells to retinoic acid and TGFβ in systemic lupus erythematosus. Arthritis Res. Ther. 13, R106 (2011).
Morel, L. et al. Genetic reconstitution of systemic lupus erythematosus immunopathology with polycongenic murine strains. Proc. Natl Acad. Sci. USA 97, 6670–6675 (2000).
Macintyre, A. N. et al. The glucose transporter Glut1 is selectively essential for CD4 T cell activation and effector function. Cell Metab. 20, 61–72 (2014).
Jacobs, S. R. et al. Glucose uptake is limiting in T cell activation and requires CD28-mediated Akt-dependent and independent pathways. J. Immunol. 180, 4476–4486 (2008).
Yang, Z. C. & Liu, Y. Hypoxia-inducible factor-1α and autoimmune lupus, arthritis. Inflammation 39, 1268–1273 (2016).
Le Buanec, H. et al. IFN-α and CD46 stimulation are associated with active lupus and skew natural T regulatory cell differentiation to type 1 regulatory T (Tr1) cells. Proc. Natl Acad. Sci. USA 108, 18995–19000 (2011).
Kolev, M. et al. Complement regulates nutrient influx and metabolic reprogramming during Th1 cell responses. Immunity 42, 1033–1047 (2015).
Kidani, Y. & Bensinger, S. J. Lipids rule: resetting lipid metabolism restores T cell function in systemic lupus erythematosus. J. Clin. Invest. 124, 482–485 (2014).
Krishnan, S. et al. Alterations in lipid raft composition and dynamics contribute to abnormal T cell responses in systemic lupus erythematosus. J. Immunol. 172, 7821–7831 (2004).
Jury, E. C., Isenberg, D. A., Mauri, C. & Ehrenstein, M. R. Atorvastatin restores Lck expression and lipid raft-associated signaling in T cells from patients with systemic lupus erythematosus. J. Immunol. 177, 7416–7422 (2006).
McDonald, G. et al. Normalizing glycosphingolipids restores function in CD4+ T cells from lupus patients. J. Clin. Invest. 124, 712–724 (2014).
Deng, G. M. & Tsokos, G. C. Cholera toxin B accelerates disease progression in lupus-prone mice by promoting lipid raft aggregation. J. Immunol. 181, 4019–4026 (2008).
Yang, W. et al. Potentiating the antitumour response of CD8+ T cells by modulating cholesterol metabolism. Nature 531, 651–655 (2016).
Wang, F., Beck-Garcia, K., Zorzin, C., Schamel, W. W. A. & Davis, M. M. Inhibition of T cell receptor signaling by cholesterol sulfate, a naturally occurring derivative of membrane cholesterol. Nat. Immunol. 17, 844–850 (2016).
Swamy, M. et al. A cholesterol-based allostery model of T cell receptor phosphorylation. Immunity 44, 1091–1101 (2016).
Hu, X. et al. Sterol metabolism controls TH17 differentiation by generating endogenous RORγ agonists. Nat. Chem. Biol. 11, 141–147 (2015).
Ulivieri, C. & Baldari, C. T. Statins: from cholesterol-lowering drugs to novel immunomodulators for the treatment of Th17-mediated autoimmune diseases. Pharmacol. Res. 88, 41–52 (2014).
Waddington, K. E., Jury, E. C. & Pineda-Torra, I. Liver X receptors in immune cell function in humans. Biochem. Soc. Trans. 43, 752–757 (2015).
Jeon, J. Y. et al. Liver X receptors alpha gene (NR1H3) promoter polymorphisms are associated with systemic lupus erythematosus in Koreans. Arthritis Res. Ther. 16, R112 (2014).
Cui, G. et al. Liver X receptor (LXR) mediates negative regulation of mouse and human Th17 differentiation. J. Clin. Invest. 121, 658–670 (2011).
Richard, E. M. et al. Reducing FLI1 levels in the MRL/lpr lupus mouse model impacts T cell function by modulating glycosphingolipid metabolism. PLoS ONE 8, e75175 (2013).
Sundararaj, K. P. et al. FLI1 levels impact CXCR3 expression and renal infiltration of T cells and renal glycosphingolipid metabolism in the MRL/lpr lupus mouse strain. J. Immunol. 195, 5551–5560 (2015).
Morris, E. E. et al. A GA microsatellite in the Fli1 promoter modulates gene expression and is associated with systemic lupus erythematosus patients without nephritis. Arthritis Res. Ther. 12, R212 (2010).
Nowling, T. K. et al. Renal glycosphingolipid metabolism is dysfunctional in lupus nephritis. J. Am. Soc. Nephrol. 26, 1402–1413 (2015).
Murray, P. J., Rathmell, J. & Pearce, E. SnapShot: immunometabolism. Cell Metab. 22, 190–190.e1 (2015).
Caro-Maldonado, A. et al. Metabolic reprogramming is required for antibody production that is suppressed in anergic but exaggerated in chronically BAFF-exposed B cells. J. Immunol. 192, 3626–3636 (2014).
Aronov, M. & Tirosh, B. Metabolic control of plasma cell differentiation — what we know and what we don't know. J. Clin. Immunol. 36 (Suppl. 1), 12–17 (2016).
Benhamron, S., Pattanayak, S. P., Berger, M. & Tirosh, B. mTOR activation promotes plasma cell differentiation and bypasses XBP-1 for immunoglobulin secretion. Mol. Cell. Biol. 35, 153–166 (2015).
Wu, T. et al. Shared signaling networks active in B cells isolated from genetically distinct mouse models of lupus. J. Clin. Invest. 117, 2186–2196 (2007).
Zeng, Q. et al. Rapamycin inhibits BAFF-stimulated cell proliferation and survival by suppressing mTOR-mediated PP2A-Erk1/2 signaling pathway in normal and neoplastic B-lymphoid cells. Cell. Mol. Life Sci. 72, 4867–4884 (2015).
Lam, W. Y. et al. Mitochondrial pyruvate import promotes long-term survival of antibody-secreting plasma cells. Immunity 45, 60–73 (2016).
Pathak, S. et al. Fatty acid amide hydrolase regulates peripheral B cell receptor revision, polyreactivity, and B1 cells in lupus. J. Immunol. 196, 1507–1516 (2016).
Lugar, P. L., Love, C., Grammer, A. C., Dave, S. S. & Lipsky, P. E. Molecular characterization of circulating plasma cells in patients with active systemic lupus erythematosus. PLoS ONE 7, e44362 (2012).
Aprahamian, T. et al. The peroxisome proliferator-activated receptor γ agonist rosiglitazone ameliorates murine lupus by induction of adiponectin. J. Immunol. 182, 340–346 (2009).
Aprahamian, T. R., Bonegio, R. G., Weitzner, Z., Gharakhanian, R. & Rifkin, I. R. Peroxisome proliferator-activated receptor gamma agonists in the prevention and treatment of murine systemic lupus erythematosus. Immunology 142, 363–373 (2014).
Venegas-Pont, M. et al. Rosiglitazone decreases blood pressure and renal injury in a female mouse model of systemic lupus erythematosus. Am. J. Physiol. Regul. Integr. Comp. Physiol. 296, R1282–R1289 (2009).
Zhao, W. et al. The peroxisome proliferator-activated receptor gamma agonist pioglitazone improves cardiometabolic risk and renal inflammation in murine lupus. J. Immunol. 183, 2729–2740 (2009).
O'Neill, L. A. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23 (2016).
Ravishankar, B. et al. Tolerance to apoptotic cells is regulated by indoleamine 2,3-dioxygenase. Proc. Natl Acad. Sci. USA 109, 3909–3914 (2012).
Ravishankar, B. et al. The amino acid sensor GCN2 inhibits inflammatory responses to apoptotic cells promoting tolerance and suppressing systemic autoimmunity. Proc. Natl Acad. Sci. USA 112, 10774–10779 (2015).
Tsalikis, J., Croitoru, D. O., Philpott, D. J. & Girardin, S. E. Nutrient sensing and metabolic stress pathways in innate immunity. Cell. Microbiol. 15, 1632–1641 (2013).
McGaha, T. L. IDO-GCN2 and autophagy in inflammation. Oncotarget 6, 21771–21772 (2015).
Eleftheriadis, T. et al. Differential effects of the two amino acid sensing systems, the GCN2 kinase and the mTOR complex 1, on primary human alloreactive CD4+ T-cells. Int. J. Mol. Med. 37, 1412–1420 (2016).
Sukhbaatar, N., Hengstschlager, M. & Weichhart, T. mTOR-mediated regulation of dendritic cell differentiation and function. Trends Immunol. 37, 778–789 (2016).
Wang, Y. et al. Tuberous sclerosis 1 (Tsc1)-dependent metabolic checkpoint controls development of dendritic cells. Proc. Natl Acad. Sci. USA 110, E4894–E4903 (2013).
Wu, D. et al. Type 1 interferons induce changes in core metabolism that are critical for immune function. Immunity 44, 1325–1336 (2016).
Berod, L. et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat. Med. 20, 1327–1333 (2014).
Smith, C. K. & Kaplan, M. J. The role of neutrophils in the pathogenesis of systemic lupus erythematosus. Curr. Opin. Rheumatol. 27, 448–453 (2015).
Lood, C. et al. Neutrophil extracellular traps enriched in oxidized mitochondrial DNA are interferogenic and contribute to lupus-like disease. Nat. Med. 22, 146–153 (2016).
Caielli, S. et al. Oxidized mitochondrial nucleoids released by neutrophils drive type I interferon production in human lupus. J. Exp. Med. 213, 697–713 (2016).
Campbell, A. M., Kashgarian, M. & Shlomchik, M. J. NADPH oxidase inhibits the pathogenesis of systemic lupus erythematosus. Sci. Transl Med. 4, 157ra141 (2012).
Bao, Y. et al. mTOR and differential activation of mitochondria orchestrate neutrophil chemotaxis. J. Cell Biol. 210, 1153–1164 (2015).
Oaks, Z. & Perl, A. Metabolic control of the epigenome in systemic lupus erythematosus. Autoimmunity 47, 256–264 (2014).
Richardson, B. C. & Patel, D. R. Epigenetics in 2013: DNA methylation and miRNA — key roles in systemic autoimmunity. Nat. Rev. Rheumatol. 10, 72–74 (2014).
Wu, T. et al. Metabolic disturbances associated with systemic lupus erythematosus. PLoS ONE 7, e37210 (2012).
Coit, P. et al. Epigenetic reprogramming in naive CD4+ T cells favoring T cell activation and non-Th1 effector T cell immune response as an early event in lupus flares. Arthritis Rheumatol. 68, 2200–2209 (2016).
Zhao, E. et al. Cancer mediates effector T cell dysfunction by targeting microRNAs and EZH2 via glycolysis restriction. Nat. Immunol. 17, 95–103 (2016).
Regna, N. L. et al. HDAC expression and activity is upregulated in diseased lupus-prone mice. Int. Immunopharmacol. 29, 494–503 (2015).
Mishra, N., Reilly, C. M., Brown, D. R., Ruiz, P. & Gilkeson, G. S. Histone deacetylase inhibitors modulate renal disease in the MRL-lpr/lpr mouse. J. Clin. Invest. 111, 539–552 (2003).
Long, H., Yin, H., Wang, L., Gershwin, M. E. & Lu, Q. The critical role of epigenetics in systemic lupus erythematosus and autoimmunity. J. Autoimmun. 74, 118–138 (2016).
Corcoran, S. E. & O'Neill, L. A. HIF1α and metabolic reprogramming in inflammation. J. Clin. Invest. 126, 3699–3707 (2016).
Shi, L. Z. et al. HIF1α–dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
Kohler, T., Reizis, B., Johnson, R. S., Weighardt, H. & Forster, I. Influence of hypoxia-inducible factor 1α on dendritic cell differentiation and migration. Eur. J. Immunol. 42, 1226–1236 (2012).
Cho, S. H. et al. Germinal centre hypoxia and regulation of antibody qualities by a hypoxia response system. Nature 537, 234–238 (2016).
Feng, C. C. et al. Lack of association between the polymorphisms of hypoxia-inducible factor 1A (HIF1A) gene and SLE susceptibility in a Chinese population. Immunogenetics 66, 9–13 (2014).
Davidson, A. What is damaging the kidney in lupus nephritis? Nat. Rev. Rheumatol. 12, 143–153 (2016).
Bethunaickan, R. et al. Identification of stage-specific genes associated with lupus nephritis and response to remission induction in (NZB x NZW)F1 and NZM2410 mice. Arthritis Rheumatol. 66, 2246–2258 (2014).
Mashmoushi, A. K. & Oates, J. C. Lipopolysaccharide induces inducible nitric oxide synthase-dependent podocyte dysfunction via a hypoxia-inducible factor 1α and cell division control protein 42 and Ras-related C3 botulinum toxin substrate 1 pathway. Free Radic. Biol. Med. 84, 185–195 (2015).
Deng, W. et al. Hypoxia inducible factor-1 alpha promotes mesangial cell proliferation in lupus nephritis. Am. J. Nephrol. 40, 507–515 (2014).
Bengtsson, A. A. et al. Metabolic profiling of systemic lupus erythematosus and comparison with primary Sjögren's syndrome and systemic sclerosis. PLoS ONE 11, e0159384 (2016).
Lood, C. et al. Type I interferon-mediated skewing of the serotonin synthesis is associated with severe disease in systemic lupus erythematosus. PLoS ONE 10, e0125109 (2015).
Pedersen, H. K. et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535, 376–381 (2016).
Hevia, A. et al. Intestinal dysbiosis associated with systemic lupus erythematosus. mBio 5, e01548-14 (2014).
Lopez, P. et al. Th17 responses and natural IgM antibodies are related to gut microbiota composition in systemic lupus erythematosus patients. Sci. Rep. 6, 24072 (2016).
Rojo, D. et al. Ranking the impact of human health disorders on gut metabolism: systemic lupus erythematosus and obesity as study cases. Sci. Rep. 5, 8310 (2015).
Serrano-Villar, S. et al. HIV infection results in metabolic alterations in the gut microbiota different from those induced by other diseases. Sci. Rep. 6, 26192 (2016).
Keller, K. E., Tan, I. S. & Lee, Y. S. SAICAR stimulates pyruvate kinase isoform M2 and promotes cancer cell survival in glucose-limited conditions. Science 338, 1069–1072 (2012).
Mills, E. & O'Neill, L. A. Succinate: a metabolic signal in inflammation. Trends Cell Biol. 24, 313–320 (2014).
Correa-Oliveira, R., Fachi, J. L., Vieira, A., Sato, F. T. & Vinolo, M. A. Regulation of immune cell function by short-chain fatty acids. Clin. Transl Immunology 5, e73 (2016).
Pasquier, B. Autophagy inhibitors. Cell. Mol. Life Sci. 73, 985–1001 (2016).
Domhan, S. et al. Molecular mechanisms of the antiangiogenic and antitumor effects of mycophenolic acid. Mol. Cancer Ther. 7, 1656–1668 (2008).
Dun, B. Y. et al. Transcriptomic changes induced by mycophenolic acid in gastric cancer cells. Am. J. Transl Res. 6, 28–42 (2014).
He, X. et al. Mycophenolic acid-mediated suppression of human CD4+ T cells: more than mere guanine nucleotide deprivation. Am. J. Transplant. 11, 439–449 (2011).
Stylianou, K. et al. The PI3K/Akt/mTOR pathway is activated in murine lupus nephritis and downregulated by rapamycin. Nephrol. Dial. Transplant. 26, 498–508 (2011).
Rhoads, J. P., Major, A. S. & Rathmell, J. C. Fine tuning of immune metabolism for the treatment of rheumatic diseases. Nat. Rev. Rheumatol. (in press).
Fernández-Ramos, A. A., Poindessous, V., Marchetti-Laurent, C., Pallet, N. & Loriot, M.-A. The effect of immunosuppressive molecules on T-cell metabolic reprogramming. Biochimie 127, 23–36 (2016).
Tanaka, N., Kusunoki, N., Kusunoki, Y., Hasunuma, T. & Kawai, S. Resistin is associated with the inflammation process in patients with systemic autoimmune diseases undergoing glucocorticoid therapy: comparison with leptin and adiponectin. Mod. Rheumatol. 23, 8–18 (2013).
Tanaka, N., Masuoka, S., Kusunoki, N., Nanki, T. & Kawai, S. Serum resistin level and progression of atherosclerosis during glucocorticoid therapy for systemic autoimmune diseases. Metabolites 6, E28 (2016).
Mejia, P. et al. Dietary restriction protects against experimental cerebral malaria via leptin modulation and T-cell mTORC1 suppression. Nat. Commun. 6, 6050 (2015).
Zhao, W. et al. The peroxisome-proliferator activated receptor-γ agonist pioglitazone modulates aberrant T cell responses in systemic lupus erythematosus. Clin. Immunol. 149, 119–132 (2013).
Bride, K. L. et al. Sirolimus is effective in relapsed/refractory autoimmune cytopenias: results of a prospective multi-institutional trial. Blood 127, 17–28 (2016).
Oaks, Z., Winans, T., Huang, N., Banki, K. & Perl, A. Activation of the mechanistic target of rapamycin in SLE: explosion of evidence in the last five years. Curr. Rheumatol. Rep. 18, 73 (2016).
Petri, M. et al. Derivation and validation of the Systemic Lupus International Collaborating Clinics classification criteria for systemic lupus erythematosus. Arthritis Rheum. 64, 2677–2686 (2012).
Acknowledgements
The author's work is supported by Alliance for Lupus Research Target Identification in Lupus grants (TIL 85521 and TIL 75018).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Glossary
- Electron transport chain
-
A series of proteins in the inner mitochondrial membrane that transfer electrons from one to the other in a series of redox reactions, resulting in the movement of protons out of the mitochondrial matrix and in the synthesis of ATP.
- Oxidative phosphorylation
-
A metabolic pathway that produces ATP from the oxidation of acetyl-CoA and the transfer of electrons to the electron transport chain via NADH and FADH2.
- Aeorbic glycolysis
-
(Also known as the 'Warburg effect') The abrupt metabolic switch from oxidative phosphorylation to glycolysis, regardless of the availability of oxygen, to provide energy for cell proliferation and effector functions.
- Glycolysis
-
An oxygen-independent metabolic pathway that generates two molecules of pyruvate, ATP and NADH from every one molecule of glucose, supporting the tricarboxylic acid cycle and providing intermediates for the pentose phosphate pathway, glycosylation reactions and for the synthesis of biomolecules (including serine, glycine, alanine and acetyl-CoA).
- Pentose phosphate pathway (PPP)
-
An anabolic metabolic pathway parallel to glycolysis that branches out from glycolysis with the conversion of glucose-6-phosphate to ribose 5-phosphate and generates the reducing equivalents NADPH, ribose 5-phosphate (used in the synthesis of nucleotides and nucleic acids) and erythrose-4-phosphosphate (used in the synthesis of amino acids).
- Lipid rafts
-
Microdomains of the plasma membrane that are enriched in cholesterol and glycosphingolipids and serve as self-organizing centres for the assembly of signalling molecules.
- Statins
-
A class of lipid-lowering drugs that inhibit a key enzyme in the synthesis of cholesterol, HMG-CoA reductase.
- Fatty acid oxidation
-
A metabolic process that produces ATP from the oxidation of acetyl-CoA derived from the mobilization of fatty acids.
- Tricarboxylic acid (TCA) cycle
-
(Also known as the Krebs cycle) A set of connected pathways in the mitochondrial matrix, which metabolize acetyl-CoA derived from glycolysis or fatty acid oxidation, producing NADH and FADH2 for the electron transport chain and precursors for amino acid and fatty acid synthesis.
- NETosis
-
A specialized form of cell death characterized by the release of neutrophil extracellular traps (NETs), which are chromatin structures loaded with granular and nucleic proteins.
Rights and permissions
About this article
Cite this article
Morel, L. Immunometabolism in systemic lupus erythematosus. Nat Rev Rheumatol 13, 280–290 (2017). https://doi.org/10.1038/nrrheum.2017.43
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrrheum.2017.43
This article is cited by
-
The immunoregulatory roles of non-haematopoietic cells in the kidney
Nature Reviews Nephrology (2024)
-
Oxidized galectin-1 in SLE fails to bind the inhibitory receptor VSTM1 and increases reactive oxygen species levels in neutrophils
Cellular & Molecular Immunology (2023)
-
The role of platelets in immune-mediated inflammatory diseases
Nature Reviews Immunology (2023)
-
Analysis of microRNA-199a-3p expression in CD4+ T cells of systemic lupus erythematosus
Clinical Rheumatology (2023)
-
COX5A as a potential biomarker for disease activity and organ damage in lupus
Clinical and Experimental Medicine (2023)