Key Points
-
Mature microRNAs (miRNAs) are single-stranded RNAs that are 19–24 nucleotides in length and are produced in a multistep process involving the ribonuclease enzymes Drosha and Dicer.
-
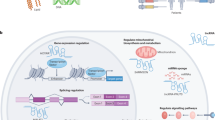
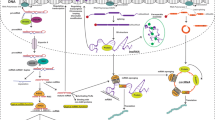
miRNAs act via diverse mechanisms. In the 'canonical' mechanism, miRNAs bind to the 3′ untranslated (3′ UTR) region of mRNAs and reduce their protein output. Several unexpected mechanisms — such as miRNA binding to other RNA regions or DNA regulatory elements — and direct or indirect upregulation of protein translation have been demonstrated recently.
-
Extracellular miRNAs such as those secreted in exosomes can act on recipient cells through hormone-like mechanisms.
-
miRNAs are drivers or cofactors of carcinogenesis as well as tumour metastasis by controlling the expression of multiple protein-coding genes.
-
Aberrant miRNA expression in cancer can be corrected by replacement using miRNA mimics or blocking with anti-miR approaches.
-
The main advantages of miRNA therapeutics are their multi-targeting effects. Despite substantial challenges associated with miRNA therapeutics, the strategies of replacing tumour suppressor miRNAs with mimics have generated the first miRNA therapeutic agent (MRX34), which is in clinical trials for the treatment of cancer.
-
The combination of miRNAs with chemotherapeutic drugs or small interfering RNAs may synergistically improve the anticancer therapeutic efficacy and can be developed for cancer treatment.
-
Long non-coding RNAs (lncRNAs), including long intergenic ncRNAs (lincRNAs), transcribed ultraconserved regions (T-UCRs) and natural antisense transcripts (NATs), have been found to be involved in human cancers.
-
Many of the lncRNAs are predominately localized in the nucleus and thus act through different mechanisms than miRNAs. The unique features of such lncRNAs can be exploited for the development of specific and novel therapeutic strategies against cancer.
Abstract
The first cancer-targeted microRNA (miRNA) drug — MRX34, a liposome-based miR-34 mimic — entered Phase I clinical trials in patients with advanced hepatocellular carcinoma in April 2013, and miRNA therapeutics are attracting special attention from both academia and biotechnology companies. Although miRNAs are the most studied non-coding RNAs (ncRNAs) to date, the importance of long non-coding RNAs (lncRNAs) is increasingly being recognized. Here, we summarize the roles of miRNAs and lncRNAs in cancer, with a focus on the recently identified novel mechanisms of action, and discuss the current strategies in designing ncRNA-targeting therapeutics, as well as the associated challenges.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mattick, J. S. The genetic signatures of noncoding RNAs. PLoS Genet. 5, e1000459 (2009).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Calin, G. A. et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl Acad. Sci. USA 99, 15524–15529 (2002).
Mendell, J. T. & Olson, E. N. MicroRNAs in stress signaling and human disease. Cell 148, 1172–1187 (2012).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs — microRNAs with a role in cancer. Nature Rev. Cancer 6, 259–269 (2006).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Volinia, S. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc. Natl Acad. Sci. USA 103, 2257–2261 (2006).
Esteller, M. Non-coding RNAs in human disease. Nature Rev. Genet. 12, 861–874 (2011).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nature Rev. Cancer 6, 857–866 (2006).
Ambros, V. MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 113, 673–676 (2003).
Croce, C. M. Causes and consequences of microRNA dysregulation in cancer. Nature Rev. Genet. 10, 704–714 (2009).
Spizzo, R., Almeida, M. I., Colombatti, A. & Calin, G. A. Long non-coding RNAs and cancer: a new frontier of translational research? Oncogene 31, 4577–4587 (2012).
Pennisi, E. Genomics. ENCODE project writes eulogy for junk DNA. Science 337, 1159–1161 (2012).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Fabian, M. R., Sonenberg, N. & Filipowicz, W. Regulation of mRNA translation and stability by microRNAs. Annu. Rev. Biochem. 79, 351–379 (2010).
Garzon, R., Marcucci, G. & Croce, C. M. Targeting microRNAs in cancer: rationale, strategies and challenges. Nature Rev. Drug Discov. 9, 775–789 (2010).
Thorsen, S. B., Obad, S., Jensen, N. F., Stenvang, J. & Kauppinen, S. The therapeutic potential of microRNAs in cancer. Cancer J. 18, 275–284 (2012).
van Kouwenhove, M., Kedde, M. & Agami, R. MicroRNA regulation by RNA-binding proteins and its implications for cancer. Nature Rev. Cancer 11, 644–656 (2011).
van Rooij, E. & Olson, E. N. MicroRNA therapeutics for cardiovascular disease: opportunities and obstacles. Nature Rev. Drug Discov. 11, 860–872 (2012).
Wahlestedt, C. Targeting long non-coding RNA to therapeutically upregulate gene expression. Nature Rev. Drug Discov. 12, 433–446 (2013).
Tay, Y., Zhang, J., Thomson, A. M., Lim, B. & Rigoutsos, I. MicroRNAs to Nanog, Oct4 and Sox2 coding regions modulate embryonic stem cell differentiation. Nature 455, 1124–1128 (2008).
Lytle, J. R., Yario, T. A. & Steitz, J. A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc. Natl Acad. Sci. USA 104, 9667–9672 (2007).
Vasudevan, S., Tong, Y. & Steitz, J. A. Switching from repression to activation: microRNAs can up-regulate translation. Science 318, 1931–1934 (2007).
Eiring, A. M. et al. miR-328 functions as an RNA decoy to modulate hnRNP E2 regulation of mRNA translation in leukemic blasts. Cell 140, 652–665 (2010).
Orom, U. A., Nielsen, F. C. & Lund, A. H. MicroRNA-10a binds the 5′UTR of ribosomal protein mRNAs and enhances their translation. Mol. Cell 30, 460–471 (2008).
Fabbri, M. et al. MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. Proc. Natl Acad. Sci. USA 109, E2110–E2116 (2012).
Mitchell, P. S. et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl Acad. Sci. USA 105, 10513–10518 (2008).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl Acad. Sci. USA 108, 5003–5008 (2011).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biol. 9, 654–659 (2007).
Vickers, K. C., Palmisano, B. T., Shoucri, B. M., Shamburek, R. D. & Remaley, A. T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nature Cell Biol. 13, 423–433 (2011).
Pigati, L. et al. Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS ONE 5, e13515 (2010).
Cortez, M. A. et al. MicroRNAs in body fluids — the mix of hormones and biomarkers. Nature Rev. Clin. Oncol. 8, 467–477 (2011).
Kasinski, A. L. & Slack, F. J. Epigenetics and genetics. MicroRNAs en route to the clinic: progress in validating and targeting microRNAs for cancer therapy. Nature Rev. Cancer 11, 849–864 (2011).
Spizzo, R., Nicoloso, M. S., Croce, C. M. & Calin, G. A. SnapShot: microRNAs in cancer. Cell 137, 586–586e1 (2009).
Volinia, S. et al. Reprogramming of miRNA networks in cancer and leukemia. Genome Res. 20, 589–599 (2010).
Medina, P. P., Nolde, M. & Slack, F. J. OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma. Nature 467, 86–90 (2010).
Hatley, M. E. et al. Modulation of K-Ras-dependent lung tumorigenesis by microRNA-21. Cancer Cell 18, 282–293 (2010).
Frankel, L. B. et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J. Biol. Chem. 283, 1026–1033 (2008).
He, L. et al. A microRNA polycistron as a potential human oncogene. Nature 435, 828–833 (2005).
Mu, P. et al. Genetic dissection of the miR-17∼92 cluster of microRNAs in Myc-induced B-cell lymphomas. Genes Dev. 23, 2806–2811 (2009).
Costinean, S. et al. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in Eμ-miR155 transgenic mice. Proc. Natl Acad. Sci. USA 103, 7024–7029 (2006).
Nicoloso, M. S., Spizzo, R., Shimizu, M., Rossi, S. & Calin, G. A. MicroRNAs — the micro steering wheel of tumour metastases. Nature Rev. Cancer 9, 293–302 (2009).
Pencheva, N. & Tavazoie, S. F. Control of metastatic progression by microRNA regulatory networks. Nature Cell Biol. 15, 546–554 (2013).
Png, K. J., Halberg, N., Yoshida, M. & Tavazoie, S. F. A microRNA regulon that mediates endothelial recruitment and metastasis by cancer cells. Nature 481, 190–194 (2012).
Song, S. J. et al. MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell 154, 311–324 (2013).
Pineau, P. et al. miR-221 overexpression contributes to liver tumorigenesis. Proc. Natl Acad. Sci. USA 107, 264–269 (2010).
Felli, N. et al. MicroRNAs 221 and 222 inhibit normal erythropoiesis and erythroleukemic cell growth via kit receptor down-modulation. Proc. Natl Acad. Sci. USA 102, 18081–18086 (2005).
Fabbri, M. et al. Association of a microRNA/TP53 feedback circuitry with pathogenesis and outcome of B-cell chronic lymphocytic leukemia. JAMA 305, 59–67 (2011).
Mavrakis, K. J. et al. A cooperative microRNA-tumor suppressor gene network in acute T-cell lymphoblastic leukemia (T-ALL). Nature Genet. 43, 673–678 (2011).
O'Donnell, K. A., Wentzel, E. A., Zeller, K. I., Dang, C. V. & Mendell, J. T. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435, 839–843 (2005).
Uhlmann, S. et al. Global microRNA level regulation of EGFR-driven cell-cycle protein network in breast cancer. Mol. Syst. Biol. 8, 570 (2012).
Wheeler, D. L., Dunn, E. F. & Harari, P. M. Understanding resistance to EGFR inhibitors — impact on future treatment strategies. Nature Rev. Clin. Oncol. 7, 493–507 (2010).
Calin, G. A. et al. MiR-15a and miR-16-1 cluster functions in human leukemia. Proc. Natl Acad. Sci. USA 105, 5166–5171 (2008).
Calin, G. A. et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N. Engl. J. Med. 353, 1793–1801 (2005).
Almeida, M. I. et al. Strand-specific miR-28-5p and miR-28-3p have distinct effects in colorectal cancer cells. Gastroenterology 142, 886–896e9 (2012).
Girardot, M. et al. miR-28 is a thrombopoietin receptor targeting microRNA detected in a fraction of myeloproliferative neoplasm patient platelets. Blood 116, 437–445 (2010).
Jiang, L. et al. Hsa-miR-125a-3p and hsa-miR-125a-5p are downregulated in non-small cell lung cancer and have inverse effects on invasion and migration of lung cancer cells. BMC Cancer 10, 318 (2010).
Galm, O., Herman, J. G. & Baylin, S. B. The fundamental role of epigenetics in hematopoietic malignancies. Blood Rev. 20, 1–13 (2006).
Lujambio, A. et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 67, 1424–1429 (2007).
Bandres, E. et al. Epigenetic regulation of microRNA expression in colorectal cancer. Int. J. Cancer 125, 2737–2743 (2009).
Saito, Y. & Jones, P. A. Epigenetic activation of tumor suppressor microRNAs in human cancer cells. Cell Cycle 5, 2220–2222 (2006).
Melo, S. et al. Small molecule enoxacin is a cancer-specific growth inhibitor that acts by enhancing TAR RNA-binding protein 2-mediated microRNA processing. Proc. Natl Acad. Sci. USA 108, 4394–4399 (2011).
Bader, A. G., Brown, D., Stoudemire, J. & Lammers, P. Developing therapeutic microRNAs for cancer. Gene Ther. 18, 1121–1126 (2011).
Tivnan, A. et al. Inhibition of neuroblastoma tumor growth by targeted delivery of microRNA-34a using anti-disialoganglioside GD2 coated nanoparticles. PLoS ONE 7, e38129 (2012).
Trang, P. et al. Systemic delivery of tumor suppressor microRNA mimics using a neutral lipid emulsion inhibits lung tumors in mice. Mol. Ther. 19, 1116–1122 (2011).
Tazawa, H., Tsuchiya, N., Izumiya, M. & Nakagama, H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc. Natl Acad. Sci. USA 104, 15472–15477 (2007).
Ji, J. et al. MicroRNA expression, survival, and response to interferon in liver cancer. N. Engl. J. Med. 361, 1437–1447 (2009).
Kota, J. et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137, 1005–1017 (2009).
Chen, L. et al. Tumor-specific expression of microRNA-26a suppresses human hepatocellular carcinoma growth via cyclin-dependent and -independent pathways. Mol. Ther. 19, 1521–1528 (2011).
Bader, A. G. miR-34 — a microRNA replacement therapy is headed to the clinic. Front. Genet. 3, 120 (2012).
Lennox, K. A. & Behlke, M. A. Chemical modification and design of anti-miRNA oligonucleotides. Gene Ther. 18, 1111–1120 (2011).
Swarbrick, A. et al. miR-380-5p represses p53 to control cellular survival and is associated with poor outcome in MYCN-amplified neuroblastoma. Nature Med. 16, 1134–1140 (2010).
Janssen, H. L. et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 368, 1685–1694 (2013).
Lieberman, J. & Sarnow, P. Micromanaging hepatitis C virus. N. Engl. J. Med. 368, 1741–1743 (2013).
Jopling, C. L., Yi, M., Lancaster, A. M., Lemon, S. M. & Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific microRNA. Science 309, 1577–1581 (2005).
Obad, S. et al. Silencing of microRNA families by seed-targeting tiny LNAs. Nature Genet. 43, 371–378 (2011).
Zhang, Y. et al. LNA-mediated anti-miR-155 silencing in low-grade B-cell lymphomas. Blood 120, 1678–1686 (2012).
Elmen, J. et al. LNA-mediated microRNA silencing in non-human primates. Nature 452, 896–899 (2008).
Krutzfeldt, J. et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature 438, 685–689 (2005).
Ma, L. et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nature Biotech. 28, 341–347 (2010).
Ebert, M. S., Neilson, J. R. & Sharp, P. A. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nature Methods 4, 721–726 (2007).
Valastyan, S. et al. A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137, 1032–1046 (2009).
Ma, L. et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nature Cell Biol. 12, 247–256 (2010).
Kluiver, J. et al. Rapid generation of microRNA sponges for microRNA inhibition. PLoS ONE 7, e29275 (2012).
Xie, J. et al. Long-term, efficient inhibition of microRNA function in mice using rAAV vectors. Nature Methods 9, 403–409 (2012).
Davis, S. et al. Potent inhibition of microRNA in vivo without degradation. Nucleic Acids Res. 37, 70–77 (2009).
Dvinge, H. et al. The shaping and functional consequences of the microRNA landscape in breast cancer. Nature 497, 378–382 (2013).
Gumireddy, K. et al. Small-molecule inhibitors of microrna miR-21 function. Angew. Chem. Int. Ed Engl. 47, 7482–7484 (2008).
Young, D. D., Connelly, C. M., Grohmann, C. & Deiters, A. Small molecule modifiers of microRNA miR-122 function for the treatment of hepatitis C virus infection and hepatocellular carcinoma. J. Am. Chem. Soc. 132, 7976–7981 (2010).
Zhang, S., Chen, L., Jung, E. J. & Calin, G. A. Targeting microRNAs with small molecules: from dream to reality. Clin. Pharmacol. Ther. 87, 754–758 (2010).
Lehmann, S. M. et al. An unconventional role for miRNA: let-7 activates Toll-like receptor 7 and causes neurodegeneration. Nature Neurosci. 15, 827–835 (2012).
O'Brien, S. et al. Randomized phase III trial of fludarabine plus cyclophosphamide with or without oblimersen sodium (Bcl-2 antisense) in patients with relapsed or refractory chronic lymphocytic leukemia. J. Clin. Oncol. 25, 1114–1120 (2007).
Jiang, K. Biotech comes to its 'antisenses' after hard-won drug approval. Nature Med. 19, 252 (2013).
Ding, L. et al. Combined transfection of Bcl-2 siRNA and miR-15a oligonucleotides enhanced methotrexate-induced apoptosis in Raji cells. Cancer Biol. Med. 10, 16–21 (2013).
Hu, X. et al. The effect of Bcl-2 siRNA combined with miR-15a oligonucleotides on the growth of Raji cells. Med. Oncol. 30, 430 (2013).
Pieters, R. et al. A treatment protocol for infants younger than 1 year with acute lymphoblastic leukaemia (Interfant-99): an observational study and a multicentre randomised trial. Lancet 370, 240–250 (2007).
Kotani, A. et al. miR-128b is a potent glucocorticoid sensitizer in MLL–AF4 acute lymphocytic leukemia cells and exerts cooperative effects with miR-221. Blood 114, 4169–4178 (2009).
Kotani, A. et al. A novel mutation in the miR-128b gene reduces miRNA processing and leads to glucocorticoid resistance of MLL-AF4 acute lymphocytic leukemia cells. Cell Cycle 9, 1037–1042 (2010).
Bockhorn, J. et al. MicroRNA-30c inhibits human breast tumour chemotherapy resistance by regulating TWF1 and IL-11. Nature Commun. 4, 1393 (2013).
Kim, S. J. et al. Development of microRNA-145 for therapeutic application in breast cancer. J. Control Release 155, 427–434 (2011).
Corsten, M. F. et al. MicroRNA-21 knockdown disrupts glioma growth in vivo and displays synergistic cytotoxicity with neural precursor cell delivered S-TRAIL in human gliomas. Cancer Res. 67, 8994–9000 (2007).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Faghihi, M. A. & Wahlestedt, C. Regulatory roles of natural antisense transcripts. Nature Rev. Mol. Cell Biol. 10, 637–643 (2009).
Calin, G. A. et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12, 215–229 (2007).
Orom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Gupta, R. A. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071–1076 (2010).
Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007).
Wang, K. C. & Chang, H. Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl Acad. Sci. USA 106, 11667–11672 (2009).
Ji, P. et al. MALAT-1, a novel noncoding RNA, and thymosin β4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22, 8031–8041 (2003).
Huarte, M. et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142, 409–419 (2010).
Faghihi, M. A. & Wahlestedt, C. RNA interference is not involved in natural antisense mediated regulation of gene expression in mammals. Genome Biol. 7, R38 (2006).
Ling, H. et al. CCAT2, a novel non-coding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 23, 1446–1461 (2013).
Salmena, L., Poliseno, L., Tay, Y., Kats, L. & Pandolfi, P. P. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146, 353–358 (2011).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Hung, T. et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nature Genet. 43, 621–629 (2011).
Wheeler, T. M. et al. Targeting nuclear RNA for in vivo correction of myotonic dystrophy. Nature 488, 111–115 (2012).
Li, C. H. & Chen, Y. Targeting long non-coding RNAs in cancers: progress and prospects. Int. J. Biochem. Cell Biol. 45, 1895–1910 (2013).
Gutschner, T. et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 73, 1180–1189 (2013).
Kino, T., Hurt, D. E., Ichijo, T., Nader, N. & Chrousos, G. P. Noncoding RNA Gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal. 3, ra8 (2010).
Rusk, N. AntagoNATs boost gene expression. Nature Methods 9, 437 (2012).
Modarresi, F. et al. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nature Biotech. 30, 453–459 (2012).
Kim, D. H. & Rossi, J. J. Strategies for silencing human disease using RNA interference. Nature Rev. Genet. 8, 173–184 (2007).
Pecot, C. V., Calin, G. A., Coleman, R. L., Lopez-Berestein, G. & Sood, A. K. RNA interference in the clinic: challenges and future directions. Nature Rev. Cancer 11, 59–67 (2011).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nature Rev. Drug Discov. 8, 129–138 (2009).
Broderick, J. A. & Zamore, P. D. MicroRNA therapeutics. Gene Ther. 18, 1104–1110 (2011).
Pai, S. I. et al. Prospects of RNA interference therapy for cancer. Gene Ther. 13, 464–477 (2006).
Flemming, A. Regulatory watch: pioneering gene therapy on brink of approval. Nature Rev. Drug Discov. 11, 664 (2012).
Gruber, K. Europe gives gene therapy the green light. Lancet 380, e10 (2012).
de Pontual, L. et al. Germline deletion of the miR-17 approximately 92 cluster causes skeletal and growth defects in humans. Nature Genet. 43, 1026–1030 (2011).
Okamura, K. & Lai, E. C. Endogenous small interfering RNAs in animals. Nature Rev. Mol. Cell Biol. 9, 673–678 (2008).
Kim, V. N. Small RNAs just got bigger: Piwi-interacting RNAs (piRNAs) in mammalian testes. Genes Dev. 20, 1993–1997 (2006).
Kiss, T. Small nucleolar RNAs: an abundant group of noncoding RNAs with diverse cellular functions. Cell 109, 145–148 (2002).
Taft, R. J. et al. Small RNAs derived from snoRNAs. RNA 15, 1233–1240 (2009).
Taft, R. J. et al. Tiny RNAs associated with transcription start sites in animals. Nature Genet. 41, 572–578 (2009).
Langenberger, D. et al. Evidence for human microRNA-offset RNAs in small RNA sequencing data. Bioinformatics 25, 2298–2301 (2009).
Wang, X. J., Gaasterland, T. & Chua, N. H. Genome-wide prediction and identification of cis-natural antisense transcripts in Arabidopsis thaliana. Genome Biol. 6, R30 (2005).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Lai, F. et al. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 494, 497–501 (2013).
Hwang, H. W., Wentzel, E. A. & Mendell, J. T. A hexanucleotide element directs microRNA nuclear import. Science 315, 97–100 (2007).
Forman, J. J., Legesse-Miller, A. & Coller, H. A. A search for conserved sequences in coding regions reveals that the let-7 microRNA targets Dicer within its coding sequence. Proc. Natl Acad. Sci. USA 105, 14879–14884 (2008).
Place, R. F., Li, L. C., Pookot, D., Noonan, E. J. & Dahiya, R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc. Natl Acad. Sci. USA 105, 1608–1613 (2008).
Redis, R. S., Calin, S., Yang, Y., You, M. J. & Calin, G. A. Cell-to-cell miRNA transfer: from body homeostasis to therapy. Pharmacol. Ther. 136, 169–174 (2012).
Yang, M. et al. Microvesicles secreted by macrophages shuttle invasion-potentiating microRNAs into breast cancer cells. Mol. Cancer 10, 117 (2011).
Liu, Y. et al. Contribution of MyD88 to the tumor exosome-mediated induction of myeloid derived suppressor cells. Am. J. Pathol. 176, 2490–2499 (2010).
Fabbri, M. et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl Acad. Sci. USA 104, 15805–15810 (2007).
Calin, G. A. et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl Acad. Sci. USA 101, 2999–3004 (2004).
Klein, U. et al. The DLEU2/miR-15a/16-1 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell 17, 28–40 (2010).
Wojcik, S. E. et al. Non-codingRNA sequence variations in human chronic lymphocytic leukemia and colorectal cancer. Carcinogenesis 31, 208–215 (2010).
Fabbri, M., Valeri, N. & Calin, G. A. MicroRNAs and genomic variations: from Proteus tricks to Prometheus gift. Carcinogenesis 30, 912–917 (2009).
Fabbri, M. MicroRNAs and cancer epigenetics. Curr. Opin. Investig. Drugs 9, 583–590 (2008).
Chen, X. et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 18, 997–1006 (2008).
Zhang, H. L. et al. Serum miRNA-21: elevated levels in patients with metastatic hormone-refractory prostate cancer and potential predictive factor for the efficacy of docetaxel-based chemotherapy. Prostate 71, 326–331 (2011).
Cheng, H. et al. Circulating plasma miR-141 is a novel biomarker for metastatic colon cancer and predicts poor prognosis. PLoS ONE 6, e17745 (2011).
Ma, L., Teruya-Feldstein, J. & Weinberg, R. A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449, 682–688 (2007).
Cimmino, A. et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl Acad. Sci. USA 102, 13944–13949 (2005).
Dews, M. et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nature Genet. 38, 1060–1065 (2006).
Mendell, J. T. miRiad roles for the miR-17-92 cluster in development and disease. Cell 133, 217–222 (2008).
Chan, J. A., Krichevsky, A. M. & Kosik, K. S. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 65, 6029–6033 (2005).
Meng, F. et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133, 647–658 (2007).
Chang, T. C. et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell 26, 745–752 (2007).
He, L. et al. A microRNA component of the p53 tumour suppressor network. Nature 447, 1130–1134 (2007).
Raver-Shapira, N. et al. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol. Cell 26, 731–743 (2007).
Liu, C. et al. The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nature Med. 17, 211–215 (2011).
Yang, S. et al. MicroRNA-34 suppresses breast cancer invasion and metastasis by directly targeting Fra-1. Oncogene 32, 4294–4303 (2012).
Eis, P. S. et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc. Natl Acad. Sci. USA 102, 3627–3632 (2005).
Tavazoie, S. F. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008).
Huang, Q. et al. The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nature Cell Biol. 10, 202–210 (2008).
Johnson, S. M. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005).
Sampson, V. B. et al. MicroRNA let-7a down-regulates MYC and reverts MYC-induced growth in Burkitt lymphoma cells. Cancer Res. 67, 9762–9770 (2007).
Tsang, W. P. & Kwok, T. T. Let-7a microRNA suppresses therapeutics-induced cancer cell death by targeting caspase-3. Apoptosis 13, 1215–1222 (2008).
Zalfa, F. et al. Fragile X mental retardation protein (FMRP) binds specifically to the brain cytoplasmic RNAs BC1/BC200 via a novel RNA-binding motif. J. Biol. Chem. 280, 33403–33410 (2005).
Muddashetty, R. et al. Poly(A)-binding protein is associated with neuronal BC1 and BC200 ribonucleoprotein particles. J. Mol. Biol. 321, 433–445 (2002).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010).
Wilusz, J. E., Freier, S. M. & Spector, D. L. 3′ end processing of a long nuclear-retained noncoding RNA yields a tRNA-like cytoplasmic RNA. Cell 135, 919–932 (2008).
Crawford, E. D. et al. Diagnostic performance of PCA3 to detect prostate cancer in men with increased prostate specific antigen: a prospective study of 1,962 cases. J. Urol. 188, 1726–1731 (2012).
Dias, N. & Stein, C. A. Antisense oligonucleotides: basic concepts and mechanisms. Mol. Cancer Ther. 1, 347–355 (2002).
Novina, C. D. & Sharp, P. A. The RNAi revolution. Nature 430, 161–164 (2004).
Acknowledgements
G.A.C. is The Alan M. Gewirtz Leukemia & Lymphoma Society Scholar. He is also supported as a Fellow at The University of Texas MD Anderson Research Trust, as a University of Texas System Regents Research Scholar and by the CLL Global Research Foundation. Work in G.A.C.'s laboratory is supported in part by the US National Institutes of Health (NIH)/US National Cancer Institute (NCI) grant CA135444; a US Department of Defense Breast Cancer Idea Award; Developmental Research Awards in breast cancer, ovarian cancer, brain cancer, prostate cancer, multiple myeloma and leukaemia (P50 CA100632) as well as head and neck cancer (P50 CA097007) from the Specialized Programs of Research Excellence (SPOREs); a Sister Institution Network Fund (SINF) grant from the MD Anderson Cancer Center and the German Cancer Research Center (DKFZ) in chronic lymphocytic leukaemia; a SINF grant in colorectal cancer, the Laura and John Arnold Foundation; the RGK Foundation; and The Estate of C. G. Johnson, Jr. H.L. is an Odyssey Fellow and his work is supported in part by the Odyssey Program and The Estate of C. G. Johnson, Jr. M.F.'s laboratory is supported by the Jean Perkins Foundation, by Nautica Malibu Triathlon Funds, by the Pablove Foundation, by the St. Baldrick's Foundation, by the Southern California Clinical and Translational Science Institute, by Funds from the Saban Research Institute, by award number P30CA014089 from the NCI, by the Hugh and Audy Lou Colvin Foundation, and by the T. J. Martell Foundation. The authors apologize to all colleagues whose work was not cited because of space restrictions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Non-coding RNAs
-
(ncRNAs). RNA molecules that are not translated into protein.
- miRNA cluster
-
A group of at least two microRNAs located close together on the genome (usually several hundreds of bases apart) that are generally transcribed in a unique transcript and also commonly regulated.
Rights and permissions
About this article
Cite this article
Ling, H., Fabbri, M. & Calin, G. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov 12, 847–865 (2013). https://doi.org/10.1038/nrd4140
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd4140
This article is cited by
-
Potential of oligonucleotide- and protein/peptide-based therapeutics in the management of toxicant/stressor-induced diseases
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
Roles of tumor-associated macrophages in anti-PD-1/PD-L1 immunotherapy for solid cancers
Molecular Cancer (2023)
-
Recent advances on high-efficiency of microRNAs in different types of lung cancer: a comprehensive review
Cancer Cell International (2023)
-
The role of noncoding RNAs in metabolic reprogramming of cancer cells
Cellular & Molecular Biology Letters (2023)
-
MicroRNA: trends in clinical trials of cancer diagnosis and therapy strategies
Experimental & Molecular Medicine (2023)