Key Points
-
Systemic sclerosis (SSc) is a complex autoimmune disease, the pathogenesis of which is influenced by genetic, epigenetic and environmental factors
-
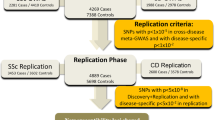
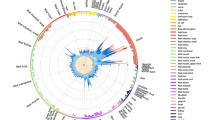
Candidate-gene studies and genome-wide association studies have identified a large number of genetic susceptibility factors for SSc and its clinical phenotypes
-
The low concordance rate for monozygotic twins demonstrates that a genetic basis cannot account exclusively for SSc pathogenesis and that epigenetic factors influenced by the environment are important
-
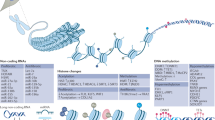
Several DNA methylation patterns, histone modifications and microRNAs (miRNAs) are altered in different cell types from patients with SSc; altered circulating miRNAs are potentially useful as disease diagnostic and prognostic biomarkers
Abstract
Systemic sclerosis (SSc) is a complex autoimmune disease of unclear aetiology. A multitude of genetic studies, ranging from candidate-gene studies to genome-wide association studies, have identified a large number of genetic susceptibility factors for SSc and its clinical phenotypes, but the contribution of these factors to disease susceptibility is only modest. However, in an endeavour to explore how the environment might affect genetic susceptibility, epigenetic research into SSc is rapidly expanding. Orchestrated by environmental factors, epigenetic modifications can drive genetically predisposed individuals to develop autoimmunity, and are thought to represent the crossroads between the environment and genetics in SSc. Therefore, in addition to providing a comprehensive description of the current understanding of genetic susceptibility underlying SSc, this Review describes the involvement of epigenetic phenomena, including DNA methylation patterns, histone modifications and microRNAs, in SSc.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Varga, J. & Abraham, D. Systemic sclerosis: a prototypic multisystem fibrotic disorder. J. Clin. Invest. 117, 557–567 (2007).
Preliminary criteria for the classification of systemic sclerosis (scleroderma). Subcommittee for scleroderma criteria of the American Rheumatism Association Diagnostic and Therapeutic Criteria Committee. Arthritis Rheum. 23, 581–590 (1980).
Steen, V. D. Autoantibodies in systemic sclerosis. Semin. Arthritis Rheum. 35, 35–42 (2005).
Broen, J. C., Coenen, M. J. & Radstake, T. R. Deciphering the genetic background of systemic sclerosis. Expert Rev. Clin. Immunol. 7, 449–462 (2011).
Joseph, C. G. et al. Association of the autoimmune disease scleroderma with an immunologic response to cancer. Science 343, 152–157 (2014).
McCormic, Z. D., Khuder, S. S., Aryal, B. K., Ames, A. L. & Khuder, S. A. Occupational silica exposure as a risk factor for scleroderma: a meta-analysis. Int. Arch. Occup. Environ. Health 83, 763–769 (2010).
Hudson, M. et al. Polyautoimmunity and familial autoimmunity in systemic sclerosis. J. Autoimmun. 31, 156–159 (2008).
Englert, H. et al. Familial risk estimation in systemic sclerosis. Aust. N. Z. J. Med. 29, 36–41 (1999).
Arora-Singh, R. K. et al. Autoimmune diseases and autoantibodies in the first degree relatives of patients with systemic sclerosis. J. Autoimmun. 35, 52–57 (2010).
Arnett, F. C. et al. Familial occurrence frequencies and relative risks for systemic sclerosis (scleroderma) in three United States cohorts. Arthritis Rheum. 44, 1359–1362 (2001).
Frech, T. et al. Heritability of vasculopathy, autoimmune disease, and fibrosis in systemic sclerosis: a population-based study. Arthritis Rheum. 62, 2109–2116 (2010).
Feghali-Bostwick, C., Medsger, T. A., Jr & Wright, T. M. Analysis of systemic sclerosis in twins reveals low concordance for disease and high concordance for the presence of antinuclear antibodies. Arthritis Rheum. 48, 1956–1963 (2003).
Broen, J. C., Coenen, M. J. & Radstake, T. R. Genetics of systemic sclerosis: an update. Curr. Rheumatol. Rep. 14, 11–21 (2012).
Tsuchiya, N. et al. Association of STAT4 polymorphism with systemic sclerosis in a Japanese population. Ann. Rheum. Dis. 68, 1375–1376 (2009).
Rueda, B. et al. The STAT4 gene influences the genetic predisposition to systemic sclerosis phenotype. Hum. Mol. Genet. 18, 2071–2077 (2009).
Gourh, P. et al. Polymorphisms in TBX21 and STAT4 increase the risk of systemic sclerosis: evidence of possible gene-gene interaction and alterations in TH1/TH2 cytokines. Arthritis Rheum. 60, 3794–3806 (2009).
Dieude, P. et al. STAT4 is a genetic risk factor for systemic sclerosis having additive effects with IRF5 on disease susceptibility and related pulmonary fibrosis. Arthritis Rheum. 60, 2472–2479 (2009).
Avouac, J. et al. Inactivation of the transcription factor STAT-4 prevents inflammation-driven fibrosis in animal models of systemic sclerosis. Arthritis Rheum. 63, 800–809 (2011).
Korman, B. D., Kastner, D. L., Gregersen, P. K. & Remmers, E. F. STAT4: genetics, mechanisms, and implications for autoimmunity. Curr. Allergy Asthma Rep. 8, 398–403 (2008).
Redmond, W. L., Ruby, C. E. & Weinberg, A. D. The role of OX40-mediated co-stimulation in T-cell activation and survival. Crit. Rev. Immunol. 29, 187–201 (2009).
Bossini-Castillo, L. et al. A replication study confirms the association of TNFSF4 (OX40L) polymorphisms with systemic sclerosis in a large European cohort. Ann. Rheum. Dis. 70, 638–641 (2011).
Gourh, P. et al. Association of the C8orf13-BLK region with systemic sclerosis in North-American and European populations. J. Autoimmun. 34, 155–162 (2010).
Aliprantis, A. O. et al. Transcription factor T-bet regulates skin sclerosis through its function in innate immunity and via IL-13. Proc. Natl Acad. Sci. USA 104, 2827–2830 (2007).
Lakos, G., Melichian, D., Wu, M. & Varga, J. Increased bleomycin-induced skin fibrosis in mice lacking the TH1-specific transcription factor T-bet. Pathobiology 73, 224–237 (2006).
Guarda, G. et al. T cells dampen innate immune responses through inhibition of NLRP1 and NLRP3 inflammasomes. Nature 460, 269–273 (2009).
Dieude, P. et al. NLRP1 influences the systemic sclerosis phenotype: a new clue for the contribution of innate immunity in systemic sclerosis-related fibrosing alveolitis pathogenesis. Ann. Rheum. Dis. 70, 668–674 (2011).
Orozco, G. et al. Study of the common genetic background for rheumatoid arthritis and systemic lupus erythematosus. Ann. Rheum. Dis. 70, 463–468 (2011).
Ito, I. et al. Association of the FAM167A-BLK region with systemic sclerosis. Arthritis Rheum. 62, 890–895 (2010).
Dymecki, S. M., Niederhuber, J. E. & Desiderio, S. V. Specific expression of a tyrosine kinase gene, blk, in B lymphoid cells. Science 247, 332–336 (1990).
Yokoyama, K. et al. BANK regulates BCR-induced calcium mobilization by promoting tyrosine phosphorylation of IP(3) receptor. EMBO J. 21, 83–92 (2002).
Gourh, P. et al. Association of the C8orf13-BLK region with systemic sclerosis in North-American and European populations. J. Autoimmun. 34, 155–162 (2010).
Kozyrev, S. V. et al. Functional variants in the B-cell gene BANK1 are associated with systemic lupus erythematosus. Nat. Genet. 40, 211–216 (2008).
Orozco, G. et al. Study of functional variants of the BANK1 gene in rheumatoid arthritis. Arthritis Rheum. 60, 372–379 (2009).
Dieude, P. et al. BANK1 is a genetic risk factor for diffuse cutaneous systemic sclerosis and has additive effects with IRF5 and STAT4. Arthritis Rheum. 60, 3447–3454 (2009).
Rueda, B. et al. BANK1 functional variants are associated with susceptibility to diffuse systemic sclerosis in Caucasians. Ann. Rheum. Dis. 69, 700–705 (2010).
Carmona, F. D. et al. New insight on the Xq28 association with systemic sclerosis. Ann. Rheum. Dis. 72, 2032–2038 (2013).
Dieudé, P. et al. Evidence of the contribution of the X chromosome to systemic sclerosis susceptibility: association with the functional IRAK1 196Phe/532Ser haplotype. Arthritis Rheum. 63, 3979–3987 (2011).
Zhou, X. et al. HLA-DPB1 and DPB2 are genetic loci for systemic sclerosis: a genome-wide association study in Koreans with replication in North Americans. Arthritis Rheum. 60, 3807–3814 (2009).
Radstake, T. R. et al. Genome-wide association study of systemic sclerosis identifies CD247 as a new susceptibility locus. Nat. Genet. 42, 426–429 (2010).
Dieude, P. et al. Independent replication establishes the CD247 gene as a genetic systemic sclerosis susceptibility factor. Ann. Rheum. Dis. 70, 1695–1696 (2011).
Allanore, Y. et al. Genome-wide scan identifies TNIP1, PSORS1C1, and RHOB as novel risk loci for systemic sclerosis. PLoS Genet. 7, e1002091 (2011).
Bossini-Castillo, L. et al. Confirmation of TNIP1 but not RHOB and PSORS1C1 as systemic sclerosis risk factors in a large independent replication study. Ann. Rheum. Dis. 72, 602–607 (2013).
López-Isac, E. et al. A genome-wide association study follow-up suggests a possible role for PPARG in systemic sclerosis susceptibility. Arthritis Res. Ther. 16, R6 (2014).
Bossini-Castillo, L. et al. A GWAS follow-up study reveals the association of the IL12RB2 gene with systemic sclerosis in Caucasian populations. Hum. Mol. Genet. 21, 926–933 (2012).
Martin, J. E. et al. A systemic sclerosis and systemic lupus erythematosus pan-meta-GWAS reveals new shared susceptibility loci. Hum. Mol. Genet. 22, 4021–4029 (2013).
Mayes, M. D. et al. Immunochip analysis identifies multiple susceptibility loci for systemic sclerosis. Am. J. Hum. Genet. 94, 47–61 (2014).
Gorlova, O. et al. Identification of novel genetic markers associated with clinical phenotypes of systemic sclerosis through a genome-wide association strategy. PLoS Genet. 7, e1002178 (2011).
Katsumoto, T. R., Whitfield, M. L. & Connolly, M. K. The pathogenesis of systemic sclerosis. Annu. Rev. Pathol. 6, 509–537 (2011).
Marie, I. et al. Prospective study to evaluate the association between systemic sclerosis and occupational exposure and review of the literature. Autoimmun. Rev. 13, 151–156 (2014).
Mora, G. F. Systemic sclerosis: environmental factors. J. Rheumatol. 36, 2383–2396 (2009).
Ballestar, E. Epigenetic alterations in autoimmune rheumatic diseases. Nat. Rev. Rheumatol. 7, 263–271 (2011).
Meda, F., Folci, M., Baccarelli, A. & Selmi, C. The epigenetics of autoimmunity. Cell Mol. Immunol. 8, 226–236 (2011).
Lei, W. et al. Abnormal DNA methylation in CD4+ T cells from patients with systemic lupus erythematosus, systemic sclerosis, and dermatomyositis. Scand. J. Rheumatol. 38, 369–374 (2009).
Jiang, H. et al. Demethylation of TNFSF7 contributes to CD70 overexpression in CD4+ T cells from patients with systemic sclerosis. Clin. Immunol. 143, 39–44 (2012).
Komura, K. et al. Blockade of CD40/CD40 ligand interactions attenuates skin fibrosis and autoimmunity in the tight-skin mouse. Ann. Rheum. Dis. 67, 867–872 (2008).
Lian, X. et al. DNA demethylation of CD40L in CD4+ T cells from women with systemic sclerosis: a possible explanation for female susceptibility. Arthritis Rheum. 64, 2338–2345 (2012).
Selmi, C. et al. X chromosome gene methylation in peripheral lymphocytes from monozygotic twins discordant for scleroderma. Clin. Exp. Immunol. 169, 253–262 (2012).
Wang, Y. Y. et al. DNA hypermethylation of the Forkhead box protein 3 (FOXP3) promoter in CD4+ T cells of patients with systemic sclerosis. Br. J. Dermatol. http://dx.doi.org/10.1111/bjd.12913.
Wang, Y., Fan, P. S. & Kahaleh, B. Association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene in scleroderma fibroblasts. Arthritis Rheum. 54, 2271–2279 (2006).
Dees, C. et al. The Wnt antagonists DKK1 and SFRP1 are downregulated by promoter hypermethylation in systemic sclerosis. Ann. Rheum. Dis. 73, 1232–1239 (2013).
Wang, Y. & Kahaleh, B. Epigenetic repression of bone morphogenetic protein receptor II expression in scleroderma. J. Cell Mol. Med. 17, 1291–1299 (2013).
Huber, L. C. et al. Trichostatin A prevents the accumulation of extracellular matrix in a mouse model of bleomycin-induced skin fibrosis. Arthritis Rheum. 56, 2755–2764 (2007).
Hemmatazad, H. et al. Histone deacetylase 7, a potential target for the antifibrotic treatment of systemic sclerosis. Arthritis Rheum. 60, 1519–1529 (2009).
Ghosh, A. K. et al. p300 is elevated in systemic sclerosis and its expression is positively regulated by TGF-β: epigenetic feed-forward amplification of fibrosis. J. Invest. Dermatol. 133, 1302–1310 (2013).
Bhattacharyya, S. et al. Fibroblast expression of the coactivator p300 governs the intensity of profibrotic response to transforming growth factor β. Arthritis Rheum. 52, 1248–1258 (2005).
Ghosh, A. K., Yuan, W., Mori, Y. & Varga, J. Smad-dependent stimulation of type I collagen gene expression in human skin fibroblasts by TGF-β involves functional cooperation with p300/CBP transcriptional coactivators. Oncogene 19, 3546–3555 (2000).
Bhattacharyya, S. et al. Egr-1 induces a profibrotic injury/repair gene program associated with systemic sclerosis. PLoS ONE 6, e23082 (2011).
Kramer, M. et al. Inhibition of H3K27 histone trimethylation activates fibroblasts and induces fibrosis. Ann. Rheum. Dis. 72, 614–620 (2013).
Wang, Y. et al. Aberrant histone modification in peripheral blood B cells from patients with systemic sclerosis. Clin. Immunol. 149, 46–54 (2013).
Li, H. et al. MicroRNA array analysis of microRNAs related to systemic scleroderma. Rheumatol. Int. 32, 307–313 (2012).
Zhu, H. et al. MicroRNA expression abnormalities in limited cutaneous scleroderma and diffuse cutaneous scleroderma. J. Clin. Immunol. 32, 514–522 (2012).
Honda, N. et al. TGF-β-mediated downregulation of microRNA-196a contributes to the constitutive upregulated type I collagen expression in scleroderma dermal fibroblasts. J. Immunol. 188, 3323–3331 (2012).
Honda, N. et al. miR-150 down-regulation contributes to the constitutive type I collagen overexpression in scleroderma dermal fibroblasts via the induction of integrin β3. Am. J. Pathol. 182, 206–216 (2013).
Zhu, H. et al. MicroRNA-21 in scleroderma fibrosis and its function in TGF-β-regulated fibrosis-related genes expression. J. Clin. Immunol. 33, 1100–1109 (2013).
Vettori, S., Gay, S. & Distler, O. Role of MicroRNAs in fibrosis. Open. Rheumatol. J. 6, 130–139 (2012).
Jiang, X., Tsitsiou, E., Herrick, S. E. & Lindsay, M. A. MicroRNAs and the regulation of fibrosis. FEBS J. 277, 2015–2021 (2010).
Peng, W. J. et al. MicroRNA-29: a potential therapeutic target for systemic sclerosis. Expert Opin. Ther. Targets. 16, 875–879 (2012).
Maurer, B. et al. MicroRNA-29, a key regulator of collagen expression in systemic sclerosis. Arthritis Rheum. 62, 1733–1743 (2010).
Bhattacharyya, S. et al. Toll-like receptor 4 signaling augments transforming growth factor-β responses: a novel mechanism for maintaining and amplifying fibrosis in scleroderma. Am. J. Pathol. 182, 192–205 (2013).
Makino, K. et al. The downregulation of microRNA let-7a contributes to the excessive expression of type I collagen in systemic and localized scleroderma. J. Immunol. 190, 3905–3915 (2013).
Makino, K. et al. Discoidin domain receptor 2-microRNA 196a-mediated negative feedback against excess type I collagen expression is impaired in scleroderma dermal fibroblasts. J. Invest Dermatol. 133, 110–119 (2013).
Wang, Z. et al. Detection of hair-microRNAs as the novel potent biomarker: evaluation of the usefulness for the diagnosis of scleroderma. J. Dermatol. Sci. 72, 134–141 (2013).
Nakashima, T. et al. Impaired IL-17 signaling pathway contributes to the increased collagen expression in scleroderma fibroblasts. J. Immunol. 188, 3573–3583 (2012).
Tanaka, S. et al. Alteration of circulating miRNAs in SSc: miR-30b regulates the expression of PDGF receptor β. Rheumatology (Oxford) 52, 1963–1972 (2013).
Yamakage, A., Kikuchi, K., Smith, E. A., LeRoy, E. C. & Trojanowska, M. Selective upregulation of platelet-derived growth factor α receptors by transforming growth factor β in scleroderma fibroblasts. J. Exp. Med. 175, 1227–1234 (1992).
Sing, T. et al. microRNA-92a expression in the sera and dermal fibroblasts increases in patients with scleroderma. Rheumatology. (Oxford) 51, 1550–1556 (2012).
Kajihara, I. et al. Increased accumulation of extracellular thrombospondin-2 due to low degradation activity stimulates type I collagen expression in scleroderma fibroblasts. Am. J. Pathol. 180, 703–714 (2012).
Koba, S. et al. Expression analysis of multiple microRNAs in each patient with scleroderma. Exp. Dermatol. 22, 489–491 (2013).
Kawashita, Y. et al. Circulating miR-29a levels in patients with scleroderma spectrum disorder. J. Dermatol. Sci. 61, 67–69 (2011).
Makino, K. et al. Circulating miR-142-3p levels in patients with systemic sclerosis. Clin. Exp. Dermatol. 37, 34–39 (2012).
Broen, J. C. & Radstake, T. R. How birds of a feather flock together: genetics in autoimmune diseases. Expert Rev. Clin. Immunol. 7, 127–128 (2011).
Schaub, M. A., Boyle, A. P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. 22, 1748–1759 (2012).
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Dozmorov, M. G., Wren, J. D. & Alarcón-Riquelme, M. E. Epigenomic elements enriched in the promoters of autoimmunity susceptibility genes. Epigenetics 9, 276–285 (2013).
Bell, J. T. et al. DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines. Genome Biol. 12, R10 (2011).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
Saunders, M. A., Liang, H. & Li, W. H. Human polymorphism at microRNAs and microRNA target sites. Proc. Natl Acad. Sci. USA 104, 3300–3305 (2007).
Sethupathy, P. & Collins, F. S. MicroRNA target site polymorphisms and human disease. Trends Genet. 24, 489–497 (2008).
Li, S. et al. Molecular signatures of antibody responses derived from a systems biology study of five human vaccines. Nat. Immunol. 15, 195–204 (2014).
Werner, H. M., Mills, G. B. & Ram, P. T. Cancer systems biology: a peek into the future of patient care? Nat. Rev. Clin. Oncol. 11, 167–176 (2014).
Farina, A. et al. Epstein–Barr virus infection induces aberrant TLR activation pathway and fibroblast-myofibroblast conversion in scleroderma. J. Invest. Dermatol. 134, 954–964 (2014).
Acknowledgements
J.C.A.B. is sponsored by the VENI Laureate from the Dutch Association of Research (NWO) and M.R. is funded by a Marie Curie Intra-European Fellowship.
Author information
Authors and Affiliations
Contributions
J.C.A.B. and M.R. researched data for the article. All authors provided substantial contributions to discussions of its content, wrote the article and undertook review and/or editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Loci associated with SSc and its clinical phenotypes in candidate-gene studies and genome-wide association studies (DOCX 30 kb)
Rights and permissions
About this article
Cite this article
Broen, J., Radstake, T. & Rossato, M. The role of genetics and epigenetics in the pathogenesis of systemic sclerosis. Nat Rev Rheumatol 10, 671–681 (2014). https://doi.org/10.1038/nrrheum.2014.128
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrrheum.2014.128
This article is cited by
-
Epigenetic Dysregulation in Autoimmune and Inflammatory Skin Diseases
Clinical Reviews in Allergy & Immunology (2022)
-
MicroRNA-320a: an important regulator in the fibrotic process in interstitial lung disease of systemic sclerosis
Arthritis Research & Therapy (2021)
-
Insights Into the Preclinical Models of SSc
Current Treatment Options in Rheumatology (2021)
-
Therapeutic Approaches to Systemic Sclerosis: Recent Approvals and Future Candidate Therapies
Clinical Reviews in Allergy & Immunology (2021)
-
CXCL4 assembles DNA into liquid crystalline complexes to amplify TLR9-mediated interferon-α production in systemic sclerosis
Nature Communications (2019)