Abstract
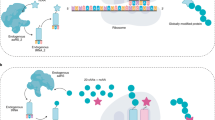
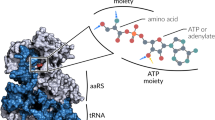
Recently, a method to encode unnatural amino acids with diverse physicochemical and biological properties genetically in bacteria, yeast and mammalian cells was developed. Over 30 unnatural amino acids have been co-translationally incorporated into proteins with high fidelity and efficiency using a unique codon and corresponding transfer-RNA:aminoacyl–tRNA-synthetase pair. This provides a powerful tool for exploring protein structure and function in vitro and in vivo, and for generating proteins with new or enhanced properties.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bock, A. et al. Selenocysteine: the 21st amino acid. Mol. Microbiol. 5, 515–520 (1991).
Srinivasan, G., James, C. M. & Krzycki, J. A. Pyrrolysine encoded by UAG in archaea: charging of a UAG-decoding specialized tRNA. Science 296, 1459–1462 (2002).
Wang, L. & Schultz, P. G. Expanding the genetic code. Angew. Chem. Int. Edn Engl. 44, 34–66 (2004).
Cornish, V. W., Mendel, D. & Schultz, P. G. Probing protein structure and function with an expanded genetic code. Angew. Chem. Int. Edn Engl. 34, 621–633 (1995).
Bain, J. D., Glabe, C. G., Dix, T. A., Chamberlin, A. R. & Diala, E. S. Biosynthetic site-specific incorporation of a non-natural amino acid into a polypeptide. J. Am. Chem. Soc. 111, 8013–8014 (1989).
Beene, D. L., Dougherty, D. A. & Lester, H. A. Unnatural amino acid mutagenesis in mapping ion channel function. Curr. Opin. Neurobiol. 13, 264–270 (2003).
Hortin, G. & Boime, I. Applications of amino acid analogs for studying co- and posttranslational modifications of proteins. Methods Enzymol. 96, 777–784 (1983).
Furter, R. Expansion of the genetic code: site-directed p-fluoro-phenylalanine incorporation in Escherichia coli. Protein Sci. 7, 419–426 (1998).
Doring, V. et al. Enlarging the amino acid set of Escherichia coli by infiltration of the valine coding pathway. Science 292, 501–504 (2001).
Kirshenbaum, K., Carrico, I. S. & Tirrell, D. A. Biosynthesis of proteins incorporating a versatile set of phenylalanine analogues. Chembiochem 3, 235–237 (2002).
Benzer, S. & Champe, S. P. A change from nonsense to sense in the genetic code. Proc. Natl Acad. Sci. USA 48, 1114–1121 (1962).
Garen, A. & Siddiqi, O. Suppression of mutations in the alkaline phosphatase structural cistron of E. coli. Proc. Natl Acad. Sci. USA 48, 1121–1127 (1962).
Wang, L., Magliery, T. J., Liu, D. R. & Schultz, P. G. A new functional suppressor tRNA/aminoacyl-tRNA synthetase pair for the in vivo incorporation of unnatural amino acids into proteins. J. Am. Chem. Soc. 122, 5010–5011 (2000).
Fechter, P., Rudinger-Thirion, J., Tukalo, M. & Giege, R. Major tyrosine identity determinants in Methanococcus jannaschii and Saccharomyces cerevisiae tRNATyr are conserved but expressed differently. Eur. J. Biochem. 268, 761–767 (2001).
Steer, B. A. & Schimmel, P. Major anticodon-binding region missing from an archaebacterial tRNA synthetase. J. Biol. Chem. 274, 35601–35606 (1999).
Jakubowski, H. & Goldman, E. Editing of errors in selection of amino acids for protein synthesis. Microbiol. Rev. 56, 412–429 (1992).
Wang, L. & Schultz, P. G. A general approach for the generation of orthogonal tRNAs. Chem. Biol. 8, 883–890 (2001).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Kobayashi, T. et al. Structural basis for orthogonal tRNA specificities of tyrosyl-tRNA synthetases for genetic code expansion. Nature Struct. Biol. 10, 425–432 (2003).
Zhang, Y., Wang, L., Schultz, P. G. & Wilson, I. A. Crystal structures of apo wild-type M. jannaschii tyrosyl-tRNA synthetase (TyrRS) and an engineered TyrRS specific for O-methyl-L-tyrosine. Protein Sci. 14, 1340–1349 (2005).
Ryu, Y. & Schultz, P. G. Efficient incorporation of unnatural amino acids into proteins in Escherichia coli. Nature Methods 3, 263–265 (2006).
Anderson, J. C. & Schultz, P. G. Adaptation of an orthogonal archaeal leucyl-tRNA and synthetase pair for four-base, amber, and opal suppression. Biochemistry 42, 9598–9608 (2003).
Anderson, J. C. et al. An expanded genetic code with a functional quadruplet codon. Proc. Natl Acad. Sci. USA 101, 7566–7571 (2004).
Kowal, A. K., Kohrer, C. & RajBhandary, U. L. Twenty-first aminoacyl-tRNA synthetase–suppressor tRNA pairs for possible use in site-specific incorporation of amino acid analogues into proteins in eukaryotes and in eubacteria. Proc. Natl Acad. Sci. USA 98, 2268–2273 (2001).
Santoro, S. W., Anderson, J. C., Lakshman, V. & Schultz, P. G. An archaebacteria-derived glutamyl-tRNA synthetase and tRNA pair for unnatural amino acid mutagenesis of proteins in Escherichia coli. Nucleic Acids Res. 31, 6700–6709 (2003).
Mehl, R. A. et al. Generation of a bacterium with a 21 amino acid genetic code. J. Am. Chem. Soc. 125, 935–939 (2003).
Chin, J. W. et al. An expanded eukaryotic genetic code. Science 301, 964–967 (2003).
Wu, N., Deiters, A., Cropp, T. A., King, D. & Schultz, P. G. A genetically encoded photocaged amino acid. J. Am. Chem. Soc. 126, 14306–14307 (2004).
Kiga, D. et al. An engineered Escherichia coli tyrosyl–tRNA synthetase for site-specific incorporation of an unnatural amino acid into proteins in eukaryotic translation and its application in a wheat germ cell-free system. Proc. Natl Acad. Sci. USA 99, 9715–9720 (2002).
Sakamoto, K. et al. Site-specific incorporation of an unnatural amino acid into proteins in mammalian cells. Nucleic Acids Res. 30, 4692–4699 (2002).
Zhang, Z. et al. Selective incorporation of 5-hydroxytryptophan into proteins in mammalian cells. Proc. Natl Acad. Sci. USA 101, 8882–8887 (2004).
Hohsaka, T., Ashizuka, Y., Taira, H., Murakami, H. & Sisido, M. Incorporation of nonnatural amino acids into proteins by using various four-base codons in an Escherichia coli in vitro translation system. Biochemistry 40, 11060–11064 (2001).
Anderson, J. C., Magliery, T. J. & Schultz, P. G. Exploring the limits of codon and anticodon size. Chem. Biol. 9, 237–244 (2002).
Feinstein, S. I. & Altman, S. Context effects on nonsense codon suppression in Escherichia coli. Genetics 88, 201–219 (1978).
Wang, L., Zhang, Z., Brock, A. & Schultz, P. G. Addition of the keto functional group to the genetic code of Escherichia coli. Proc. Natl Acad. Sci. USA 100, 56–61 (2003).
Zhang, Z. et al. A new strategy for the site-specific modification of proteins in vivo. Biochemistry 42, 6735–6746 (2003).
Chin, J. W. et al. Addition of p-azido-L-phenylalanine to the genetic code of Escherichia coli. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Deiters, A. et al. Adding amino acids with novel reactivity to the genetic code of Saccharomyces cerevisiae. J. Am. Chem. Soc. 125, 11782–11783 (2003).
Deiters, A., Cropp, T. A., Summerer, D., Mukherji, M. & Schultz, P. G. Site-specific PEGylation of proteins containing unnatural amino acids. Bioorg. Med. Chem. Lett. 14, 5743–5745 (2004).
Tsao, M. L., Tian, F. & Schultz, P. G. Selective Staudinger modification of proteins containing p-azidophenylalanine. Chembiochem 6, 2147–2149 (2005).
Farrell, I. S., Toroney, R., Hazen, J. L., Mehl, R. A. & Chin, J. W. Photo-cross-linking interacting proteins with a genetically encoded benzophenone. Nature Methods 2, 377–384 (2005).
Chin, J. W., Martin, A. B., King, D. S., Wang, L. & Schultz, P. G. Addition of a photocrosslinking amino acid to the genetic code of Escherichia coli. Proc. Natl Acad. Sci. USA 99, 11020–11024 (2002).
Chin, J. W. & Schultz, P. G. In vivo photocrosslinking with unnatural amino acid mutagenesis. Chembiochem 3, 1135–1137 (2002).
Kauer, J. C., Erickson-Viitanen, S., Wolfe, H. R. Jr & DeGrado, W. F. p-Benzoyl-L-phenylalanine, a new photoreactive amino acid. Photolabeling of calmodulin with a synthetic calmodulin-binding peptide. J. Biol. Chem. 261, 10695–10700 (1986).
Schlieker, C. et al. Substrate recognition by the AAA+ chaperone ClpB. Nature Struct. Mol. Biol. 11, 607–615 (2004).
Weibezahn, J. et al. Thermotolerance requires refolding of aggregated proteins by substrate translocation through the central pore of ClpB. Cell 119, 653–665 (2004).
Hino, N. et al. Protein photo-cross-linking in mammalian cells by site-specific incorporation of a photoreactive amino acid. Nature Methods 2, 201–206 (2005).
Deiters, A., Groff, D., Ryu, Y., Xie, J. & Schultz, P. G. A genetically encoded photocaged tyrosine. Angew. Chem. Int. Edn Engl. 45, 2728–2731 (2006).
Bartels, E., Wassermann, N. H. & Erlanger, B. F. Photochromic activators of the acetylcholine receptor. Proc. Natl Acad. Sci. USA 68, 1820–1823 (1971).
Bose, M., Groff, D., Xie, J., Brustad, E. & Schultz, P. G. The incorporation of a photoisomerizable amino acid into proteins in E. coli. J. Am. Chem. Soc. 128, 388–389 (2006).
Rudd, P. M. & Dwek, R. A. Glycosylation: heterogeneity and the 3D structure of proteins. Crit. Rev. Biochem. Mol. Biol. 32, 1–100 (1997).
Zhang, Z. et al. A new strategy for the synthesis of glycoproteins. Science 303, 371–373 (2004).
Xu, R. et al. Site-specific incorporation of the mucin-type N-acetylgalactosamine-α-O-threonine into protein in Escherichia coli. J. Am. Chem. Soc. 126, 15654–15655 (2004).
Lu, W., Gong, D., Bar-Sagi, D. & Cole, P. A. Site-specific incorporation of a phosphotyrosine mimetic reveals a role for tyrosine phosphorylation of SHP-2 in cell signaling. Mol. Cell 8, 759–769 (2001).
Xie, J. Adding Unnatural Amino Acids to the Genetic Repertoire. Thesis, Scripps Research Institute, La Jolla (2006).
Wang, J., Xie, J. & Schultz, P. G. A genetically encoded fluorescent amino acid. J. Am. Chem. Soc. 128, 8738–8739 (2006).
Summerer, D. et al. A genetically encoded fluorescent amino acid. Proc. Natl Acad. Sci. USA 103, 9785–9789 (2006).
Tsao, M. L., Summerer, D., Ryu, Y. & Schultz, P. G. The genetic incorporation of a distance probe into proteins in Escherichia coli. J. Am. Chem. Soc. 128, 4572–4573 (2006).
Xie, J. et al. The site-specific incorporation of p-iodo-L-phenylalanine into proteins for structure determination. Nature Biotechnol. 22, 1297–1301 (2004).
Deiters, A., Geierstanger, B. H. & Schultz, P. G. Site-specific in vivo labeling of proteins for NMR studies. Chembiochem 6, 55–58 (2005).
Suydam, I. T. & Boxer, S. G. Vibrational Stark effects calibrate the sensitivity of vibrational probes for electric fields in proteins. Biochemistry 42, 12050–12055 (2003).
Alfonta, L., Zhang, Z., Uryu, S., Loo, J. A. & Schultz, P. G. Site-specific incorporation of a redox-active amino acid into proteins. J. Am. Chem. Soc. 125, 14662–14663 (2003).
Wang, L., Xie, J., Deniz, A. A. & Schultz, P. G. Unnatural amino acid mutagenesis of green fluorescent protein. J. Org. Chem. 68, 174–176 (2003).
Wang, L., Brock, A. & Schultz, P. G. Adding L-3-(2-naphthyl)alanine to the genetic code of E. coli. J. Am. Chem. Soc. 124, 1836–1837 (2002).
Turner, J. M., Graziano, J., Spraggon, G. & Schultz, P. G. Structural characterization of a p-acetylphenylalanyl aminoacyl-tRNA synthetase. J. Am. Chem. Soc. 127, 14976–14977 (2005).
Turner, J. M., Graziano, J., Spraggon, G. & Schultz, P. G. Structural plasticity of an aminoacyl-tRNA synthetase active site. Proc. Natl Acad. Sci. USA 103, 6483–6488 (2006).
Tian, F., Tsao, M. L. & Schultz, P. G. A phage display system with unnatural amino acids. J. Am. Chem. Soc. 126, 15962–15963 (2004).
Castelli, D. D., Lovera, E., Ascenzi, P. & Fasano, M. Unfolding of the loggerhead sea turtle (Caretta caretta) myoglobin: a 1H-NMR and electronic absorbance study. Protein Sci. 11, 2273–2278 (2002).
Acknowledgements
Work in the laboratory of P.G.S. is supported by the National Institutes of Health, the United States Department of Energy and the Skaggs Institute for Chemical Biology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Xie, J., Schultz, P. A chemical toolkit for proteins — an expanded genetic code. Nat Rev Mol Cell Biol 7, 775–782 (2006). https://doi.org/10.1038/nrm2005
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm2005
This article is cited by
-
Synthesis and applications of mirror-image proteins
Nature Reviews Chemistry (2023)
-
Ribosome-mediated biosynthesis of pyridazinone oligomers in vitro
Nature Communications (2022)
-
Light control of the peptide-loading complex synchronizes antigen translocation and MHC I trafficking
Communications Biology (2021)
-
N-Terminal selective modification of peptides and proteins using 2-ethynylbenzaldehydes
Communications Chemistry (2020)