Key Points
-
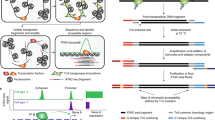
In next-generation sequencing (NGS) chromatin profiling experiments technical artefacts may be introduced at any stage, most importantly in fragmenting DNA, selecting the fragment population of interest, DNA amplification, DNA sequencing itself and read mapping to a reference genome.
-
The effect of technical biases on experimental results will depend, to a large extent, on the genomic scale of the feature being analysed and the scale on which the bias is manifested. Bias will have the greatest effect when the length scale of the bias is similar to the scale of the feature.
-
Genomic experiments should be planned to recognize the potential confounding effects of biases and the limits of the technology. Proper controls to understand and characterize the potential biases in chromatin profiling should be included and sequenced to sufficient depth in such experiments.
-
Nuclease-induced fragmentation is usually biased by DNA sequence in ways that can produce patterns that might seem to have biological importance.
-
Basic principles of statistical analysis should be applied to the analysis of chromatin profiling experiments: variability and bias should be taken into account, and the fit of statistical models to observed data should be characterized.
Abstract
Next-generation sequencing (NGS) technologies have been used in diverse ways to investigate various aspects of chromatin biology by identifying genomic loci that are bound by transcription factors, occupied by nucleosomes or accessible to nuclease cleavage, or loci that physically interact with remote genomic loci. However, reaching sound biological conclusions from such NGS enrichment profiles requires many potential biases to be taken into account. In this Review, we discuss common ways in which biases may be introduced into NGS chromatin profiling data, approaches to diagnose these biases and analytical techniques to mitigate their effect.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007). This paper reports the first use of MNase digestion followed by ChIP–seq to characterize genome-wide patterns of 20 varieties of histone lysine and arginine methylation.It identifies common modifications that are associated with active and repressed regions of the genome, transcription start sites, enhancers and insulator elements.
Johnson, D., Mortazavi, A., Myers, R. & Wold, B. Genome-wide mapping of in vivo protein–DNA interactions. Science 80, 1497–1502 (2007).
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Kharchenko, P. V., Tolstorukov, M. Y. & Park, P. J. Design and analysis of ChIP–seq experiments for DNA-binding proteins. Nature Biotech. 26, 1351–1359 (2008). This study proposes using the distribution of oriented reads to discriminate between real TF binding sites and artefacts.
Schones, D. E. et al. Dynamic regulation of nucleosome positioning in the human genome. Cell 132, 887–898 (2008).
He, H. H. et al. Nucleosome dynamics define transcriptional enhancers. Nature Genet. 42, 343–347 (2010).
Boyle, A. P. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011).
Hesselberth, J. R. et al. Global mapping of protein–DNA interactions in vivo by digital genomic footprinting. Nature Methods 6, 283–289 (2009).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nature Methods 10, 1213–1218 (2013).
Landt, S. G. et al. ChIP–seq guidelines and practices of the ENCODE and modENCODE consortia. Genome Res. 22, 1813–1831 (2012).
Teytelman, L. et al. Impact of chromatin structures on DNA processing for genomic analyses. PLoS ONE 4, e6700 (2009).
Modak, S. P. & Beard, P. Analysis of DNA double- and single-strand breaks by two dimensional electrophoresis: action of micrococcal nuclease on chromatin and DNA, and degradation in vivo of lens fiber chromatin. Nucleic Acids Res. 8, 2665–2678 (1980).
Zentner, G. E. & Henikoff, S. Surveying the epigenomic landscape, one base at a time. Genome Biol. 13, 250 (2012).
Telford, D. J. & Stewart, B. W. Micrococcal nuclease: its specificity and use for chromatin analysis. Int. J. Biochem. 21, 127–137 (1989).
Henikoff, J. G., Belsky, J. A., Krassovsky, K., Macalpine, D. M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl Acad. Sci. USA 108, 18318–18323 (2011).
Tillo, D. et al. High nucleosome occupancy is encoded at human regulatory sequences. PLoS ONE 5, e9129 (2010).
Valouev, A. et al. Determinants of nucleosome organization in primary human cells. Nature 474, 516–520 (2011).
Gaffney, D. J. et al. Controls of nucleosome positioning in the human genome. PLoS Genet. 8, e1003036 (2012).
Fan, X. et al. Nucleosome depletion at yeast terminators is not intrinsic and can occur by a transcriptional mechanism linked to 3′-end formation. Proc. Natl Acad. Sci. USA 107, 17945–17950 (2010).
Chung, H.-R. et al. The effect of micrococcal nuclease digestion on nucleosome positioning data. PLoS ONE 5, e15754 (2010).
Campbell, V. W. & Jackson, D. A. The effect of divalent cations on the mode of action of DNase I. The initial reaction products produced from covalently closed circular DNA. J. Biol. Chem. 255, 3726–3735 (1980).
He, H. H. et al. Refined DNase-seq protocol and data analysis reveals intrinsic bias in transcription factor footprint identification. Nature Methods 11, 73–78 (2014). This study shows how fragment size selection in DNase-seq can have a large impact on peak identification and that intrinsic DNase I cleavage bias can be mistaken as TF binding footprints.
Vierstra, J. Wang, H., John, S., Sandstrom, R. & Stamatoyannopoulos, J. A. Coupling transcription factor occupancy to nucleosome architecture with DNase–FLASH. Nature Methods 11, 66–72 (2014).
Lazarovici, A. et al. Probing DNA shape and methylation state on a genomic scale with DNase I. Proc. Natl Acad. Sci. USA 110, 6376–6381 (2013).
Grøntved, L. et al. Rapid genome-scale mapping of chromatin accessibility in tissue. Epigenetics Chromatin 5, 10 (2012).
Van Heesch, S. et al. Systematic biases in DNA copy number originate from isolation procedures. Genome Biol. 14, R33 (2013).
Giresi, P. G. & Lieb, J. D. Isolation of active regulatory elements from eukaryotic chromatin using FAIRE (formaldehyde assisted isolation of regulatory elements). Methods 48, 233–239 (2009).
Gilfillan, G. D. et al. Limitations and possibilities of low cell number ChIP–seq. BMC Genomics 13, 645 (2012).
Dabney, J. & Meyer, M. Length and GC-biases during sequencing library amplification: a comparison of various polymerase-buffer systems with ancient and modern DNA sequencing libraries. Biotechniques 52, 87–94 (2012).
Benjamini, Y. & Speed, T. P. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 40, e72 (2012). This study shows the importance of selecting the correct genomic interval for bias analysis, as some sources of bias are best modelled using properties of DNA fragments rather than DNA reads.
Wheeler, T. J. et al. Dfam: a database of repetitive DNA based on profile hidden Markov models. Nucleic Acids Res. 41, D70–D82 (2013).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Alkan, C. et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nature Genet. 41, 1061–1067 (2009).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Derrien, T. et al. Fast computation and applications of genome mappability. PLoS ONE 7, e30377 (2012).
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nature Genet. 42, 631–634 (2010).
Chung, D. et al. Discovering transcription factor binding sites in highly repetitive regions of genomes with multi-read analysis of ChIP–seq data. PLoS Comput. Biol. 7, e1002111 (2011).
Day, D. S., Luquette, L. J., Park, P. J. & Kharchenko, P. V. Estimating enrichment of repetitive elements from high-throughput sequence data. Genome Biol. 11, R69 (2010).
Wang, T. et al. Species-specific endogenous retroviruses shape the transcriptional network of the human tumor suppressor protein p53. Proc. Natl Acad. Sci. USA 104, 18613–18618 (2007).
Pickrell, J. K., Gaffney, D. J., Gilad, Y. & Pritchard, J. K. False positive peaks in ChIP–seq and other sequencing-based functional assays caused by unannotated high copy number regions. Bioinformatics 27, 2144–2146 (2011).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Rashid, N. U., Giresi, P. G., Ibrahim, J. G., Sun, W. & Lieb, J. D. ZINBA integrates local covariates with DNA-seq data to identify broad and narrow regions of enrichment, even within amplified genomic regions. Genome Biol. 12, R67 (2011).
Degner, J. F. et al. Effect of read-mapping biases on detecting allele-specific expression from RNA-sequencing data. Bioinformatics 25, 3207–3212 (2009).
Rozowsky, J. et al. AlleleSeq: analysis of allele-specific expression and binding in a network framework. Mol. Syst. Biol. 7, 522 (2011).
Sherwood, R. I. et al. Discovery of directional and nondirectional pioneer transcription factors by modeling DNase profile magnitude and shape. Nature Biotech. 32, 171–178 (2014).
König, J. et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nature Struct. Mol. Biol. 17, 909–915 (2010).
Daley, T. & Smith, A. D. Predicting the molecular complexity of sequencing libraries. Nature Methods 10, 325–327 (2013).
Marinov, G. K., Kundaje, A., Park, P. J. & Wold, B. J. Large-scale quality analysis of published ChIP–seq data. G3 (Bethesda) 4, 209–223 (2014).
Chen, Y. et al. Systematic evaluation of factors influencing ChIP–seq fidelity. Nature Methods 9, 609–614 (2012).
Ho, J. W. K. et al. ChIP–chip versus ChIP–seq: lessons for experimental design and data analysis. BMC Genomics 12, 134 (2011).
Bonhoure, N. et al. Quantifying ChIP–seq data: a spiking method providing an internal reference for sample-to-sample normalization. Genome Res. 24, 1157–1168 (2014).
Kidder, B. L., Hu, G. & Zhao, K. ChIP–seq: technical considerations for obtaining high-quality data. Nature Immunol. 12, 918–922 (2011).
Lassmann, T., Hayashizaki, Y. & Daub, C. O. SAMStat: monitoring biases in next generation sequencing data. Bioinformatics 27, 130–131 (2010).
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
Wang, L., Wang, S. & Li, W. RSeQC: quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185 (2012).
Planet, E. & Attolini, C. S., Reina, O., Flores, O. & Rossell, D. htSeqTools: high-throughput sequencing quality control, processing and visualization in R. Bioinformatics 28, 589–590 (2012).
Diaz, A., Nellore, A. & Song, J. S. CHANCE: comprehensive software for quality control and validation of ChIP–seq data. Genome Biol. 13, R98 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Hansen, K. D., Irizarry, R. A. & Wu, Z. Removing technical variability in RNA-seq data using conditional quantile normalization. Biostatistics 13, 204–216 (2012).
Cleveland, W. S. Robust locally and smoothing weighted regression scatterplots. J. Am. Stat. Soc. 74, 829–836 (2013).
Koenker, R. & Hallock, K. F. Quantile regression. J. Econ. Perspect. 15, 143–156 (2013).
Rozowsky, J. et al. PeakSeq enables systematic scoring of ChIP–seq experiments relative to controls. Nature Biotech. 27, 66–75 (2009).
Liang, K. & Keles, S. Detecting differential binding of transcription factors with ChIP–seq. Bioinformatics 28, 121–122 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Dillies, M.-A. et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief. Bioinform. 14, 671–683 (2012).
Shao, Z., Zhang, Y., Yuan, G.-C., Orkin, S. H. & Waxman, D. J. MAnorm: a robust model for quantitative comparison of ChIP–seq data sets. Genome Biol. 13, R16 (2012).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008). This study introduces the idea of estimating background effects using sliding windows on multiple scales. MACS remains one of the most widely used and best-performing algorithms for ChIP–seq peak calling.
Hashimoto, T. B., Edwards, M. D. & Gifford, D. K. Universal count correction for high-throughput sequencing. PLoS Comput. Biol. 10, 14–18 (2014).
Anders, S. et al. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nature Protoc. 8, 1765–1786 (2013).
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nature Methods 4, 651–657 (2007).
Ji, H. et al. An integrated software system for analyzing ChIP–chip and ChIP–seq data. Nature Biotech. 26, 1293–1300 (2008).
Nix, D. A., Courdy, S. J. & Boucher, K. M. Empirical methods for controlling false positives and estimating confidence in ChIP–seq peaks. BMC Bioinformatics 9, 1–9 (2008).
Valouev, A. et al. Genome-wide analysis of transcription factor binding sites based on ChIP–seq data. Nature Methods 5, 829–834 (2008).
Sun, G., Chung, D. & Liang, K. Statistical analysis of ChIP–seq data with MOSAiCS. Methods Mol. Biol. 1038, 193–212 (2013).
Zhang, X. et al. PICS: probabilistic inference for ChIP–seq. Biometrics 67, 151–163 (2011).
Kornacker, K., Rye, M. B., Håndstad, T. & Drabløs, F. The Triform algorithm: improved sensitivity and specificity in ChIP–seq peak finding BMC Bioinformatics 13, 176 (2012).
Kumar, V. et al. Uniform, optimal signal processing of mapped deep-sequencing data. Nature Biotech. 31, 615–622 (2013).
Chen, X., Hoffman, M. M., Bilmes, J. A., Hesselberth, J. R. & Noble, W. S. A dynamic Bayesian network for identifying protein-binding footprints from single molecule-based sequencing data. Bioinformatics 26, i334–i342 (2010).
Piper, J. et al. Wellington: a novel method for the accurate identification of digital genomic footprints from DNase-seq data. Nucleic Acids Res. 41, e201 (2013).
Fu, Y., Sinha, M., Peterson, C. L. & Weng, Z. The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome. PLoS Genet. 4, e1000138 (2008).
He, H. H. et al. Differential DNase I hypersensitivity reveals factor-dependent chromatin dynamics. Genome Res. 22, 1015–1025 (2012).
Pique-Regi, R. et al. Accurate inference of transcription factor binding from DNA sequence and chromatin accessibility data. Genome Res. 21, 447–455 (2011).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP–seq data. Bioinformatics 25, 1952–1958 (2009).
Song, Q. & Smith, A. D. Identifying dispersed epigenomic domains from ChIP–seq data. Bioinformatics 27, 870–871 (2011).
Wang, J., Lunyak, V. V. & Jordan, I. K. BroadPeak: a novel algorithm for identifying broad peaks in diffuse ChIP–seq datasets. Bioinformatics 29, 492–493 (2013).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nature Biotech. 28, 817–825 (2010).
Hoffman, M. M. et al. Unsupervised pattern discovery in human chromatin structure through genomic segmentation. Nature Methods 9, 473–476 (2012).
Lun, D. S., Sherrid, A., Weiner, B., Sherman, D. R. & Galagan, J. E. A blind deconvolution approach to high-resolution mapping of transcription factor binding sites from ChIP–seq data. 12, 1–12 (2009).
Guo, Y. et al. Discovering homotypic binding events at high spatial resolution. Bioinformatics 26, 3028–3034 (2010).
Chung, D. et al. dPeak: high resolution identification of transcription factor binding sites from PET and SET ChIP–seq data. PLos Comput. Biol. 9, 9–11 (2013).
Li, J., Jiang, H. & Wong, W. H. Modeling non-uniformity in short-read rates in RNA-seq data. Genome Biol. 11, 1–11 (2010).
Leek, J. T. et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nature Rev. Genet. 11, 733–739 (2010). This review discusses the importance of modelling batch effects in genome-wide analyses and statistical techniques for such analyses.
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Leek, J. T. & Storey, J. D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, 1724–1735 (2007).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Hu, M. et al. HiCNorm: removing biases in Hi-C data via Poisson regression. Bioinformatics 28, 3131–3133 (2012).
Hu, M. et al. Bayesian inference of spatial organizations of chromosomes. PLoS Comput. Biol. 9, e1002893 (2013).
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization Nature Methods 9, 999–1003 (2012). This study proposes a novel decomposition scheme for the analysis of Hi-C data that separates visibility and interaction components.
Dostie, J. et al. Chromosome conformation capture carbon copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Degner, J. F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Zeng, W. & Mortazavi, A. Technical considerations for functional sequencing assays. Nature Immunol. 13, 802–807 (2012).
Jung, Y. L. et al. Impact of sequencing depth in ChIP–seq experiments. Nucleic Acids Res. 42, e74 (2014).
Zhang, Y. et al. Intrinsic histone–DNA interactions are not the major determinant of nucleosome positions in vivo. Nature Struct. Mol. Biol. 16, 847–852 (2009).
Bravo, H. C. & Irizarry, R. A. Model-based quality assessment and base-calling for second-generation sequencing data. Biometrics 66, 665–674 (2010).
Pickrell, J. K., Gilad, Y. & Pritchard, J. K. Comment on “Widespread RNA & DNA sequence differences in the human transcriptome”. Science 335, 1302 (2012).
Teytelman, L., Thurtle, D. M., Rine, J. & van Oudenaarden, A. Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins. Proc. Natl Acad. Sci. USA 110, 18602–18607 (2013).
Wang, J. et al. Sequence features and chromatin structure around the genomic regions bound by 119 human transcription factors. Genome Res. 22, 1798–1812 (2012).
Park, P. J. ChIP–seq: advantages and challenges of a maturing technology. Nature Rev. Genet. 10, 669–680 (2009).
Pickrell, J. K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464, 768–772 (2010).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning. (Springer, 2001).
Acknowledgements
The authors thank members of X.S.L and M. Brown's laboratories for their discussions. This work is supported by the US National Institutes of Health grant R01GM099409.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
FURTHER INFORMATION
Glossary
- ChIP–seq
-
(Chromatin immunoprecipitation followed by next-generation DNA sequencing). A method to identify DNA-associated protein-binding sites.
- MNase-seq
-
A method in which micrococcal nuclease (MNase) digestion of chromatin is followed by next-generation sequencing to identify loci of high nucleosome occupancy.
- FAIRE–seq
-
(Formaldehyde-assisted isolation of regulatory elements followed by sequencing). A method to determine regulatory regions of the genome.
- DNase-seq
-
A method in which DNase I digestion of chromatin is combined with next-generation sequencing to identify regulatory regions of the genome, including enhancers and promoters.
- Hi-C
-
An extension of chromosome conformation capture that uses next-generation sequencing to observe long-range interaction frequencies between different regions of the genome.
- ChIA-PET
-
(Chromatin interaction analysis by paired-end tag sequencing). A method that combines chromatin immunoprecipitation-based enrichment and chromatin proximity ligation with paired-end next-generation sequencing to determine genome-wide chromatin interactions.
- ATAC-seq
-
(Assay for transposase-accessible chromatin using sequencing). A method that combines next-generation sequencing with in vitro transposition of sequencing adapters into native chromatin.
- Random barcoding
-
A technique that ligates a diverse assortment of short random DNA sequences to an unamplified DNA sample, which can be used to distinguish duplicates produced by PCR from those originating from the unamplified DNA.
- Spike-in
-
Controls that are known quantities of readily identifiable nucleic acids, which are added to a sample prior to critical steps in an experimental protocol. Such controls may be used for bias assessment and calibration purposes.
- Splines
-
Flexible smooth nonlinear functions that are defined piecewise by polynomials for fitting nonlinear trends.
- Locally estimated scatterplot smoothing
-
(LOESS). A simple yet robust method for fitting nonlinear trends.
- Quantile regression
-
A statistical regression method that estimates the median or other quantile of the response variables and that is robust against outliers.
- Surrogate variable analysis
-
A statistical analysis to identify and model variables that are not explicitly annotated but that have measureable effects.
Rights and permissions
About this article
Cite this article
Meyer, C., Liu, X. Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. Nat Rev Genet 15, 709–721 (2014). https://doi.org/10.1038/nrg3788
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrg3788
This article is cited by
-
RGT: a toolbox for the integrative analysis of high throughput regulatory genomics data
BMC Bioinformatics (2023)
-
Cell-specific and shared regulatory elements control a multigene locus active in mammary and salivary glands
Nature Communications (2023)
-
Nanopore microscope identifies RNA isoforms with structural colours
Nature Chemistry (2022)
-
Intrinsic bias estimation for improved analysis of bulk and single-cell chromatin accessibility profiles using SELMA
Nature Communications (2022)
-
NUCOME: A comprehensive database of nucleosome organization referenced landscapes in mammalian genomes
BMC Bioinformatics (2021)