Abstract
The dynamic nature of microtubules allows them to search the three-dimensional space of the cell. But what are they looking for? During cellular morphogenesis, microtubules are captured at sites just under the plasma membrane, and this polarizes the microtubule array and associated organelles. Recent data indicate that the signalling pathways that are involved in regulating the different microtubule–cortical interactions are not only conserved between species, but also that they function in diverse processes.
Similar content being viewed by others
Main
Microtubules are one of the main elements of the cytoskeleton, and are essential for cell division, cell migration, vesicle transport and cell polarity. Since the 1980s, it has been clear that microtubules are highly dynamic structures (see the Perspective by Manfred Schliwa in this issue). In cells, they turn over every few minutes — or even seconds — due to an intrinsic property termed dynamic instability (Box 1).
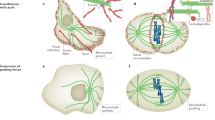
What has not been clear is why cells keep microtubules in such a highly dynamic state. GTP is hydrolysed during microtubule polymerization to allow for dynamic instability, so cells must derive some benefit from maintaining highly dynamic microtubules. In 1986, Marc Kirschner and Timothy Mitchison1 proposed that dynamic instability allows microtubules to respond rapidly to external stimuli such as growth factors or cytokines. They proposed that dynamic microtubule ends could sample the three-dimensional space of the cell to interact with — and be stabilized by — sites just under the membrane (cortical sites) that had been activated by signals (Fig. 1).
In an unpolarized cell, an external signal is thought to activate stabilizing factors (red) in the cell cortex. Dynamic microtubules that encounter these sites become stabilized. Stabilized microtubules (green) are shown to acquire post-translationally modified tubulin. This, in turn, distinguishes them from their dynamic counterparts and allows for the preferential interaction of vesicles and organelles with the stabilized microtubules leading to a polarization of the cell. Adapted from Kirschner and Mitchison1.
This model is conceptually appealing because it does not require further information to explain how microtubules might respond in a localized fashion to external signals; the dynamics of microtubules is sufficient to convey them to signal-activated targets. When this model was proposed, there were no known cases in which dynamic microtubules responded to external signals. Instead, Kirschner and Mitchison illustrated the importance of dynamic microtubules for finding targets with the capture of spindle microtubules by chromosomes during mitosis — and further evidence for chromosome capture of microtubules has accumulated2.
Evidence is beginning to emerge that microtubule searching is also important for the generation of asymmetric cell shape. For example, when microtubule antagonists are applied at nanomolar concentrations, they decrease microtubule dynamics without causing microtubule breakdown or disorganization, and inhibit processes that depend on polarized cell morphology, such as growth-cone motility3 and fibroblast migration4. Evidence has also accumulated that dynamic microtubules are selectively stabilized during cell polarization and differentiation5. However, crucial features of the model — including the identities of the external signals and the cortical targets, and how they are regulated — have remained mysterious.
Rho, Rac and Cdc42 are members of the Rho subfamily of Ras-related small GTPases, which have well-characterized roles in regulating the actin cytoskeleton, cell adhesion, gene expression and cell proliferation6,7. (There are several isoforms of both Rho and Rac in mammalian cells; unless indicated otherwise, all references to Rho and Rac refer to the well-characterized isoforms, RhoA and Rac1.) Recent studies in mammalian cells indicate that the Rho and Cdc42 GTPases also regulate signalling pathways that have distinct effects on the microtubule cytoskeleton.
Examination of the Rho and Cdc42 signalling pathways that regulate microtubules in mammalian cells shows a striking similarity to pathways that were previously identified in budding yeast — both in the proteins involved and in the microtubule–cortical interactions that the GTPases regulate. This indicates that basic mechanisms for microtubule–cortical interactions might have been evolutionarily conserved between yeast and mammalian cells. The cellular processes that are regulated by these signalling pathways in yeast and mammalian cells are different (budding and directed migration, respectively), which indicates that there are only a limited number of basic mechanisms for regulating microtubule–cortical interactions, and that these are then adapted to carry out diverse cell functions. The characteristics of the proteins that are involved in the two pathways point to a general model for the regulated interaction of microtubules with the cell cortex.
Polarized responses of microtubules
In yeast, cytoplasmic microtubules that emanate from the spindle pole body — the yeast microtubule-organizing centre (MTOC) — become polarized along the mother–bud axis during cell division (Fig. 2a). Two cortical mechanisms contribute to the alignment of microtubules along this axis. One mechanism, known as 'microtubule capture and shrinkage', involves the end-on interaction of dynamic cytoplasmic microtubules with activated targets at the bud cortex8,9,10,11,12,13,14,15. This capture results in a transient stabilization followed by shrinkage of the attached microtubule8. The second mechanism, referred to here as 'microtubule capture and sliding', involves the lateral interaction of microtubules with activated targets in the bud cortex, but results in sliding of the microtubules8,16,17,18. Both types of cortical interaction contribute to alignment of the associated nucleus along the mother–bud axis, and proper segregation of nuclei to the mother and bud. But whereas capture and shrinkage occurs early in the cell cycle, aligning the spindle along the mother–bud axis and moving the nucleus towards the bud neck, capture and sliding functions later, and is the predominant mechanism for pulling the nucleus through the bud neck16.
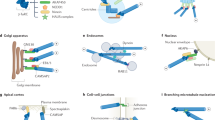
a | Microtubule capture and shrinkage, and microtubule capture and sliding in budding yeast. During cell division in budding yeast, two microtubule-capture mechanisms polarize microtubules along the mother–bud axis, and contribute to orientation of the nucleus and movements of the nucleus towards and into the bud. Microtubule capture and shrinkage (top) involves end-on capture of microtubules in the bud cortex followed by shrinkage of the captured microtubule. This aligns the spindle along the mother–bud axis and moves the nucleus to the bud neck. (Spindle pole bodies are indicated by boxes on the nucleus; spindle microtubules are not shown for clarity.) Microtubule capture and sliding (bottom) involves lateral capture of microtubules in the bud cortex, followed by sliding of the captured microtubule along the bud cortex. This pulls the nucleus through the bud neck. b | Microtubule stabilization and reorientation of the microtubule-organizing centre (MTOC) in migrating wound-edge cells. On receiving stimuli for migration, unpolarized cells (left) adopt a polarized morphology for directed cell migration (right). During the acquisition of the polarized morphology, microtubule arrays become polarized in two ways: by selectively stabilizing a subset of microtubules (top; shown in red) towards the direction of migration; and by reorienting the MTOC (bottom; shown in red) to a position between the leading edge and the nucleus.
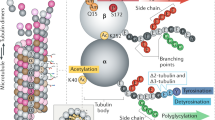
Fibroblasts migrating to in vitro wounds have been a useful model for studying microtubule polarization, as microtubules become polarized in two ways in response to wounding: by reorientation of the MTOC to a position between the nucleus and the leading edge19,20; and by the selective stabilization of microtubules in the leading lamella19,20,21,22 (Fig. 2b). Each of these contributes to the polarization of microtubules towards the leading edge, and might contribute to directed cell migration20,23. MTOC reorientation biases the bulk of the radial microtubule array towards the leading edge, whereas microtubule stabilization generates a localized subset of microtubules that accumulate post-translationally detyrosinated tubulin (Box 2), and so are biochemically distinct from their dynamic counterparts.
MTOC reorientation is accompanied by reorientation of the Golgi apparatus, and this could contribute to the biased delivery of membrane precursors to the leading edge that has been observed in wound-edge fibroblasts24. Microtubule stabilization might also bias membrane transport to the leading edge, as microtubules with elevated detyrosinated tubulin bind the microtubule motor protein kinesin better in vitro and participate in the recycling of endocytic vesicles to the cell surface25,26. As discussed below, the molecular pathways that regulate selective microtubule stabilization and MTOC reorientation in fibroblasts are similar to those that regulate microtubule polarization during budding in yeast, and are likely to involve similar microtubule–cortical interactions.
Microtubule capture and shrinkage
The pathway that mediates capture and shrinkage in yeast includes proteins called Bni1, Kar9, Myo2, Bim1/Yeb1 and Kip3 (Fig. 3). Bni1 belongs to the Diaphanous-related formin (DRF) family of adaptor proteins that are involved in cell polarity through their ability to regulate the formation of actin fibres (known as 'cables' in yeast27,28 or 'stress fibres' in mammalian cells29,30,31) and to regulate the microtubule cytoskeleton (discussed below). DRFs bind to, and are regulated by, Rho and Cdc42 GTPases. They contain conserved domains, termed formin-homology domains, that bind to other factors (such as profilin, Src and bud6) that might mediate effects on the actin and microtubule cytoskeletons27,28,29,30,31,32,33,34.
In mammalian cells, Rho GTPase stimulates the formin mDia. mDia is thought to activate a protein (possibly adenomatous polyposis coli; APC), which then interacts with dynamic microtubules bearing EB1 to cap them. In yeast, the Rho or Cdc42 GTPases stimulate the formin Bni1. Bni1 localizes Kar9 and might activate it to interact with dynamic microtubules bearing Bim1/Yeb1 to capture them. Mammalian proteins that are homologous to those in yeast are known for each step, except for Kar9. APC is proposed to be the functional homologue of Kar9 on the basis of its binding to EB1 and other criteria.
In yeast, Bni1 is localized in the bud by cell-cycle cues and localizes Kar9 to the bud tip9,10. Bni1 does not bind Kar9 directly, but polarizes actin cables towards the bud. Kar9 binds Myo2, a type-V myosin motor, and this might carry Kar9 to the bud tip on the polarized actin cables35. Kar9, in turn, interacts with the microtubule-end-binding protein Bim1/Yeb1, and can mediate the binding of Kar9 to microtubules11,12. Bim1/Yeb1 is localized to microtubule growing ends in yeast13, as is its mammalian homologue EB1 (Ref. 36), and this indicates a model in which Bim1/Yeb1 at the ends of microtubules interacts with Kar9 positioned at the bud site by Bni1. In support of this, direct imaging has shown that deletion of Bim1/Yeb1 leads to reduced capture and shrinkage8, and that Kar9 labelled with green fluorescent protein (GFP) is found at the ends of captured microtubules in the bud14.
Microtubule stabilization in fibroblasts
The pathway that regulates selective microtubule stabilization in migrating fibroblasts involves Rho, mDia and, possibly, EB1 and the adenomatous polyposis coli (APC) protein (Fig. 3). This pathway has been deciphered with serum-starved wounded 3T3 fibroblasts, which require serum or a specific serum factor, lysophosphatidic acid (LPA), to generate stable microtubules after wounding22,37. Evidence for this pathway includes measurements showing that LPA stimulates the formation of Rho·GTP in fibroblasts38, and that Rho is both necessary and sufficient for LPA-induced stabilization of microtubules in serum-starved fibroblasts37. Cdc42 and Rac are not involved (Ref. 39; A. F. Palazzo and G.G.G., unpublished observations). mDia1, which is a mouse homologue of Bni1, is the only Rho effector that is necessary for microtubule stabilization in serum-starved fibroblasts, and active forms of mDia1 or mDia2 can stimulate stable microtubule formation in the absence of LPA or active Rho, which shows that mDia functions downstream of Rho40.
Evolutionary conservation?
Given that Bni1 regulates a similar — albeit transient — microtubule stabilization in yeast, could the mammalian homologues of other proteins in the yeast capture and shrinkage pathway be involved in microtubule stabilization? Overexpressed mammalian EB1 induces selective stabilization of microtubules in serum-starved fibroblasts, and fragments of EB1 block the formation of stable microtubules that are induced by LPA or active Rho (Y. Wen, N. Cabrera-Poch and G.G.G., unpublished observations). So, EB1 might contribute to selective microtubule stabilization and could function in an analogous way to Bim1/Yeb1 in yeast microtubule capture and shrinkage.
There are no known homologues of Kar9 in mammals, Caenorhabditis elegans or Drosophila melanogaster. However, APC has been proposed as its functional homologue41. APC binds to EB1 (Ref. 42), has a region with limited homology to Kar9 (Ref. 41), and is found at the periphery of cells near microtubule ends43. Overexpressed APC in proliferating cells bundles microtubules and makes them resistant to the microtubule-depolymerizing drug nocodazole (although this might be due to its microtubule-binding domain rather than its EB1-binding domain44). In preliminary experiments, APC expression in serum-starved fibroblasts induces selective microtubule stability without bundling, and we are testing whether this is due to its EB1-binding domain. This supports the idea that APC might function in fibroblasts in an analogous way to Kar9 in yeast.
These results point to a striking similarity between the molecules that are involved in microtubule capture and shrinkage in yeast, and those that are involved in selective microtubule stabilization in mammalian cells. In addition, both pathways regulate events at the plus ends of the microtubules to prevent dynamic instability. In yeast, this has been shown directly by imaging the capture and subsequent shrinkage of microtubules at the bud cortex8. In migrating fibroblasts, selectively stabilized microtubules do not lose or gain subunits45, and the mechanism that is responsible for this behaviour has recently been shown to be capping of the normally dynamic plus ends of the microtubule46. LPA, active Rho or active mDia37,40 induce stable microtubules in fibroblasts that are capped in a similar way to those that are seen in epithelial cells, where capping was initially characterized46. Capped microtubules form within five minutes of treatment with LPA37, so they are probably formed from dynamic microtubules that are captured at cortical sites. Indeed, direct recordings of fibroblasts activated by LPA show that a subset of dynamic microtubules, which encounter the leading edge, become non-dynamic and show long states of pausing, in which no subunit addition or loss is detected37. This is analogous to capture of yeast microtubules at the bud tip. As these long-paused microtubules are likely to be precursors of capped microtubules, selective stabilization of microtubules in fibroblasts can be considered to occur by a microtubule capture and capping mechanism.
Importantly, captured microtubules have different fates in yeast and fibroblasts. In budding yeast, captured microtubules are only transiently stabilized, and subsequently shrink. In fibroblasts, by contrast, captured microtubules become capped and do not shrink for hours. We do not yet know whether captured microtubules in fibroblasts always become capped or whether they can sometimes shrink; capped microtubules in fibroblasts will shrink under certain experimental conditions in vitro46 and in vivo47. So, differences in the longevity of captured microtubules in yeast and fibroblasts might reflect differences in the degree of regulation rather than fundamental differences in the processes.
Further features of the yeast and mammalian pathways remain to be explored. For example, the Rho GTPase that regulates the activity of Bni1 in microtubule capture and shrinkage is unknown. Bni1 binds to Cdc42, Rho1, Rho3 and Rho4 in yeast6,32,33. We also need to determine the role of Bni1/mDia in capture. In yeast, Bni1 might mediate its effects on microtubule capture through actin35, but in fibroblasts, mDia binds microtubules, which indicates a more direct role40. And how do Bim1/Yeb1 and EB1 function in the capture events? EB1 is localized on the growing ends of microtubules, but it is not detected on the ends of capped microtubules in TC-7 cells46 or wounded 3T3 cells (Y. Wen and G.G.G, unpublished observations). Perhaps EB1 acts transiently by initiating capture, but not by contributing to subsequent events? There are no reports so far as to whether Bim1/Yeb1 is on the ends of captured and shrinking microtubules in budding yeast. Nonetheless, during mating in yeast, microtubules are captured at the tip of the cell extension that forms to mediate mating ('shmoo tip'). Capture at the shmoo tip is analogous to that at the bud tip; however, at the shmoo tip the captured microtubules both shrink and grow48. Moreover, Bim1/Yeb1is detectable on the ends of these captured microtubules only during periods of growth (P. Maddox and E. Salmon, unpublished observations).
Finally, the possibility that other factors participate in the capture pathways needs to be explored. Presumably, different proteins are necessary for captured microtubules to shrink (yeast budding and mating), grow (yeast mating) or become capped (fibroblasts). Genetic studies in yeast have implicated the kinesin motor Kip3 — which is localized along cytoplasmic and spindle microtubules — in capture and shrinkage15. Kip3 mutants have longer microtubules, which indicates that Kip3 might act as the 'shrinkage factor'. Kinesins can remain attached to the ends of depolymerizing microtubules in reconstitution experiments49, making them candidates to modify the activity of captured microtubules. The kinesin inhibitor AMP–PNP blocks the ATP-induced shrinking of capped microtubules in extracted fibroblasts, which indicates that the mammalian cap might also contain a kinesin-like molecule46.
Microtubule capture and sliding in yeast
The principal components in budding yeast microtubule capture and sliding are the microtubule motor protein dynein and its regulator, dynactin (Fig. 4). This pathway is less defined than the capture and shrinkage pathway, but clearly involves distinct proteins and a distinct interaction between microtubules and the cortex (Fig. 2).
In mammalian cells, the Cdc42 GTPase stimulates Par-6/protein kinase C (PKC)ζ, which then regulates the activity of dynein or dynactin to induce MTOC reorientation. Dynein might interact with dynactin localized on the ends of microtubules. In yeast, dynein and dynactin are known to be involved in microtubule attachment and sliding. On the basis of comparisons with the mammalian pathway, it is proposed that the Rho or Cdc42 GTPases regulate this pathway — perhaps by interacting with the yeast homologue of PKC, Pkc1.
Observations that support the idea of a separate pathway for microtubule capture and sliding in yeast include the fact that deletions in subunits of either dynein or dynactin are lethal in combination with deletions in members of the capture and shrinkage pathway, but deletions of more than one of the dynein or dynactin components do not increase the severity of the phenotype of single mutants8,16,17,18. Cortical microtubule sliding in the bud has been observed directly by fluorescent speckle analysis. Moreover, knockouts of Arp1 — a subunit of dynactin — reduce cortical-microtubule sliding, which links dynein and dynactin in this process8.
MTOC reorientation in fibroblasts
Studies with wounded monolayers of serum-starved fibroblasts39 and astrocytes50 show that MTOC reorientation is controlled by a pathway that involves Cdc42, dynein and dynactin (Fig. 4). MTOC reorientation in serum-starved fibroblasts requires serum and, as with microtubule capture and capping, LPA is the main serum factor that triggers it39. However, for MTOC reorientation, LPA triggers a Cdc42-regulated pathway that involves dynein and dynactin as probable downstream components39. Although the downstream Cdc42 effector in fibroblasts has not been identified, in astrocytes it seems to be the adaptor protein Par-6 (Ref. 50), a PDZ-domain-containing protein that specifically binds the GTP-bound forms of Cdc42 and Rac. Par-6 interacts with the atypical protein kinase C (PKC)-ζ, and inhibitors of PKCζ inhibit MTOC reorientation50. These studies indicate that there is a regulatory pathway for MTOC reorientation in migrating cells that is stimulated by LPA and involves Cdc42, Par-6/PKCζ, dynein and dynactin (Fig. 4). It is not yet clear whether Par-6/PKCζ directly or indirectly regulates dynein/dynactin.
Genetic studies have shown that the two pathways that regulate microtubule capture and shrinkage, and microtubule capture and sliding in yeast act independently. Palazzo et al.39 have shown that the pathways that regulate MTOC reorientation and selective microtubule stabilization in 3T3 fibroblasts also act independently — treatments that activate or inhibit one process do not activate or inhibit the other process. For example, stimulating microtubule stabilization with active Rho, or with an active form or autoinhibitory domain of mDia, does not induce MTOC reorientation39. Conversely, inhibiting MTOC reorientation with dominant-negative Cdc42, with antibodies to dynein, or by overexpression of the dynactin subunit dynamitin, does not block the formation of stabilized microtubules39. This rules out the possibility that microtubule stabilization is required for MTOC reorientation, or vice versa, and indicates that the two pathways are independently controlled in a similar manner to the two capture pathways in yeast.
A second conserved pathway?
The results discussed above point to a similarity in the molecular pathways that regulate microtubule capture and sliding in yeast and MTOC reorientation in mammalian cells. On the basis of the role of dynein and dynactin in both processes, the microtubule–cortical interaction that mediates MTOC reorientation in mammalian cells is probably analogous to that involved in capture and sliding in yeast. Indeed, sliding (or pulling) of microtubules that are captured by active dynein anchored near the leading edge is a straightforward mechanism that can account for MTOC reorientation in migrating cells. However, direct imaging has not yet been done in mammalian cells, and this will be necessary to determine whether microtubule sliding occurs during MTOC reorientation.
Questions about microtubule capture and sliding remain to be addressed in both systems. The upstream regulators in the microtubule capture and sliding pathway in yeast have not been identified. On the basis of the work in mammalian cells, it is possible that Rho GTPases regulate dynein/dynactin in yeast. Although there is no equivalent to Par-6/PKCζ in yeast, Pkc1 is a yeast PKC homologue that is regulated directly by Rho during budding51. The dynactin subunit Nip100 binds to active Rho1 and Rho2 in yeast two-hybrid screens, and this is another possibility for regulation52.
Another question is how the cortical capture that is regulated by this pathway occurs. Most models propose that cortically localized dynein pulls on microtubules. Support for this hypothesis comes from the discovery that a yeast protein called Num1 acts as a cortical anchor for dynein and functions genetically in the dynein pathway17,18. In mammalian cells, Par-6/PKCζ could have an analogous function, or it could interact with other cortical proteins that serve as the dynein anchor. For example, β-catenin has recently been shown53 to bind dynein. Localization studies in both systems should help refine the mechanisms that are involved.
A general model: TMAPs and CMAPs
The pathways that regulate microtubule capture and shrinkage/capping, and microtubule capture and sliding use different molecules and result in different types of cortical interaction. Nonetheless, a general model for the interaction of microtubules with the cortex can be proposed.
There are four basic components of the model: a Rho family GTPase; an effector of the GTPase; a cortical microtubule attachment protein (CMAP), which acts as a microtubule receptor; and a tip-only microtubule-associated protein (TMAP), which acts as a CMAP ligand (Fig. 5). The Rho-family GTPase acts as a regulatory switch and allows microtubule–cortical interactions to be initiated by external or internal signals and integrated with the actin cytoskeleton. In mammals, Rho GTPases can be switched on by soluble activators (such as LPA), and also by other molecules such as integrins. Indeed, we have found that integrin stimulation affects microtubule capture and capping (A. F. Palazzo, J. Yoon and G.G.G., unpublished observations). Integrins also seem to be involved in the MTOC-reorientation pathway in astrocytes50. In yeast, the factors that activate Rho-family GTPases are less clear, although cell-cycle and mating factors control the activation of Cdc42 (Refs 33,54). Cell-cycle regulation of microtubule capture and capping might also occur in mammalian cells; for example, during cytokinesis, where Rho GTPases are required.
This model is based on the two pathways for regulating microtubule–cortical interactions in mammalian cells and yeast. The first component is a member of the Rho family of GTPases, which functions as a switch in response to external, cell-cycle or cell-adhesion signals. The activated Rho GTPase then interacts with an effector, which might alter the activity of the effector, its location or its interaction with other proteins. The activated effector then stimulates the activity of the cortical microtubule attachment protein (CMAP). CMAPs initiate the interaction with microtubules through their ability to bind tip-only microtubule-associated proteins (TMAPs). This interaction might be modified further to generate attached microtubules that shrink, grow, remain stabilized or slide along the cortex.
Rho GTPases do not seem to regulate the activity of the CMAPs directly; instead, they act through their GTPase effectors. This might reflect the complexity of microtubule attachment to cortical sites, and the effectors that have been identified so far — Bni1/mDia and Par-6/PKCζ — all associate with the cortex and bind other proteins through interaction domains. These effectors also interact with proteins that regulate the actin cytoskeleton, which points to an integrative function and the importance of interacting with cortical actin.
The least-understood aspect of the model is the nature of the CMAPs, and how they become associated with the cortex. Kar9 and APC are candidate CMAPs for the microtubule capture and shrinkage/capping pathways, although it is not yet clear whether they act as the structural tether to microtubules. Kar9 and APC bind to the TMAP EB1, and this could mediate the initial interaction between a growing microtubule and the cortex. However, EB1 is not detectable on captured and shrinking microtubules in yeast, or on captured and capped microtubules in mammalian cells. Perhaps Kar9 (and APC) promote the interaction of microtubule ends with still-to-be-identified CMAPs that mediate the actual tethering.
Dynactin is localized on the ends of growing microtubules55, so it is a TMAP-like EB1. Dynactin that is localized at microtubule tips could stimulate cortically localized dynein and/or provide interaction sites for dynein to initiate capture and sliding of microtubules. Such a model extends an earlier idea — that the localization of dynactin at microtubule tips provides docking sites from which vesicles that contain dynein initiate their movement towards the minus end (retrograde movement)55. For example, endosomes are moved towards the centre of the cell by dynein. In the case of microtubule–cortical sliding, dynactin on the growing ends of microtubules would interact with dynein that is immobilized in the cortex rather than on mobile vesicles, and this would cause the microtubule (and attached MTOC) to move towards the active dynein. One test of this idea will be to see whether dynactin is localized on the ends of captured and sliding microtubules.
CLIP-170 (cytoplasmic linker protein p170) is another candidate TMAP. CLIP-170 was originally identified as a protein that binds to microtubules in a nucleotide-dependent fashion and was shown to be specifically localized to the growing ends of microtubules56,57. Recently, Galjart and colleagues58 identified CLASPs (CLIP-associating proteins) as a new family of CLIP-170-binding proteins. CLASPs might act as CMAPs, as they are localized near microtubule ends and are redistributed to the leading edge of wounded fibroblasts in response to serum58. Although the factors that regulate CLASPs are not yet known, CLASPs themselves seem to be involved in serum-induced microtubule stabilization. CLIP-170 also interacts with dynein and modifies its activity, which indicates that it might participate in the microtubule capture and sliding pathway. In yeast, the CLIP-170 homologue Bik1 seems to function in dynein microtubule capture and sliding59. Whether CLIP-170 and CLASPs regulate a new microtubule–cortical interaction or modify known cortical interactions will require further study.
In summary, Kirschner and Mitchison's selective stabilization model has provided a useful theoretical framework for exploring the importance of dynamic microtubules and their possible interaction with cortical sites that are regulated by signalling pathways. By identifying the molecules that mediate and regulate the cortical interactions, more recent studies have highlighted the conservation of the regulatory pathways from yeast to mammals. As the conserved pathways regulate different cellular activities in yeast and mammalian cells, it could be that there are only a few basic microtubule–cortical interactions, and that these are then modified to carry out specific cellular functions. Indeed, there are a number of other systems in which molecules that regulate microtubule capture in yeast and fibroblasts affect cellular asymmetry (Box 3; Table 1). In any case, the repertoire of cortical interactions is much richer than previously predicted in the original Kirschner and Mitchison model: interactions involve not only microtubule capture, but also distinct consequences for the captured microtubules — shrinkage, growth, capping and sliding.
The regulatory pathways that control microtubule–cortical interactions have also turned out to be more intricate than could have been predicted, and this might allow for fine tuning and integration with other cellular responses. Given the repertoire of cortical interactions and the regulatory mechanisms that control them, it is likely that the contribution of microtubules to cell polarity is more varied than previously thought.
References
Kirschner, M. & Mitchison, T. Beyond self-assembly: from microtubules to morphogenesis. Cell 45, 329–342 (1986).
Rieder, C. L. & Salmon, E. D. The vertebrate cell kinetochore and its roles during mitosis. Trends Cell Biol. 8, 310–318 (1998).
Tanaka, E., Ho, T. & Kirschner, M. W. The role of microtubule dynamics in growth cone motility and axonal growth. J. Cell Biol. 128, 139–155 (1995).
Liao, G., Nagasaki, T. & Gundersen, G. G. Low concentrations of nocodazole interfere with fibroblast locomotion without significantly affecting MT level: implications for the role of dynamic microtubules in cell locomotion. J. Cell Sci. 108, 3473–3483 (1995).
Bulinski, J. C. & Gundersen, G. G. Stabilization and post-translational modification of microtubules during cellular morphogenesis. Bioessays 13, 285–293 (1991).
Hall, A. Rho GTPases and the actin cytoskeleton. Science 279, 509–514 (1998).
Takai, Y., Sasaki, T. & Matozaki, T. Small GTP-binding proteins. Physiol. Rev. 81, 153–208 (2001).
Adames, N. R. & Cooper, J. A. Microtubule interactions with the cell cortex causing nuclear movements in Saccharomyces cerevisiae. J. Cell Biol. 149, 863–874 (2000).
Lee, L., Klee, S. K., Evangelista, M., Boone, C. & Pellman, D. Control of mitotic spindle position by the Saccharomyces cerevisiae formin Bni1p. J. Cell Biol. 144, 947–961 (1999).
Miller, R., Matheos, D. & Rose, M. The cortical location of the microtubule orientation protein, Kar9p, is dependent upon actin and proteins required for polarization. J. Cell Biol. 144, 963–975 (1999).
Korinek, W. S., Copeland, M. J., Chadhuri, A. & Chant, J. Molecular linkage underlying microtubule orientation toward cortical sites in yeast. Science 287, 2257–2259 (2000).
Lee, L. et al. Positioning of the mitotic spindle by a cortical–microtubule capture mechanism. Science 287, 2260–2262 (2000).
Tirnauer, J. S., O'Toole, E., Berrueta, L., Bierer, B. E. & Pellman, D. Yeast Bim1p promotes the G1-specific dynamics of microtubules. J. Cell Biol. 145, 993–1007 (1999).
Beach, D. L., Thibodeaux, J., Maddox, P., Yeh, E. & Bloom, K. The role of the proteins Kar9 and Myo2 in orienting the mitotic spindle of budding yeast. Curr. Biol. 10, 1497–1506 (2000).
Miller, R. K. et al. The kinesin-related proteins, Kip2p and Kip3p, function differently in nuclear migration in yeast. Mol. Biol. Cell 9, 2051–2068 (1998).
Yeh, E. et al. Dynamic positioning of mitotic spindles in yeast: role of microtubule motors and cortical determinants. Mol. Biol. Cell 11, 3949–3961 (2000).
Heil-Chapdelaine, R. A., Oberle, J. R. & Cooper, J. A. The cortical protein Num1p is essential for dynein-dependent interactions of microtubules with the cortex. J. Cell Biol. 151, 1337–1344 (2000).
Farkasovsky, M. & Kuntzel, H. Cortical Num1p interacts with the dynein intermediate chain Pac11p and cytoplasmic microtubules in budding yeast. J. Cell Biol. 152, 251–262 (2001).
Kupfer, A., Louvard, D. & Singer, S. J. Polarization of the Golgi apparatus and the microtubule-organizing center in cultured fibroblasts at the edge of an experimental wound. Proc. Natl Acad. Sci. USA 79, 2603–2607 (1982).
Gundersen, G. G. & Bulinski, J. C. Selective stabilization of microtubules oriented toward the direction of cell migration. Proc. Natl Acad. Sci. USA 85, 5946–5950 (1988).
Nagasaki, T., Chapin, C. J. & Gundersen, G. G. Distribution of detyrosinated microtubules in motile NRK fibroblasts is rapidly altered upon cell–cell contact: implications for contact inhibition of locomotion. Cell Motil. Cytoskeleton 23, 45–60 (1992).
Gundersen, G. G., Kim, I. & Chapin, C. J. Induction of stable microtubules in 3T3 fibroblasts by TGF-β and serum. J. Cell Sci. 107, 645–659 (1994).
Ueda, M., Graf, R., MacWilliams, H. K., Schliwa, M. & Euteneuer, U. Centrosome positioning and directionality of cell movements. Proc. Natl Acad. Sci. USA 94, 9674–9678 (1997).
Bergmann, J. E., Kupfer, A. & Singer, S. J. Membrane insertion at the leading edge of motile fibroblasts. Proc. Natl Acad. Sci. USA 80, 1367–1371 (1983).
Liao, G. & Gundersen, G. G. Kinesin is a candidate for cross-bridging microtubules and intermediate filaments: selective binding of kinesin to detyrosinated tubulin and vimentin. J. Biol. Chem. 273, 9797–9803 (1998).
Lin, S. X., Gundersen, G. G. & Maxfield, F. R. Export from endocytic recycling compartments to cell surface depends on stable, detyrosinated (Glu) microtubules and kinesin. Mol. Biol. Cell 13, 96–109 (2002).
Evangelista, M., Pruyne, D., Amberg, D. C., Boone, C. & Bretscher, A. Formins direct Arp2/3-independent actin filament assembly to polarize cell growth in yeast. Nature Cell Biol. 4, 32–41 (2002).
Sagot, I., Klee, S. K. & Pellman, D. Yeast formins regulate cell polarity by controlling the assembly of actin cables. Nature Cell Biol. 4, 42–50 (2002).
Tominaga, T. et al. Diaphanous-related formins bridge Rho GTPase and Src tyrosine kinase signaling. Mol. Cell 5, 13–25 (2000).
Watanabe, N., Kato, T., Fujita, A., Ishizaki, T. & Narumiya, S. Cooperation between mDia1 and ROCK in Rho-induced actin reorganization. Nature Cell Biol. 1, 136–143 (1999).
Nakano, K. et al. Distinct actions and cooperative roles of ROCK and mDia in Rho small G protein-induced reorganization of the actin cytoskeleton in Madin–Darby canine kidney cells. Mol. Biol. Cell 10, 2481–2491 (1999).
Kohno, H. et al. Bni1p implicated in cytoskeletal control is a putative target of Rho1p small GTP binding protein in Saccharomyces cerevisiae. EMBO J. 15, 6060–6068 (1996).
Evangelista, M. et al. Bni1p, a yeast formin linking cdc42p and the actin cytoskeleton during polarized morphogenesis. Science 276, 118–122 (1997).
Watanabe, N. et al. p140mDia, a mammalian homolog of Drosophila diaphanous, is a target protein for Rho small GTPase and is a ligand for profilin. EMBO J. 16, 3044–3056 (1997).
Yin, H., Pruyne, D., Huffaker, T. C. & Bretscher, A. Myosin V orientates the mitotic spindle in yeast. Nature 406, 1013–1015 (2000).
Berrueta, L. et al. The adenomatous polyposis coli-binding protein EB1 is associated with cytoplasmic and spindle microtubules. Proc. Natl Acad. Sci. USA 95, 10596–10601 (1998).
Cook, T. A., Nagasaki, T. & Gundersen, G. G. Rho guanosine triphosphatase mediates the selective stabilization of microtubules induced by lysophosphatidic acid. J. Cell Biol. 141,175–185 (1998).
Ren, X. D., Kiosses, W. B. & Schwartz, M. A. Regulation of the small GTP-binding protein Rho by cell adhesion and the cytoskeleton. EMBO J. 18, 578–585 (1999).
Palazzo, A. F. et al. Cdc42, dynein, and dynactin regulate MTOC reorientation independent of Rho-regulated microtubule stabilization. Curr. Biol. 11, 1536–1541 (2001).
Palazzo, A. F., Cook, T. A., Alberts, A. S. & Gundersen, G. G. mDia mediates Rho regulated formation and orientation of stable microtubules. Nature Cell Biol. 3, 723–729 (2001).
Bienz, M. Spindles cotton on to junctions, APC and EB1. Nature Cell Biol. 3, E67–E68 (2001).
Su, L. K. et al. APC binds to the novel protein EB1. Cancer Res. 55, 2971–2977 (1995).
Nathke, I. S., Adams, C. L., Polakis, P., Sellin, J. H. & Nelson, W. J. The adenomatous polyposis coli tumor suppressor protein localizes to plasma membrane sites involved in active cell migration. J. Cell Biol. 134, 165–179 (1996).
Zumbrunn, J., Kinoshita, K., Hyman, A. A. & Nathke, I. S. Binding of the adenomatous polyposis coli protein to microtubules increases microtubule stability and is regulated by GSK3β phosphorylation. Curr. Biol. 11, 4–9 (2001).
Webster, D. R., Gundersen, G. G., Bulinski, J. C. & Borisy, G. G. Differential turnover of tyrosinated and detyrosinated microtubules. Proc. Natl Acad. Sci. USA 84, 9040–9044 (1987).
Infante, A. S., Stein, M., Zhai, Y., Borisy, G. G. & Gundersen, G. G. Detyrosinated (Glu) microtubules are stabilized by an ATP-sensitive plus-end cap. J. Cell Sci. 113, 3907–3919 (2000).
Gurland, G. & Gundersen, G. G. Protein phosphatase inhibitors induce the selective breakdown of stable microtubules in fibroblasts and epithelial cells. Proc. Natl Acad. Sci. USA 90, 8827–8831 (1993).
Maddox, P. et al. Microtubule dynamics from mating through the first zygotic division in the budding yeast Saccharomyces cerevisiae. J. Cell Biol. 144, 977–987 (1999).
Lombillo, V. A., Syewart, R. J. & McIntosh, J. R. Minus-end-directed motion of kinesin-coated microspheres driven by microtubule depolymerization. Nature 373, 161–164 (1995).
Etienne-Manneville, S. & Hall, A. Integrin-mediated activation of Cdc42 controls cell polarity in migrating astrocytes through PKCζ. Cell 106, 489–498 (2001).
Nonaka, H. et al. A downstream target of RHO1 small GTP-binding protein is PKC1, a homolog of protein kinase C, which leads to activation of the MAP kinase cascade in Saccharomyces cerevisiae. EMBO J. 14, 5931–5938 (1995).
Drees, B. L. et al. A protein interaction map for cell polarity development. J. Cell Biol. 154, 549–571 (2001).
Lignon, L. A., Karki, S., Tokito, M. & Holzbaur, E. L. Dynein binds to β-catenin and may tether microtubules at adherins junctions. Nature Cell Biol. 3, 913–917 (2001).
Gulli, M. P. & Peter, M. Temporal and spatial regulation of Rho-type guanine-nucleotide exchange factors: the yeast perspective. Genes Dev. 15, 365–379 (2001).
Vaughan, K. T., Tynan, S. H., Faulkner, N. E., Echeverri, C. J. & Vallee, R. B. Colocalization of cytoplasmic dynein with dynactin and CLIP-170 at microtubule distal ends. J. Cell Sci. 112, 1437–1447 (1999).
Rickard, J. E. & Kreis, T. E. Binding of pp170 to microtubules is regulated by phosphorylation. J. Biol. Chem. 266, 17597–17605 (1991).
Perez, F., Diamantopoulos, G. S., Stalder, R. & Kreis, T. E. CLIP-170 highlights growing microtubule ends in vivo. Cell 96, 517–527 (1999).
Akhmanova, A. et al. Clasps are CLIP-115 and -170 associating proteins involved in the regional regulation of microtubule dynamics in motile fibroblasts. Cell 104, 923–935 (2001).
Lin, H. et al. Polyploids require Bik1 for kinetochore-microtubule attachment. J. Cell Biol. 155, 1173–1184 (2001).
Saxton, W. M. et al. Tubulin dynamics in cultured mammalian cells. J. Cell Biol. 99, 2175–2186 (1984).
Schulze, E. & Kirschner, M. Microtubule dynamics in interphase cells. J. Cell Biol. 102, 1020–1031 (1986).
Mitchison, T. & Kirschner, M. Dynamic instability of microtubule growth. Nature 312, 237–242 (1984).
Walczak, C. E., Mitchison, T. J. & Desai, A. XKCM1: a Xenopus kinesin-related protein that regulates microtubule dynamics during mitotic spindle assembly. Cell 84, 37–47 (1996).
Keating, T. J. & Borisy, G. G. Centrosomal and non-centrosomal microtubules. Biol. Cell 91, 321–329 (1999).
MacRae, T. H. Tubulin post-translational modifications — enzymes and their mechanisms of action. Eur. J. Biochem. 244, 265–278 (1997).
Idriss, H. T. Man to trypanosome: the tubulin tyrosination/detyrosination cycle revisited. Cell Motil. Cytoskeleton 45, 173–184 (2000).
Kemphues, K. PARsing embryonic polarity. Cell 101, 345–348 (2000).
Knoblich, J. A. Asymmetric cell division during development. Nature Rev. Mol. Cell Biol. 2, 11–20 (2001).
Doe, C. Q. & Bowerman, B. Asymmetric cell division: fly neuroblast meets worm zygote. Curr. Opin. Cell Biol. 13, 68–75 (2001).
Dujardin, D. L. & Vallee, R. B. Dynein at the cortex. Curr. Opin. Cell Biol. 14, 44–49 (2002).
Dikovskaya, D., Zumbrunn, J., Penman, G. A. & Nathke, I. S. The adenomatous polyposis coli protein: in the limelight out at the edge. Trends Cell Biol. 11, 378–384 (2001).
Hayles, J. & Nurse, P. A journey into space. Nature Rev. Mol. Cell Biol. 2, 647–656 (2001).
Watts, J. L. et al. par-6, a gene involved in the establishment of asymmetry in early C. elegans embryos, mediates the asymmetric localization of PAR-3. Development 122, 3133–3140 (1996).
Etemad-Moghadam, B., Guo, S. & Kemphues, K. J. Asymmetrically distributed PAR-3 protein contributes to cell polarity and spindle alignment in early C. elegans embryos. Cell 83, 743–752 (1995).
Tabuse, Y. et al. Atypical protein kinase C cooperates with PAR-3 to establish embryonic polarity in Caenorhabditis elegans. Development 125, 3607–3614 (1998).
Kay, A. J. & Hunter, C. P. CDC-42 regulates PAR protein localization and function to control cellular and embryonic polarity in C. elegans. Curr. Biol. 11, 474–481 (2001).
Gotta, M., Abraham, M. C. & Ahringer, J. CDC-42 controls early cell polarity and spindle orientation in C. elegans. Curr. Biol. 11, 482–488 (2001).
Skop, A. R. & White, J. G. The dynactin complex is required for cleavage plane specification in early Caenorhabditis elegans embryos. Curr. Biol. 8, 1110–1116 (1998).
Gonczy, P., Pichler, S., Kirkham, M. & Hyman, A. A. Cytoplasmic dynein is required for distinct aspects of MTOC positioning, including centrosome separation, in the one cell stage Caenorhabditis elegans embryo. J. Cell Biol. 147, 135–150 (1999).
Kuchinke, U., Grawe, F. & Knust, E. Control of spindle orientation in Drosophila by the Par-3-related PDZ-domain protein Bazooka. Curr. Biol. 8, 1357–1365 (1998).
Petronczki, M. & Knoblich, J. A. DmPAR-6 directs epithelial polarity and asymmetric cell division of neuroblasts in Drosophila. Nature Cell Biol. 3, 43–49 (2001).
Wodarz, A., Ramrath, A., Grimm, A. & Knust, E. Drosophila atypical protein kinase C associates with Bazooka and controls polarity of epithelia and neuroblasts. J. Cell Biol. 150, 1361–1374 (2000).
Suzuki, A. et al. Atypical protein kinase C is involved in the evolutionarily conserved par protein complex and plays a critical role in establishing epithelia-specific junctional structures. J. Cell Biol. 152, 1183–1196 (2001).
Lu, B., Roegiers, F., Jan, L. Y. & Jan, Y. N. Adherens junctions inhibit asymmetric division in the Drosophila epithelium. Nature 409, 522–525 (2001).
McCartney, B. M. et al. Drosophila APC2 and Armadillo participate in tethering mitotic spindles to cortical actin. Nature Cell Biol. 3, 933–938 (2001).
Brunner, D. & Nurse, P. CLIP-170-like tip1p spatially organizes microtubular dynamics in fission yeast. Cell 102, 696–704 (2000).
Drummond, D. R. & Cross, R. A. Dynamics of interphase microtubules in Schizosaccharomyces pombe. Curr. Biol. 10, 766–775 (2000).
Tran, P. T., Marsh, L., Doye, V., Inoue, S. & Chang, F. A mechanism for nuclear positioning in fission yeast based on microtubule pushing. J. Cell Biol. 153, 397–411 (2001).
Kupfer, A., Dennert, G. & Singer, S. J. Polarization of the Golgi apparatus and the microtubule-organizing center within cloned natural killer cells bound to their targets. Proc. Natl Acad. Sci. USA 80, 7224–7228 (1983).
Stowers, L., Yelon, D., Berg, L. J. & Chant, J. Regulation of the polarization of T cells toward antigen-presenting cells by Ras-related GTPase CDC42. Proc. Natl Acad. Sci. USA 92, 5027–5031 (1995).
Nemere, I., Kupfer, A. & Singer, S. J. Reorientation of the Golgi apparatus and the microtubule-organizing center inside macrophages subjected to a chemotactic gradient. Cell Motil. 5, 17–29 (1985).
Gotlieb, A. I., May, L. M., Subrahmanyan, L. & Kalnins, V. I. Distribution of microtubule organizing centers in migrating sheets of endothelial cells. J. Cell Biol. 91, 589–594 (1981).
Han, G. et al. The Aspergillus cytoplasmic dynein heavy chain and NUDF localize to microtubule ends and affect microtubule dynamics. Curr. Biol. 11, 719–724 (2001).
Morris, N. R. Nuclear migration. From fungi to the mammalian brain. J. Cell Biol. 148, 1097–1101 (2000).
O'Connell, C. B. & Wang, Y. L. Mammalian spindle orientation and position respond to changes in cell shape in a dynein-dependent fashion. Mol. Biol. Cell 11, 1765–1774 (2000).
Faulkner, N. E. et al. A role for the lissencephaly gene LIS1 in mitosis and cytoplasmic dynein function. Nature Cell Biol. 2, 784–791 (2000).
Busson, S., Dujardin, D., Moreau, A., Dompierre, J. & De Mey, J. R. Dynein and dynactin are localized to astral microtubules and at cortical sites in mitotic epithelial cells. Curr. Biol. 8, 541–544 (1998).
Yamamoto, A., West, R. R., McIntosh, J. R. & Hiraoka, Y. A cytoplasmic dynein heavy chain is required for oscillatory nuclear movement of meiotic prophase and efficient meiotic recombination in fission yeast. J. Cell Biol. 145, 1233–1249 (1999).
Acknowledgements
I thank R. Kessin and A. Palazzo for comments on the manuscript. During the time this article was written, G.G.G. was supported in part by a grant from the National Institutes of Health.
Author information
Authors and Affiliations
Related links
Related links
DATABASES
<i>Saccharomyces</i> Genome Database
Swiss-Prot
Encyclopedia of Life Science
FURTHER READING
Rights and permissions
About this article
Cite this article
Gundersen, G. Evolutionary conservation of microtubule-capture mechanisms. Nat Rev Mol Cell Biol 3, 296–304 (2002). https://doi.org/10.1038/nrm777
Issue Date:
DOI: https://doi.org/10.1038/nrm777
This article is cited by
-
HIV-1 capsid exploitation of the host microtubule cytoskeleton during early infection
Retrovirology (2021)
-
Critical role of lipid membranes in polarization and migration of cells: a biophysical view
Biophysical Reviews (2021)
-
Real-time analysis of quantum dot labeled single porcine epidemic diarrhea virus moving along the microtubules using single particle tracking
Scientific Reports (2019)
-
Emerging regulators of vascular smooth muscle cell migration
Journal of Muscle Research and Cell Motility (2019)
-
Tankyrase inhibition impairs directional migration and invasion of lung cancer cells by affecting microtubule dynamics and polarity signals
BMC Biology (2016)