Key Points
-
Bile acids are critical components of the gastrointestinal tract that link the gut microbiota to hepatic and intestinal metabolism and therefore influence gastrointestinal motility, intestinal permeability and carcinogenesis
-
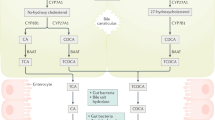
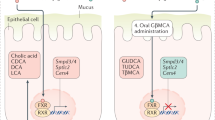
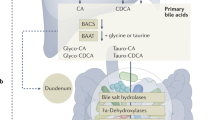
The gut microbiota regulates bile acid production and signalling via the biotransformation of intestinal bile acids to unconjugated and secondary forms that readily activate bile acid receptors
-
Bile acids are ligands for the G protein-coupled bile acid receptor 1 (TGR5) and for the nuclear hormone receptor farnesoid X receptor (FXR)
-
The profiles of bile acids and gut microbiota influence each other; bile acids can modulate microbiota composition, which in turn regulates the size and composition of the bile acid pool
-
Disruption of bile acid–microbiota crosstalk promotes inflammation and a gastrointestinal disease phenotype, which can contribute to the development of gastrointestinal cancers, including colorectal cancer and hepatocellular carcinoma
-
The modulation of gut microbiota and bile acid profiles holds promise as a novel therapeutic approach for the treatment of gastrointestinal cancers and represents the next frontier for gastrointestinal cancer research
Abstract
Emerging evidence points to a strong association between the gut microbiota and the risk, development and progression of gastrointestinal cancers such as colorectal cancer (CRC) and hepatocellular carcinoma (HCC). Bile acids, produced in the liver, are metabolized by enzymes derived from intestinal bacteria and are critically important for maintaining a healthy gut microbiota, balanced lipid and carbohydrate metabolism, insulin sensitivity and innate immunity. Given the complexity of bile acid signalling and the direct biochemical interactions between the gut microbiota and the host, a systems biology perspective is required to understand the liver–bile acid–microbiota axis and its role in gastrointestinal carcinogenesis to reverse the microbiota-mediated alterations in bile acid metabolism that occur in disease states. An examination of recent research progress in this area is urgently needed. In this Review, we discuss the mechanistic links between bile acids and gastrointestinal carcinogenesis in CRC and HCC, which involve two major bile acid-sensing receptors, farnesoid X receptor (FXR) and G protein-coupled bile acid receptor 1 (TGR5). We also highlight the strategies and cutting-edge technologies to target gut-microbiota-dependent alterations in bile acid metabolism in the context of cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mucci, L. A., Wedren, S., Tamimi, R. M., Trichopoulos, D. & Adami, H. O. The role of gene-environment interaction in the aetiology of human cancer: examples from cancers of the large bowel, lung and breast. J. Intern. Med. 249, 477–493 (2001).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
Siegel, R. L. et al. Colorectal cancer statistics, 2017. CA Cancer J. Clini. 67, 177–193 (2017).
International Agency for Research on Cancer. Liver cancer. GLOBOCAN 2012: Estimated Incidence Mortality and Prevalence Worldwide in 2012 http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspx (2017).
Jemal, A. et al. Global cancer statistics. CA Cancer J. Clin. 61, 69–90 (2011).
Jia, W., Li, H., Zhao, L. & Nicholson, J. K. Gut microbiota: a potential new territory for drug targeting. Nat. Rev. Drug Discov. 7, 123–129 (2008).
Nicholson, J. K. et al. Host-gut microbiota metabolic interactions. Science 336, 1262–1267 (2012).
Monte, M. J., Marin, J. J. G., Antelo, A. & Vazquez-Tato, J. Bile acids: chemistry, physiology, and pathophysiology. World J. Gastroenterol. 15, 804–816 (2009).
Swann, J. R. et al. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. Proc. Natl Acad. Sci. USA 108 (Suppl. 1), 4523–4530 (2011).
Dumas, M. E. et al. Metabolic profiling reveals a contribution of gut microbiota to fatty liver phenotype in insulin-resistant mice. Proc. Natl Acad. Sci. USA 103, 12511–12516 (2006).
Holmes, E. et al. Therapeutic modulation of microbiota-host metabolic interactions. Sci. Transl Med. 4, 137rv136 (2012).
Musso, G., Gambino, R. & Cassader, M. Interactions between gut microbiota and host metabolism predisposing to obesity and diabetes. Annu. Rev. Med. 62, 361–380 (2011).
Corrêa-Oliveira, R., Fachi, J. L., Vieira, A., Sato, F. T. & Vinolo, M. A. R. Regulation of immune cell function by short-chain fatty acids. Clin. Transl Immunology 5, e73 (2016).
Iida, N. et al. Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342, 967–970 (2013).
Zheng, X. et al. Melamine-induced renal toxicity is mediated by the gut microbiota. Sci. Transl Med. 5, 172ra122 (2013).
Ahn, J. et al. Human gut microbiome and risk for colorectal cancer. J. Natl Cancer Inst. 105, 1907–1911 (2013).
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013).
Sheh, A. & Fox, J. G. The role of the gastrointestinal microbiome in Helicobacter pylori pathogenesis. Gut Microbes 4, 505–531 (2013).
Moss, S. F. The clinical evidence linking Helicobacter pylori to gastric cancer. Cell. Mol. Gastroenterol. Hepatol. 3, 183–191 (2017).
Carboni, M. et al. Chronic atrophic gastritis and risk of N-nitroso compounds carcinogenesis. Langenbecks Arch. Chir. 373, 82–90 (1988).
Caygill, C. P., Hill, M. J., Braddick, M. & Sharp, J. C. Cancer mortality in chronic typhoid and paratyphoid carriers. Lancet 343, 83–84 (1994).
Prieto, A. I., Ramos-Morales, F. & Casadesús, J. Bile-induced DNA damage in Salmonella enterica. Genetics 168, 1787–1794 (2004).
Boleij, A. et al. The Bacteroides fragilis toxin gene is prevalent in the colon mucosa of colorectal cancer patients. Clin. Infect. Dis. 60, 208–215 (2015).
McCoy, A. N. et al. Fusobacterium is associated with colorectal adenomas. PLoS ONE 8, 15 (2013).
Schwabe, R. F. & Jobin, C. The microbiome and cancer. Nat. Rev. Cancer 13, 800–812 (2013).
Chen, G. Y., Shaw, M. H., Redondo, G. & Nunez, G. The innate immune receptor Nod1 protects the intestine from inflammation-induced tumorigenesis. Cancer Res. 68, 10060–10067 (2008).
Quante, M. et al. Bile acid and inflammation activate gastric cardia stem cells in a mouse model of Barrett-like metaplasia. Cancer Cell 21, 36–51 (2012).
Bernstein, C. et al. Carcinogenicity of deoxycholate, a secondary bile acid. Arch. Toxicol. 85, 863–871 (2011).
Lozano, E. et al. Cocarcinogenic effects of intrahepatic bile acid accumulation in cholangiocarcinoma development. Mol. Cancer Res. 12, 91–100 (2014).
Forman, B. M. et al. Identification of a nuclear receptor that is activated by farnesol metabolites. Cell 81, 687–693 (1995).
Seol, W., Choi, H. S. & Moore, D. D. Isolation of proteins that interact specifically with the retinoid X receptor: two novel orphan receptors. Mol. Endocrinol. 9, 72–85 (1995).
Hylemon, P. B. et al. Bile acids as regulatory molecules. J. Lipid Res. 50, 1509–1520 (2009).
Thomas, C., Pellicciari, R., Pruzanski, M., Auwerx, J. & Schoonjans, K. Targeting bile-acid signalling for metabolic diseases. Nat. Rev. Drug Discov. 7, 678–693 (2008).
Zollner, G., Wagner, M. & Trauner, M. Nuclear receptors as drug targets in cholestasis and drug-induced hepatotoxicity. Pharmacol. Ther. 126, 228–243 (2010).
Mora, J. R., Iwata, M. & von Andrian, U. H. Vitamin effects on the immune system: vitamins A and D take centre stage. Nat. Rev. Immunol. 8, 685–698 (2008).
Allen, K., Jaeschke, H. & Copple, B. L. Bile acids induce inflammatory genes in hepatocytes: a novel mechanism of inflammation during obstructive cholestasis. Am. J. Pathol. 178, 175–186 (2011).
Dawson, P. A., Lan, T. & Rao, A. Bile acid transporters. J. Lipid Res. 50, 2340–2357 (2009).
Ridlon, J. M., Kang, D. J. & Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J. Lipid. Res. 47, 241–259 (2006).
Axelson, M. et al. Bile acid synthesis in cultured human hepatocytes: support for an alternative biosynthetic pathway to cholic acid. Hepatology 31, 1305–1312 (2000).
Trauner, M. & Boyer, J. L. Bile salt transporters: molecular characterization, function, and regulation. Physiol. Rev. 83, 633–671 (2003).
Meier, P. J. & Stieger, B. Bile salt transporters. Annu. Rev. Physiol. 64, 635–661 (2002).
Zollner, G., Marschall, H. U., Wagner, M. & Trauner, M. Role of nuclear receptors in the adaptive response to bile acids and cholestasis: pathogenetic and therapeutic considerations. Mol. Pharm. 3, 231–251 (2006).
Lemoy, M.-J. M. F., Westworth, D. R., Ardeshir, A. & Tarara, R. P. Reference intervals for preprandial and postprandial serum bile acid in adult rhesus macaques (Macaca mulatta). J. Am. Assoc. Lab. Anim. Sci. 52, 444–447 (2013).
Holm, R., Mullertz, A. & Mu, H. Bile salts and their importance for drug absorption. Int. J. Pharm. 453, 44–55 (2013).
Geyer, J., Wilke, T. & Petzinger, E. The solute carrier family SLC10: more than a family of bile acid transporters regarding function and phylogenetic relationships. Naunyn Schmiedebergs Arch. Pharmacol. 372, 413–431 (2006).
Dawson, P. A. Role of the intestinal bile acid transporters in bile acid and drug disposition. Handb. Exp. Pharmacol. http://dx.doi.org/10.1007/978-3-642-14541-4_4 (2011).
Gadaleta, R. M., Cariello, M., Sabba, C. & Moschetta, A. Tissue-specific actions of FXR in metabolism and cancer. Biochim. Biophys. Acta 1851, 30–39 (2015).
Matsubara, T., Li, F. & Gonzalez, F. J. FXR signaling in the enterohepatic system. Mol. Cell. Endocrinol. 368, 17–29 (2013).
Kullak-Ublick, G. A., Stieger, B. & Meier, P. J. Enterohepatic bile salt transporters in normal physiology and liver disease. Gastroenterology 126, 322–342 (2004).
Hata, S. et al. Substrate specificities of rat oatp1 and ntcp: implications for hepatic organic anion uptake. Am. J. Physiol. Gastrointest. Liver Physiol. 285, G829–G839 (2003).
Chiang, J. Y. Bile acids: regulation of synthesis. J. Lipid Res. 50, 1955–1966 (2009).
Stanimirov, B., Stankov, K. & Mikov, M. Bile acid signaling through farnesoid X and TGR5 receptors in hepatobiliary and intestinal diseases. Hepatobiliary Pancreat. Dis. Int. 14, 18–33 (2015).
Botham, K. M. & Boyd, G. S. The metabolism of chenodeoxycholic acid to beta-muricholic acid in rat liver. Eur. J. Biochem. 134, 191–196 (1983).
Bergstrom, S., Danielsson, H. & Goransson, A. On the bile acid meataboism in the pig. Acta Chem. Scand. 13, 776–783 (1959).
Martin, F. P. et al. A top-down systems biology view of microbiome-mammalian metabolic interactions in a mouse model. Mol. Syst. Biol. 3, 112 (2007).
Gao, J. et al. Association between serum bile acid profiles and gestational diabetes mellitus: a targeted metabolomics study. Clin. Chim. Acta 459, 63–72 (2016).
García-Cañaveras, J. C., Donato, M. T., Castell, J. V. & Lahoz, A. Targeted profiling of circulating and hepatic bile acids in human, mouse, and rat using a UPLC-MRM-MS-validated method. J. Lipid Res. 53, 2231–2241 (2012).
Sayin, S. I. et al. Gut microbiota regulates bile acid metabolism by reducing the levels of tauro-beta-muricholic acid, a naturally occurring FXR antagonist. Cell Metab. 17, 225–235 (2013).
Stanimirov, B., Stankov, K. & Mikov, M. Pleiotropic functions of bile acids mediated by the farnesoid X receptor. Acta Gastroenterol. Belg. 75, 389–398 (2012).
Gérard, P. Metabolism of cholesterol and bile acids by the gut microbiota. Pathogens 3, 14–24 (2014).
Parks, D. J. et al. Bile acids: natural ligands for an orphan nuclear receptor. Science 284, 1365–1368 (1999).
Makishima, M. et al. Identification of a nuclear receptor for bile acids. Science 284, 1362–1365 (1999).
Vaquero, J., Monte, M. J., Dominguez, M., Muntane, J. & Marin, J. J. Differential activation of the human farnesoid X receptor depends on the pattern of expressed isoforms and the bile acid pool composition. Biochem. Pharmacol. 86, 926–939 (2013).
Song, P., Rockwell, C. E., Cui, J. Y. & Klaassen, C. D. Individual bile acids have differential effects on bile acid signaling in mice. Toxicol. Appl. Pharmacol. 283, 57–64 (2015).
Kanda, T. et al. Regulation of expression of human intestinal bile acid-binding protein in Caco-2 cells. Biochem. J. 330, 261–265 (1998).
Beuers, U., Trauner, M., Jansen, P. & Poupon, R. New paradigms in the treatment of hepatic cholestasis: from UDCA to FXR, PXR and beyond. J. Hepatol. 62, S25–S37 (2015).
Duboc, H., Taché, Y. & Hofmann, A. F. The bile acid TGR5 membrane receptor: from basic research to clinical application. Dig. Liver Dis. 46, 302–312 (2014).
Maruyama, T. et al. Identification of membrane-type receptor for bile acids (M-BAR). Biochem. Biophys. Res. Commun. 298, 714–719 (2002).
Kawamata, Y. et al. A G protein-coupled receptor responsive to bile acids. J. Biol. Chem. 278, 9435–9440 (2003).
Stanley, L. A., Horsburgh, B. C., Ross, J., Scheer, N. & Wolf, C. R. PXR and CAR: nuclear receptors which play a pivotal role in drug disposition and chemical toxicity. Drug Metab. Rev. 38, 515–597 (2006).
Yoneno, K. et al. TGR5 signalling inhibits the production of pro-inflammatory cytokines by in vitro differentiated inflammatory and intestinal macrophages in Crohn's disease. Immunology 139, 19–29 (2013).
Kliewer, S. A. & Willson, T. M. Regulation of xenobiotic and bile acid metabolism by the nuclear pregnane X receptor. J. Lipid Res. 43, 359–364 (2002).
Wagner, M. et al. CAR and PXR agonists stimulate hepatic bile acid and bilirubin detoxification and elimination pathways in mice. Hepatology 42, 420–430 (2005).
Zhou, J., Liu, M., Zhai, Y. & Xie, W. The antiapoptotic role of pregnane X receptor in human colon cancer cells. Mol. Endocrinol. 22, 868–880 (2008).
Uppal, H. et al. Combined loss of orphan receptors PXR and CAR heightens sensitivity to toxic bile acids in mice. Hepatology 41, 168–176 (2005).
Stedman, C. A. et al. Nuclear receptors constitutive androstane receptor and pregnane X receptor ameliorate cholestatic liver injury. Proc. Natl Acad. Sci. USA 102, 2063–2068 (2005).
Wang, H. et al. Pregnane X receptor activation induces FGF19-dependent tumor aggressiveness in humans and mice. J. Clin. Invest. 121, 3220–3232 (2011).
He, J., Nishida, S., Xu, M., Makishima, M. & Xie, W. PXR prevents cholesterol gallstone disease by regulating biosynthesis and transport of bile salts. Gastroenterology 140, 2095–2106 (2011).
Chatterjee, B., Echchgadda, I. & Song, C. S. Vitamin D receptor regulation of the steroid/bile acid sulfotransferase SULT2A1. Methods Enzymol. 400, 165–191 (2005).
Chen, X. et al. Transactivation of rat apical sodium-dependent bile acid transporter and increased bile acid transport by 1alpha, 25-dihydroxyvitamin D3 via the vitamin D receptor. Mol. Pharmacol. 69, 1913–1923 (2006).
Schmidt, D. R. et al. Regulation of bile acid synthesis by fat-soluble vitamins A and D. J. Biol. Chem. 285, 14486–14494 (2010).
Ding, N. et al. A vitamin D receptor/SMAD genomic circuit gates hepatic fibrotic response. Cell 153, 601–613 (2013).
Massague, J. TGFbeta in Cancer. Cell 134, 215–230 (2008).
Gadaleta, R. M. et al. Activation of bile salt nuclear receptor FXR is repressed by pro-inflammatory cytokines activating NF-kappaB signaling in the intestine. Biochim. Biophys. Acta 1812, 851–858 (2011).
Lefebvre, P., Cariou, B., Lien, F., Kuipers, F. & Staels, B. Role of bile acids and bile acid receptors in metabolic regulation. Physiol. Rev. 89, 147–191 (2009).
Chiang, J. Y. Bile acid metabolism and signaling. Compr. Physiol. 3, 1191–1212 (2013).
Chen, T. L. et al. Serum and urine metabolite profiling reveals potential biomarkers of human hepatocellular carcinoma. Mol. Cell. Proteomics 10, M110.004945 (2011).
Wang, X. N. et al. Urinary metabolite variation is associated with pathological progression of the post-hepatitis B cirrhosis patients. J. Proteome Res. 11, 3838–3847 (2012).
Ananthanarayanan, M., Balasubramanian, N., Makishima, M., Mangelsdorf, D. J. & Suchy, F. J. Human bile salt export pump promoter is transactivated by the farnesoid X receptor/bile acid receptor. J. Biol. Chem. 276, 28857–28865 (2001).
Landrier, J. F., Eloranta, J. J., Vavricka, S. R. & Kullak-Ublick, G. A. The nuclear receptor for bile acids, FXR, transactivates human organic solute transporter-alpha and -beta genes. Am. J. Physiol. Gastrointest. Liver Physiol. 290, G476–G485 (2006).
Boyer, J. L. et al. Upregulation of a basolateral FXR-dependent bile acid efflux transporter OSTalpha-OSTbeta in cholestasis in humans and rodents. Am. J. Physiol. Gastrointest. Liver Physiol. 290, G1124–G1130 (2006).
Zhao, L. et al. High throughput and quantitative measurement of microbial metabolome by gas chromatography/mass spectrometry using automated alkyl chloroformate derivatization. Anal. Chem. 89, 5565–5577 (2017).
Sinal, C. J. et al. Targeted disruption of the nuclear receptor FXR/BAR impairs bile acid and lipid homeostasis. Cell 102, 731–744 (2000).
Yang, F. et al. Spontaneous development of liver tumors in the absence of the bile acid receptor farnesoid X receptor. Cancer Res. 67, 863–867 (2007).
Cuperus, F. J., Claudel, T., Gautherot, J., Halilbasic, E. & Trauner, M. The role of canalicular ABC transporters in cholestasis. Drug Metab. Dispos. 42, 546–560 (2014).
Halilbasic, E., Claudel, T. & Trauner, M. Bile acid transporters and regulatory nuclear receptors in the liver and beyond. J. Hepatol. 58, 155–168 (2013).
Zollner, G. et al. Hepatobiliary transporter expression in human hepatocellular carcinoma. Liver Int. 25, 367–379 (2005).
Anwer, M. S. Intracellular signaling by bile acids. J. Biosci. (Rajshari) 20, 1–23 (2014).
Pikarsky, E. et al. NF-[kappa]B functions as a tumour promoter in inflammation-associated cancer. Nature 431, 461–466 (2004).
Luedde, T. & Schwabe, R. F. NF-κB in the liver—linking injury, fibrosis and hepatocellular carcinoma. Nat. Rev. Gastroenterol. Hepatol. 8, 108–118 (2011).
Jung, I. H. et al. Predominant activation of JAK/STAT3 pathway by interleukin-6 is implicated in hepatocarcinogenesis. Neoplasia 17, 586–597 (2015).
Nag, S., Qin, J., Srivenugopal, K. S., Wang, M. & Zhang, R. The MDM2-p53 pathway revisited. J. Biomed. Res. 27, 254–271 (2013).
Bernstein, H., Bernstein, C., Payne, C. M., Dvorakova, K. & Garewal, H. Bile acids as carcinogens in human gastrointestinal cancers. Mutat. Res. 589, 47–65 (2005).
Bernstein, H., Bernstein, C., Payne, C. M. & Dvorak, K. Bile acids as endogenous etiologic agents in gastrointestinal cancer. World J. Gastroenterol. 15, 3329–3340 (2009).
Ouyang, X., Ghani, A. & Mehal, W. Z. Inflammasome biology in fibrogenesis. Biochim. Biophys. Acta 1832, 979–988 (2013).
Degirolamo, C. et al. Prevention of spontaneous hepatocarcinogenesis in farnesoid X receptor-null mice by intestinal-specific farnesoid X receptor reactivation. Hepatology 61, 161–170 (2015).
Kong, B. et al. Mice with hepatocyte-specific FXR deficiency are resistant to spontaneous but susceptible to cholic acid-induced hepatocarcinogenesis. Am. J. Physiol. Gastrointest. Liver Physiol. 310, G295–G302 (2016).
Knisely, A. S. et al. Hepatocellular carcinoma in ten children under five years of age with bile salt export pump deficiency. Hepatology 44, 478–486 (2006).
Iannelli, F. et al. Massive gene amplification drives paediatric hepatocellular carcinoma caused by bile salt export pump deficiency. Nat. Commun. 5, 3850 (2014).
Amaral, J. D., Viana, R. J., Ramalho, R. M., Steer, C. J. & Rodrigues, C. M. Bile acids: regulation of apoptosis by ursodeoxycholic acid. J. Lipid Res. 50, 1721–1734 (2009).
Woolbright, B. L. & Jaeschke, H. Novel insight into mechanisms of cholestatic liver injury. World J. Gastroenterol. 18, 4985–4993 (2012).
Pean, N., Doignon, I. & Tordjmann, T. Bile acids and liver carcinogenesis: TGR5 as a novel piece in the puzzle? Clin. Res. Hepatol. Gastroenterol. 37, 226–229 (2013).
Gadaleta, R. M. et al. Bile acids and their nuclear receptor FXR: relevance for hepatobiliary and gastrointestinal disease. Biochim. Biophys. Acta 7, 683–692 (2010).
Zhang, Y. & Wang, L. Nuclear receptor small heterodimer partner in apoptosis signaling and liver cancer. Cancers 3, 198–212 (2011).
Li, G. et al. Small heterodimer partner overexpression partially protects against liver tumor development in farnesoid X receptor knockout mice. Toxicol. Appl. Pharmacol. 272, 299–305 (2013).
Wolfe, A. et al. Increased activation of the Wnt/beta-catenin pathway in spontaneous hepatocellular carcinoma observed in farnesoid X receptor knockout mice. J. Pharmacol. Exp. Ther. 338, 12–21 (2011).
Anakk, S. et al. Bile acids activate YAP to promote liver carcinogenesis. Cell Rep. 5, 1060–1069 (2013).
Cook, J. W., Kennaway, E. L. & Kennaway, N. M. Production of tumours in mice by deoxycholic acid. Nature 145, 627–627 (1940).
Xie, G. et al. Dysregulated hepatic bile acids collaboratively promote liver carcinogenesis. Int. J. Cancer 139, 1764–1775 (2016).
Xie, G. et al. Distinctly altered gut microbiota in the progression of liver disease. Oncotarget 7, 19355–19366 (2016).
Frank, D. N. et al. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc. Natl Acad. Sci. USA 104, 13780–13785 (2007).
Sokol, H. et al. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 15, 1183–1189 (2009).
Swidsinski, A., Loening-Baucke, V., Vaneechoutte, M. & Doerffel, Y. Active Crohn's disease and ulcerative colitis can be specifically diagnosed and monitored based on the biostructure of the fecal flora. Inflamm. Bowel Dis. 14, 147–161 (2008).
Dawson, P. A. & Karpen, S. J. Intestinal transport and metabolism of bile acids. J. Lipid Res. 56, 1085–1099 (2015).
Degirolamo, C., Rainaldi, S., Bovenga, F., Murzilli, S. & Moschetta, A. Microbiota modification with probiotics induces hepatic bile acid synthesis via downregulation of the Fxr-Fgf15 axis in mice. Cell Rep. 7, 12–18 (2014).
Nagengast, F. M., Grubben, M. J. & van Munster, I. P. Role of bile acids in colorectal carcinogenesis. Eur. J. Cancer 31A, 1067–1070 (1995).
Hofmann, A. F., Cravetto, C., Molino, G., Belforte, G. & Bona, B. Simulation of the metabolism and enterohepatic circulation of endogenous deoxycholic acid in humans using a physiologic pharmacokinetic model for bile acid metabolism. Gastroenterology 93, 693–709 (1987).
Huang, Y. & Chen, Z. Inflammatory bowel disease related innate immunity and adaptive immunity. Am. J. Transl Res. 8, 2490–2497 (2016).
Wang, Y. D., Chen, W. D., Yu, D., Forman, B. M. & Huang, W. The G-protein-coupled bile acid receptor, Gpbar1 (TGR5), negatively regulates hepatic inflammatory response through antagonizing nuclear factor kappa light-chain enhancer of activated B cells (NF-kappaB) in mice. Hepatology 54, 1421–1432 (2011).
Jensen, D. D. et al. The bile acid receptor TGR5 does not interact with beta-arrestins or traffic to endosomes but transmits sustained signals from plasma membrane rafts. J. Biol. Chem. 288, 22942–22960 (2013).
Cao, W. et al. Expression of bile acid receptor TGR5 in gastric adenocarcinoma. Am. J. Physiol. Gastrointest. Liver Physiol. 304, G322–G327 (2013).
Komohara, Y., Fujiwara, Y., Ohnishi, K. & Takeya, M. Tumor-associated macrophages: potential therapeutic targets for anti-cancer therapy. Adv. Drug Deliv. Rev. 99, 180–185 (2016).
Algars, A. et al. Type and location of tumor-infiltrating macrophages and lymphatic vessels predict survival of colorectal cancer patients. Int. J. Cancer 131, 864–873 (2012).
Cook, J. & Hagemann, T. Tumour-associated macrophages and cancer. Curr. Opin. Pharmacol. 13, 595–601 (2013).
McMahan, R. H. et al. Bile acid receptor activation modulates hepatic monocyte activity and improves nonalcoholic fatty liver disease. J. Bio. Chem. 288, 11761–11770 (2013).
Zhang, Y., Sime, W., Juhas, M. & Sjolander, A. Crosstalk between colon cancer cells and macrophages via inflammatory mediators and CD47 promotes tumour cell migration. Eur. J. Cancer 49, 3320–3334 (2013).
Calmus, Y. & Poupon, R. Shaping macrophages function and innate immunity by bile acids: mechanisms and implication in cholestatic liver diseases. Clin. Res. Hepatol. Gastroenterol. 38, 550–556 (2014).
Haselow, K. et al. Bile acids PKA-dependently induce a switch of the IL-10/IL-12 ratio and reduce proinflammatory capability of human macrophages. J. Leukoc. Biol. 94, 1253–1264 (2013).
Kamada, N., Seo, S. U., Chen, G. Y. & Nunez, G. Role of the gut microbiota in immunity and inflammatory disease. Nat. Rev. Immunol. 13, 321–335 (2013).
Littman, D. R. & Rudensky, A. Y. Th17 and regulatory T cells in mediating and restraining inflammation. Cell 140, 845–858 (2010).
Shapiro, H., Thaiss, C. A., Levy, M. & Elinav, E. The cross talk between microbiota and the immune system: metabolites take center stage. Curr. Opin. Immunol. 30, 54–62 (2014).
Stenman, L. K., Holma, R., Eggert, A. & Korpela, R. A novel mechanism for gut barrier dysfunction by dietary fat: epithelial disruption by hydrophobic bile acids. Am. J. Physiol. Gastrointest. Liver Physiol. 304, 29 (2013).
Brestoff, J. R. & Artis, D. Commensal bacteria at the interface of host metabolism and the immune system. Nat. Immunol. 14, 676–684 (2013).
Duboc, H. et al. Connecting dysbiosis, bile-acid dysmetabolism and gut inflammation in inflammatory bowel diseases. Gut 62, 531–539 (2013).
Attili, A. F., Angelico, M., Cantafora, A., Alvaro, D. & Capocaccia, L. Bile acid-induced liver toxicity: relation to the hydrophobic-hydrophilic balance of bile acids. Med. Hypotheses 19, 57–69 (1986).
Radominska, A. et al. Human liver steroid sulphotransferase sulphates bile acids. Biochem. J. 272, 597–604 (1990).
Gerard, P. Metabolism of cholesterol and bile acids by the gut microbiota. Pathogens 3, 14–24 (2013).
Van den Abbeele, P. et al. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. ISME J. 7, 949–961 (2013).
Wang, T. et al. Structural segregation of gut microbiota between colorectal cancer patients and healthy volunteers. ISME J. 6, 320–329 (2012).
Goncalves, P., Araujo, J. R., Pinho, M. J. & Martel, F. In vitro studies on the inhibition of colon cancer by butyrate and polyphenolic compounds. Nutr. Cancer 63, 282–294 (2011).
Hague, A., Elder, D. J., Hicks, D. J. & Paraskeva, C. Apoptosis in colorectal tumour cells: induction by the short chain fatty acids butyrate, propionate and acetate and by the bile salt deoxycholate. Int. J. Cancer 60, 400–406 (1995).
Bingham, S. A. et al. Dietary fibre in food and protection against colorectal cancer in the European Prospective Investigation into Cancer and Nutrition (EPIC): an observational study. Lancet 361, 1496–1501 (2003).
Clausen, M. R., Bonnen, H. & Mortensen, P. B. Colonic fermentation of dietary fibre to short chain fatty acids in patients with adenomatous polyps and colonic cancer. Gut 32, 923–928 (1991).
Grivennikov, S. I., Greten, F. R. & Karin, M. Immunity, inflammation, and cancer. Cell 140, 883–899 (2010).
Wang, Y. D. et al. Farnesoid X receptor antagonizes nuclear factor kappaB in hepatic inflammatory response. Hepatology 48, 1632–1643 (2008).
Fiorucci, S. et al. Protective effects of 6-ethyl chenodeoxycholic acid, a farnesoid X receptor ligand, in estrogen-induced cholestasis. J. Pharmacol. Exp. Ther. 313, 604–612 (2005).
Vavassori, P., Mencarelli, A., Renga, B., Distrutti, E. & Fiorucci, S. The bile acid receptor FXR is a modulator of intestinal innate immunity. J. Immunol. 183, 6251–6261 (2009).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Armaghany, T., Wilson, J. D., Chu, Q. & Mills, G. Genetic alterations in colorectal cancer. Gastrointest. Cancer Res. 5, 19–27 (2012).
Sears, C. L. & Garrett, W. S. Microbes, microbiota, and colon cancer. Cell Host Microbe 15, 317–328 (2014).
De Gottardi, A. et al. The bile acid nuclear receptor FXR and the bile acid binding protein IBABP are differently expressed in colon cancer. Dig. Dis. Sci. 49, 982–989 (2004).
Modica, S., Murzilli, S., Salvatore, L., Schmidt, D. R. & Moschetta, A. Nuclear bile acid receptor FXR protects against intestinal tumorigenesis. Cancer Res. 68, 9589–9594 (2008).
Torres, J. et al. Farnesoid X receptor expression is decreased in colonic mucosa of patients with primary sclerosing cholangitis and colitis-associated neoplasia. Inflamm. Bowel Dis. 19, 275–282 (2013).
Bailey, A. M. et al. FXR silencing in human colon cancer by DNA methylation and KRAS signaling. Am. J. Physiol. Gastrointest. Liver Physiol. 306, G48–G58 (2014).
Lax, S. et al. Expression of the nuclear bile acid receptor/farnesoid X receptor is reduced in human colon carcinoma compared to nonneoplastic mucosa independent from site and may be associated with adverse prognosis. Int. J. Cancer 130, 2232–2239 (2012).
Maran, R. R. et al. Farnesoid X receptor deficiency in mice leads to increased intestinal epithelial cell proliferation and tumor development. J. Pharm. Exp. Ther. 328, 469–477 (2009).
Ajouz, H., Mukherji, D. & Shamseddine, A. Secondary bile acids: an underrecognized cause of colon cancer. World J. Surg. Oncol. 12, 164–164 (2014).
Payne, C. M., Bernstein, C., Dvorak, K. & Bernstein, H. Hydrophobic bile acids, genomic instability, Darwinian selection, and colon carcinogenesis. Clin. Exp. Gastroenterol. 1, 19–47 (2008).
Washo-Stultz, D. et al. Role of mitochondrial complexes I and II, reactive oxygen species and arachidonic acid metabolism in deoxycholate-induced apoptosis. Cancer Lett. 177, 129–144 (2002).
Ridlon, J. M. & Bajaj, J. S. The human gut sterolbiome: bile acid-microbiome endocrine aspects and therapeutics. Acta Pharm. Sin. B 5, 99–105 (2015).
Labbe, A., Ganopolsky, J. G., Martoni, C. J., Prakash, S. & Jones, M. L. Bacterial bile metabolising gene abundance in Crohn's, ulcerative colitis and type 2 diabetes metagenomes. PLoS ONE 9, e115175 (2014).
Barrasa, J. I., Olmo, N., Lizarbe, M. A. & Turnay, J. Bile acids in the colon, from healthy to cytotoxic molecules. Toxicol. In Vitro 27, 964–977 (2013).
Axelrad, J. E., Lichtiger, S. & Yajnik, V. Inflammatory bowel disease and cancer: the role of inflammation, immunosuppression, and cancer treatment. World J. Gastroenterol. 22, 4794–4801 (2016).
Hold, G. L. et al. Role of the gut microbiota in inflammatory bowel disease pathogenesis: what have we learnt in the past 10 years? World J. Gastroenterol. 20, 1192–1210 (2014).
Mattar, M. C., Lough, D., Pishvaian, M. J. & Charabaty, A. Current management of inflammatory bowel disease and colorectal cancer. Gastrointest. Cancer Res. 4, 53–61 (2011).
Morelli, L. & Capurso, L. FAO/WHO guidelines on probiotics: 10 years later. J. Clin. Gastroenterol. 46, S1–S2 (2012).
Louis, P., Hold, G. L. & Flint, H. J. The gut microbiota, bacterial metabolites and colorectal cancer. Nat. Rev. Microbiol. 12, 661–672 (2014).
Ghouri, Y. A. et al. Systematic review of randomized controlled trials of probiotics, prebiotics, and synbiotics in inflammatory bowel disease. Clin. Exp. Gastroenterol. 7, 473–487 (2014).
Scaldaferri, F. et al. Gut microbial flora, prebiotics, and probiotics in IBD: their current usage and utility. Biomed. Res. Int. 2013, 435268 (2013).
Mallon, P., McKay, D., Kirk, S. & Gardiner, K. Probiotics for induction of remission in ulcerative colitis. Cochrane Database Syst. Rev. 4, CD005573 (2007).
Boyer, J. L. Nuclear receptor ligands: rational and effective therapy for chronic cholestatic liver disease? Gastroenterology 129, 735–740 (2005).
Zollner, G. & Trauner, M. Nuclear receptors as therapeutic targets in cholestatic liver diseases. Br. J. Pharmacol. 156, 7–27 (2009).
Hohenester, S. et al. A biliary HCO3- umbrella constitutes a protective mechanism against bile acid-induced injury in human cholangiocytes. Hepatology 55, 173–183 (2012).
Fickert, P. et al. 24-norUrsodeoxycholic acid is superior to ursodeoxycholic acid in the treatment of sclerosing cholangitis in Mdr2 (Abcb4) knockout mice. Gastroenterology 130, 465–481 (2006).
Ali, A. H., Carey, E. J. & Lindor, K. D. Recent advances in the development of farnesoid X receptor agonists. Ann. Transl Med. 3, 5 (2015).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02516605 (2017).
Nevens, F. et al. A placebo-controlled trial of obeticholic acid in primary biliary cholangitis. N. Engl. J. Med. 375, 631–643 (2016).
De Magalhaes Filho, C. D., Downes, M. & Evans, R. Bile acid analog intercepts liver fibrosis. Cell 166, 789 (2016)
Perino, A. & Schoonjans, K. TGR5 and immunometabolism: insights from physiology and pharmacology. Trends Pharmacol. Sci. 36, 847–857 (2015).
Festa, C. et al. Exploitation of cholane scaffold for the discovery of potent and selective farnesoid x receptor (FXR) and G-protein coupled bile acid receptor 1 (GP-BAR1) ligands. J. Med. Chem. 57, 8477–8495 (2014).
Pellicciari, R. et al. Nongenomic actions of bile acids. Synthesis and preliminary characterization of 23- and 6,23-alkyl-substituted bile acid derivatives as selective modulators for the G-protein coupled receptor TGR5. J. Med. Chem. 50, 4265–4268 (2007).
Pellicciari, R. et al. Discovery of 6alpha-ethyl-23(S)-methylcholic acid (S-EMCA, INT-777) as a potent and selective agonist for the TGR5 receptor, a novel target for diabesity. J. Med. Chem. 52, 7958–7961 (2009).
Kobayashi, M. et al. Prevention and treatment of obesity, insulin resistance, and diabetes by bile acid-binding resin. Diabetes 56, 239–247 (2007).
Rao, A. et al. Inhibition of ileal bile acid uptake protects against nonalcoholic fatty liver disease in high-fat diet-fed mice. Sci. Transl Med. 8, 357ra122 (2016).
Baghdasaryan, A. et al. Inhibition of intestinal bile acid absorption improves cholestatic liver and bile duct injury in a mouse model of sclerosing cholangitis. J. Hepatol. 64, 674–681 (2016).
Nicaise, C. et al. Control of acute, chronic, and constitutive hyperammonemia by wild-type and genetically engineered Lactobacillus plantarum in rodents. Hepatology 48, 1184–1192 (2008).
Jones, M. L., Chen, H., Ouyang, W., Metz, T. & Prakash, S. Microencapsulated genetically engineered Lactobacillus plantarum 80 (pCBH1) for bile acid deconjugation and its implication in lowering cholesterol. J. Biomed. Biotechnol. 2004, 61–69 (2004).
Xie, G. et al. Sex-dependent effects on gut microbiota regulate hepatic carcinogenic outcomes. Sci. Rep. 7, 45232 (2017).
Zheng, M. M., Wang, R. F., Li, C. X. & Xu, J. H. Two-step enzymatic synthesis of ursodeoxycholic acid with a new 7β-hydroxysteroid dehydrogenase from Ruminococcus torques. Process Biochem. 50, 598–604 (2015).
Palmer, R. H. Bile acid sulfates. II. Formation, metabolism, and excretion of lithocholic acid sulfates in the rat. J. Lipid Res. 12, 680–687 (1971).
Lee, J. et al. Adaptive regulation of bile salt transporters in kidney and liver in obstructive cholestasis in the rat. Gastroenterology 121, 1473–1484 (2001).
Nijmeijer, R. M. et al. Farnesoid X receptor (FXR) activation and FXR genetic variation in inflammatory bowel disease. PLoS ONE 6, e23745 (2011).
Li, T. & Chiang, J. Y. Bile acid signaling in metabolic disease and drug therapy. Pharmacol. Rev. 66, 948–983 (2014).
Holt, J. A. et al. Definition of a novel growth factor-dependent signal cascade for the suppression of bile acid biosynthesis. Genes Dev. 17, 1581–1591 (2003).
Inagaki, T. et al. Fibroblast growth factor 15 functions as an enterohepatic signal to regulate bile acid homeostasis. Cell Metab. 2, 217–225 (2005).
Mellor, H. R. Targeted inhibition of the FGF19-FGFR4 pathway in hepatocellular carcinoma; translational safety considerations. Liver Int. 34, e1–9 (2014).
Halilbasic, E., Baghdasaryan, A. & Trauner, M. Nuclear receptors as drug targets in cholestatic liver diseases. Clin. Liver Dis. 17, 161–189 (2013).
Prenzel, N. et al. EGF receptor transactivation by G-protein-coupled receptors requires metalloproteinase cleavage of proHB-EGF. Nature 402, 884–888 (1999).
Pelengaris, S., Khan, M. & Evan, G. c-MYC: more than just a matter of life and death. Nat. Rev. Cancer 2, 764–776 (2002).
Lee, H. Y., Crawley, S., Hokari, R., Kwon, S. & Kim, Y. S. Bile acid regulates MUC2 transcription in colon cancer cells via positive EGFR/PKC/Ras/ERK/CREB, PI3K/Akt/IkappaB/NF-kappaB and p38/MSK1/CREB pathways and negative JNK/c-Jun/AP-1 pathway. Int. J. Oncol. 36, 941–953 (2010).
Pai, R., Tarnawski, A. S. & Tran, T. Deoxycholic acid activates beta-catenin signaling pathway and increases colon cell cancer growth and invasiveness. Mol. Biol. Cell 15, 2156–2163 (2004).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02308111 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01999101 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02177136 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02548351 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02039219 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01265498 (2015).
Intercept Pharmaceuticals. Intercept Pharmaceuticals initiates phase 1 study of INT-767, a dual FXR and TGR5 agonist. InterceptPharma. http://ir.interceptpharma.com/releasedetail.cfm?releaseid=944736 (2015).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01877577 (2014).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03162432 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01755507 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02021110 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03004118 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01857284 (2013).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT00808743 (2013).
Acknowledgements
Wei J. and G.X. are supported by grants from the US NIH (1U01CA188387-01A1).
Author information
Authors and Affiliations
Contributions
Wei J. and G.X. researched data for the article and wrote the article. Wei J., G.X. and Weiping J. made substantial contributions to discussion of content and reviewed/edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Hypochlorhydria
-
A deficiency of hydrochloric acid in the stomach.
- Enterotoxigenic bacteria
-
Bacteria that cause disease in humans and domestic animals by producing enterotoxin.
- Enterohepatic circulation
-
The circulation of bile salts, bilirubin, drugs or other substances from the liver to the bile, followed by entry into the small intestine, absorption by enterocytes and return to the liver via the portal circulation.
- Bile canaliculus
-
A thin tube that collects bile secreted by hepatocytes.
- Lithogenic diet
-
A diet designed to increase the likelihood of stone formation, particularly gallstones.
- Senescence-associated secretory phenotype
-
A phenotype in which cells induced to senesce by genotoxic stress secrete pro-inflammatory cytokines, chemokines, and proteases associated with inflammation and malignancy.
- Primary biliary cholangitis
-
A type of liver disease caused by damage to the bile ducts in the liver.
- Primary sclerosing cholangitis
-
A chronic liver disease characterized by a progressive course of cholestasis with inflammation and fibrosis of the intrahepatic and extrahepatic bile ducts.
- Hyperammonaemia
-
A metabolic disturbance characterized by an excess of ammonia in the blood.
Rights and permissions
About this article
Cite this article
Jia, W., Xie, G. & Jia, W. Bile acid–microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat Rev Gastroenterol Hepatol 15, 111–128 (2018). https://doi.org/10.1038/nrgastro.2017.119
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrgastro.2017.119
This article is cited by
-
Exploration of the application potential of serum multi-biomarker model in colorectal cancer screening
Scientific Reports (2024)
-
Bile acid metabolism and signaling in health and disease: molecular mechanisms and therapeutic targets
Signal Transduction and Targeted Therapy (2024)
-
Ginseng extracts improve circadian clock gene expression and reduce inflammation directly and indirectly through gut microbiota and PI3K signaling pathway
npj Biofilms and Microbiomes (2024)
-
Metabolic phenotyping reveals an emerging role of ammonia abnormality in Alzheimer’s disease
Nature Communications (2024)
-
Defining the biogeographical map and potential bacterial translocation of microbiome in human ‘surface organs’
Nature Communications (2024)