Abstract
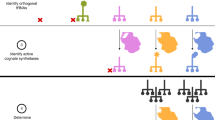
Natural organisms use a four-letter genetic alphabet that makes available 64 triplet codons, of which 61 are sense codons used to encode proteins with the 20 canonical amino acids. We have shown that the unnatural nucleotides dNaM and dTPT3 can pair to form an unnatural base pair (UBP) and allow for the creation of semisynthetic organisms (SSOs) with additional sense codons. Here, we report a systematic analysis of the unnatural codons. We identify nine unnatural codons that can produce unnatural protein with nearly complete incorporation of an encoded noncanonical amino acid (ncAA). We also show that at least three of the codons are orthogonal and can be simultaneously decoded in the SSO, affording the first 67-codon organism. The ability to incorporate multiple, different ncAAs site specifically into a protein should now allow the development of proteins with novel activities, and possibly even SSOs with new forms and functions.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Annotated plasmid sequences from this study are available via Genbank (accession numbers MN882182–MN882190) as detailed in Supplementary Table 4. All data supporting the findings of this study are available within the paper and the supplementary information or from the corresponding author upon reasonable request.
References
Leader, B., Baca, Q. J. & Golan, D. E. Protein therapeutics: a summary and pharmacological classification. Nat. Rev. Drug Discov. 7, 21–39 (2008).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Blight, S. K. et al. Direct charging of tRNA(CUA) with pyrrolysine in vitro and in vivo. Nature 431, 333–335 (2004).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J. W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Chatterjee, A., Sun, S. B., Furman, J. L., Xiao, H. & Schultz, P. G. A versatile platform for single- and multiple-unnatural amino acid mutagenesis in Escherichia coli. Biochemistry 52, 1828–1837 (2013).
Italia, J. S. et al. Mutually orthogonal nonsense-suppression systems and conjugation chemistries for precise protein labeling at up to three distinct sites. J. Am. Chem. Soc. 141, 6204–6212 (2019).
Lajoie, M. J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Aerni, H. R., Shifman, M. A., Rogulina, S., O’Donoghue, P. & Rinehart, J. Revealing the amino acid composition of proteins within an expanded genetic code. Nucleic Acids Res. 43, e8 (2015).
Hamashima, K., Kimoto, M. & Hirao, I. Creation of unnatural base pairs for genetic alphabet expansion toward synthetic xenobiology. Curr. Opin. Chem. Biol. 46, 108–114 (2018).
Biondi, E. & Benner, S. A. Artificially expanded genetic information systems for new aptamer technologies. Biomedicines 6, 53 (2018).
Li, L. et al. Natural-like replication of an unnatural base pair for the expansion of the genetic alphabet and biotechnology applications. J. Am. Chem. Soc. 136, 826–829 (2014).
Morris, S. E., Feldman, A. W. & Romesberg, F. E. Synthetic biology parts for the storage of increased genetic information in cells. ACS Synth. Biol. 6, 1834–1840 (2017).
Seo, Y. J., Hwang, G. T., Ordoukhanian, P. & Romesberg, F. E. Optimization of an unnatural base pair toward natural-like replication. J. Am. Chem. Soc. 131, 3246–3252 (2009).
Ast, M. et al. Diatom plastids depend on nucleotide import from the cytosol. Proc. Natl Acad. Sci. USA 106, 3621–3626 (2009).
Malyshev, D. A. et al. A semi-synthetic organism with an expanded genetic alphabet. Nature 509, 385–388 (2014).
Ledbetter, M. P., Karadeema, R. J. & Romesberg, F. E. Reprograming the replisome of a semisynthetic organism for the expansion of the genetic alphabet. J. Am. Chem. Soc. 140, 758–765 (2018).
Zhang, Y. et al. A semisynthetic organism engineered for the stable expansion of the genetic alphabet. Proc. Natl Acad. Sci. USA 114, 1317–1322 (2017).
Zhang, Y. et al. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 551, 644–647 (2017).
Pedelacq, J. D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Nguyen, D. P. et al. Genetic encoding and labeling of aliphatic azides and alkynes in recombinant proteins via a pyrrolysyl-tRNA synthetase/tRNACUA pair and click chemistry. J. Am. Chem. Soc. 131, 8720–8721 (2009).
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. 13, 1253–1260 (2017).
Dien, V. T. et al. Progress toward a semi-synthetic organism with an unrestricted expanded genetic alphabet. J. Am. Chem. Soc. 140, 16115–16123 (2018).
Chin, J. W. et al. Addition of p-azido-l-phenylalanine to the genetic code of Escherichia coli. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Shimizu, M., Asahara, H., Tamura, K., Hasegawa, T. & Himeno, H. The role of anticodon bases and the discriminator nucleotide in the recognition of some E. coli tRNAs by their aminoacyl-tRNA synthetases. J. Mol. Evol. 35, 436–443 (1992).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Ostrov, N. et al. Design, synthesis, and testing toward a 57-codon genome. Science 353, 819–822 (2016).
Nissen, P., Ippolito, J. A., Ban, N., Moore, P. B. & Steitz, T. A. RNA tertiary interactions in the large ribosomal subunit: the A-minor motif. Proc. Natl Acad. Sci. USA 98, 4899–4903 (2001).
Ramakrishnan, V. Ribosome structure and the mechanism of translation. Cell 108, 557–572 (2002).
Betz, K. et al. Structural insights into DNA replication without hydrogen bonds. J. Am. Chem. Soc. 135, 18637–18643 (2013).
Betz, K. et al. KlenTaq polymerase replicates unnatural base pairs by inducing a Watson–Crick geometry. Nat. Chem. Biol. 8, 612–614 (2012).
Hirao, I. et al. An unnatural base pair for incorporating amino acid analogs into proteins. Nat. Biotechnol. 20, 177–182 (2002).
Bain, J. D., Switzer, C., Chamberlin, A. R. & Benner, S. A. Ribosome-mediated incorporation of a non-standard amino acid into a peptide through expansion of the genetic code. Nature 356, 537–539 (1992).
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001).
Hoernes, T. P. et al. Translation of non-standard codon nucleotides reveals minimal requirements for codon–anticodon interactions. Nat. Commun. 9, 4865 (2018).
Feldman, A. W. & Romesberg, F. E. In vivo structure–activity relationships and optimization of an unnatural base pair for replication in a semi-synthetic organism. J. Am. Chem. Soc. 139, 11427–11433 (2017).
Schwark, D. G., Schmitt, M. A. & Fisk, J. D. Dissecting the contribution of release factor interactions to amber stop codon reassignment efficiencies of the Methanocaldococcus jannaschii orthogonal pair. Genes 9, E546 (2018).
O’Donoghue, P., Ling, J., Wang, Y. S. & Soll, D. Upgrading protein synthesis for synthetic biology. Nat. Chem. Biol. 9, 594–598 (2013).
Feldman, A. W. et al. Optimization of replication, transcription, and translation in a semi-synthetic organism. J. Am. Chem. Soc. 141, 10644–10653 (2019).
Gibson, D. G. Enzymatic assembly of overlapping DNA fragments. Methods Enzymol. 498, 349–361 (2011).
Acknowledgements
This work was supported by the National Institutes of Health (GM118178 to F.E.R., GM123735 to Y.Z. and GM128376 to R.J.K.). E.C.F. was supported by a Boehringer Ingelheim Fonds PhD Fellowship. K.H. was supported by a JSPS Overseas Research Fellowship. A.W.F. and M.P.L. were supported by a National Science Foundation Graduate Research Fellowship (NSF/DGE-1346837). R.K. was supported by NASA Exobiology (NNX14AP59G).
Author information
Authors and Affiliations
Contributions
F.E.R., Y.Z. and E.C.F. conceived the project. E.C.F. and F.E.R. designed experiments. E.C.F., K.H., A.W.F. and V.T.D. performed and analyzed experiments. R.J.K. and R.K. synthesized unnatural DNA oligonucleotides. A.W.F., M.P.L. and R.A. provided technical assistance. F.E.R. provided project leadership. E.C.F. and F.E.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing financial interests: a patent application has been filed based on the use of UBPs in SSOs (PCT/US2018/041509).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11 and Tables 1–4.
Rights and permissions
About this article
Cite this article
Fischer, E.C., Hashimoto, K., Zhang, Y. et al. New codons for efficient production of unnatural proteins in a semisynthetic organism. Nat Chem Biol 16, 570–576 (2020). https://doi.org/10.1038/s41589-020-0507-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-020-0507-z
This article is cited by
-
Quintuply orthogonal pyrrolysyl-tRNA synthetase/tRNAPyl pairs
Nature Chemistry (2023)
-
Consistent Clustering Pattern of Prokaryotic Genes Based on Base Frequency at the Second Codon Position and its Association with Functional Category Preference
Interdisciplinary Sciences: Computational Life Sciences (2022)
-
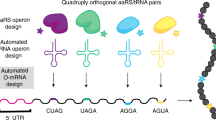
A 68-codon genetic code to incorporate four distinct non-canonical amino acids enabled by automated orthogonal mRNA design
Nature Chemistry (2021)
-
An engineered IL-2 reprogrammed for anti-tumor therapy using a semi-synthetic organism
Nature Communications (2021)
-
A call for caution in analysing mammalian co-transfection experiments and implications of resource competition in data misinterpretation
Nature Communications (2021)