Abstract
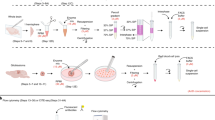
Immune cells comprise a diverse and dynamic cell population that is responsible for a broad range of immunological activities. They act in concert with other immune and nonimmune cells via cytokine-mediated communication and direct cell–cell interactions. Understanding the complex immune network requires a broad characterization of its individual cellular components. This is especially relevant for the brain compartment, which is an active immunological site, composed of resident and infiltrating immune cells that affect brain development, tissue homeostasis and neuronal activity. Mass cytometry, or CyTOF (cytometry by time-of-flight), uses metal-conjugated antibodies to enable a high-dimensional description of tens of markers at the single-cell level, thereby providing a bird's-eye view of the immune system. This technique has been successfully applied to the discovery of novel immune populations in humans and rodents. Here, we provide a step-by-step description of a mass cytometry approach for the analysis of the mouse brain compartment. The different stages of the procedure include brain perfusion, extraction of the brain tissue and its dissociation into a single-cell suspension, followed by cell staining with metal-tagged antibodies, sample reading using a mass cytometer, and data analysis using SPADE and viSNE. This procedure takes <5 h (excluding data analysis) and can be applied to study modifications in the brain's immune populations under normal and pathological conditions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Paolicelli, R.C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Welberg, L. Synaptic plasticity: a synaptic role for microglia. Nat. Rev. Neurosci. 15, 68–69 (2014).
Wu, Y., Dissing-Olesen, L., MacVicar, B.A. & Stevens, B. Microglia: dynamic mediators of synapse development and plasticity. Trends Immunol. 36, 605–613 (2015).
Iori, V., Frigerio, F. & Vezzani, A. Modulation of neuronal excitability by immune mediators in epilepsy. Curr. Opin. Pharmacol. 26, 118–123 (2016).
Ransohoff, R.M., Schafer, D., Vincent, A., Blachère, N.E. & Bar-Or, A. Neuroinflammation: ways in which the immune system affects the brain. Neurotherapeutics 12, 896–909 (2015).
Lampron, A., Pimentel-Coelho, P.M. & Rivest, S. Migration of bone marrow-derived cells into the central nervous system in models of neurodegeneration. J. Comp. Neurol. 521, 3863–3876 (2013).
Dendrou, C.A., Fugger, L. & Friese, M.A. Immunopathology of multiple sclerosis. Nat. Rev. Immunol. 15, 545–558 (2015).
Heppner, F.L., Ransohoff, R.M. & Becher, B. Immune attack: the role of inflammation in Alzheimer disease. Nat. Rev. Neurosci. 16, 358–372 (2015).
Baruch, K. et al. PD-1 immune checkpoint blockade reduces pathology and improves memory in mouse models of Alzheimer's disease. Nat. Med. 22, 135–137 (2016).
Teeling, J.L. & Asuni, A.A. Immune to brain communication in health, age and disease: implications for understanding age-related neurodegeneration. in The Ageing Immune System and Health 125–139 (Springer, 2017).
Arlehamn, C.S.L. et al. Immune response in Parkinson's disease driven by HLA display of α-synuclein peptides. J. Immunol. 198, 55.26 (2017).
Baruch, K. et al. Breaking immune tolerance by targeting Foxp3+ regulatory T cells mitigates Alzheimer's disease pathology. Nat. Commun. 6, 7967 (2015).
Stern, J.N.H. et al. B cells populating the multiple sclerosis brain mature in the draining cervical lymph nodes. Sci. Transl. Med. 6, 248ra107 (2014).
Pikor, N.B., Prat, A., Bar-Or, A. & Gommerman, J.L. Meningeal tertiary lymphoid tissues and multiplesclerosis: a gathering place for diverse types of immune cells during CNS autoimmunity. Front. Immunol. 6, 2015.00657 (2016).
Mosley, R.L., Hutter-Saunders, J.A., Stone, D.K. & Gendelman, H.E. Inflammation and adaptive immunity in Parkinson's disease. Cold Spring Harb. Perspect. Med. 2, a009381 (2012).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Grabert, K. et al. Microglial brain region-dependent diversity and selective regional sensitivities to aging. Nat. Neurosci. 19, 504–516 (2016).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer's disease. Cell 169, 1276–1290.e17 (2017).
Cheung, R.K. & Utz, P.J. Screening: CyTOF—the next generation of cell detection. Nat. Rev. Rheumatol. 7, 502–503 (2011).
Bendall, S.C. et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell 157, 714–725 (2014).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Simoni, Y. et al. Human innate lymphoid cell subsets possess tissue-type based heterogeneity in phenotype and frequency. Immunity 46, 148–161 (2017).
See, P. et al. Mapping the human DC lineage through the integration of high-dimensional techniques. Science 356, eaag3009 (2017).
Yao, Y. & Montgomery, R.R. Role of immune aging in susceptibility to West Nile Virus. Methods Mol. Biol. 1435, 235–247 (2016).
Whiting, C.C. et al. Large-scale and comprehensive immune profiling and functional analysis of normal human aging. PLoS One 10, e0133627 (2015).
Baughn, L.B. et al. Phenotypic and functional characterization of a bortezomib-resistant multiple myeloma cell line by flow and mass cytometry. Leuk. Lymphoma 58, 1931–1940 (2017).
Aquino-López, A., Senyukov, V.V., Vlasic, Z., Kleinerman, E.S. & Lee, D.A. Interferon gamma induces changes in natural killer (NK) cell ligand expression and alters NK cell-mediated lysis of pediatric cancer cell lines. Front. Immunol. 8, 391 (2017).
Korin, B. et al. High-dimensional, single-cell characterization of the brain's immune compartment. Nat. Neurosci. 20, 1300–1309 (2017).
Becher, B. et al. High-dimensional analysis of the murine myeloid cell system. Nat. Immunol. 15, 1181–1189 (2014).
Mrdjen, D., Hartmann, F. & Becher, B. High dimensional cytometry of central nervous system leukocytes during neuroinflammation. in Inflammation (eds. E. Clausen, B., Laman, J. D. & Clausen, B. E.) 321–332 (Springer, 2017).
Garcia, J.A., Cardona, S.M. & Cardona, A.E. Isolation and analysis of mouse microglial cells. Curr. Protoc. Immunol. 104, Unit 14.35 (2014).
Beaudet, M.-J. et al. High yield extraction of pure spinal motor neurons, astrocytes and microglia from single embryo and adult mouse spinal cord. Sci. Rep. 5, srep16763 (2015).
Mills, K., McManus, R. & Dungan, L. Isolation and FACS analysis on mononuclear cells from CNS tissue. Bio-Protoc. 4(18), 1240 (2014).
Günther, R. et al. Clinical testing and spinal cord removal in a mouse model for amyotrophic lateral sclerosis (ALS). J. Vis. Exp. (61), e3936 (2012).
Pino, P.A. & Cardona, A.E. Isolation of brain and spinal cord mononuclear cells using Percoll gradients. J. Vis. Exp. (48), e2348 (2011).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Ofengeim, D., Giagtzoglou, N., Huh, D., Zou, C. & Yuan, J. Single-cell RNA sequencing: unraveling the brain one cell at a time. Trends Mol. Med. 23, 563–576 (2017).
Matcovitch-Natan, O. et al. Microglia development follows a stepwise program to regulate brain homeostasis. Science 353, aad8670 (2016).
Gerner, M.Y., Kastenmuller, W., Ifrim, I., Kabat, J. & Germain, R.N. Histo-cytometry: a method for highly multiplex quantitative tissue imaging analysis applied to dendritic cell subset microanatomy in lymph nodes. Immunity 37, 364–376 (2012).
Gerdes, M.J. et al. Highly multiplexed single-cell analysis of formalin-fixed, paraffin-embedded cancer tissue. Proc. Natl. Acad. Sci. USA 110, 11982–11987 (2013).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Kwon, S. Single-molecule fluorescence in situ hybridization: quantitative imaging of single RNA molecules. BMB Rep. 46, 65–72 (2013).
Chen, K.H., Boettiger, A.N., Moffitt, J.R., Wang, S. & Zhuang, X. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Shah, S., Lubeck, E., Zhou, W. & Cai, L. In situ transcription profiling of single cells reveals spatial organization of cells in the mouse hippocampus. Neuron 92, 342–357 (2016).
Bendall, S.C., Nolan, G.P., Roederer, M. & Chattopadhyay, P.K. A deep profiler's guide to cytometry. Trends Immunol. 33, 323–332 (2012).
Brodie, T.M. & Tosevski, V. High-dimensional single-cell analysis with mass cytometry. Curr. Protoc. Immunol. 118, 5.11.1–5.11.25 (2017).
McCarthy, R.L., Mak, D.H., Burks, J.K. & Barton, M.C. Rapid monoisotopic cisplatin based barcoding for multiplexed mass cytometry. Sci. Rep. 7, 3779 (2017).
Lai, L., Ong, R., Li, J. & Albani, S. A CD45-based barcoding approach to multiplex mass-cytometry (CyTOF). Cytometry A 87, 369–374 (2015).
Nassar, A.F., Wisnewski, A.V. & Raddassi, K. Automation of sample preparation for mass cytometry barcoding in support of clinical research: protocol optimization. Anal. Bioanal. Chem. 409, 2363–2372 (2017).
Bodenmiller, B. et al. Multiplexed mass cytometry profiling of cellular states perturbed by small-molecule regulators. Nat. Biotechnol. 30, 858–867 (2012).
Zunder, E.R. et al. Palladium-based mass tag cell barcoding with a doublet-filtering scheme and single-cell deconvolution algorithm. Nat. Protoc. 10, 316–333 (2015).
Elghetany, M.T. & Davis, B.H. Impact of preanalytical variables on granulocytic surface antigen expression: a review. Cytometry B Clin. Cytom. 65, 1–5 (2005).
Kappelmayer, J. et al. Flow cytometric detection of intracellular myeloperoxidase, CD3 and CD79a. Interaction between monoclonal antibody clones, fluorochromes and sample preparation protocols. J. Immunol. Methods 242, 53–65 (2000).
Fienberg, H.G., Simonds, E.F., Fantl, W.J., Nolan, G.P. & Bodenmiller, B. A platinum-based covalent viability reagent for single-cell mass cytometry. Cytometry A 81, 467–475 (2012).
Anderson, K.G. et al. Intravascular staining for discrimination of vascular and tissue leukocytes. Nat. Protoc. 9, 209–222 (2014).
Stern, A.D., Rahman, A.H. & Birtwistle, M.R. Cell size assays for mass cytometry. Cytometry A 91, 14–24 (2017).
Bennett, M.L. et al. New tools for studying microglia in the mouse and human CNS. Proc. Natl. Acad. Sci. USA 113, E1738–1746 (2016).
Butovsky, O. et al. Identification of a unique TGF-β-dependent molecular and functional signature in microglia. Nat. Neurosci. 17, 131–143 (2014).
Qiu, P. et al. Extracting a cellular hierarchy from high-dimensional cytometry data with SPADE. Nat. Biotechnol. 29, 886–891 (2011).
Anchang, B. et al. Visualization and cellular hierarchy inference of single-cell data using SPADE. Nat. Protoc. 11, 1264–1279 (2016).
Amir, E.D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013).
Samusik, N., Good, Z., Spitzer, M.H., Davis, K.L. & Nolan, G.P. Automated mapping of phenotype space with single-cell data. Nat. Methods 13, 493–496 (2016).
Weber, L.M. & Robinson, M.D. Comparison of clustering methods for high-dimensional single-cell flow and mass cytometry data. Cytometry A 89, 1084–1096 (2016).
Mair, F. et al. The end of gating? An introduction to automated analysis of high dimensional cytometry data. Eur. J. Immunol. 46, 34–43 (2016).
Bruggner, R.V., Bodenmiller, B., Dill, D.L., Tibshirani, R.J. & Nolan, G.P. Automated identification of stratifying signatures in cellular subpopulations. Proc. Natl. Acad. Sci. USA 111, E2770–E2777 (2014).
Van Gassen, S. et al. FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87, 636–645 (2015).
Newell, E.W. & Cheng, Y. Mass cytometry: blessed with the curse of dimensionality. Nat. Immunol. 17, 890–895 (2016).
Melchiotti, R., Gracio, F., Kordasti, S., Todd, A.K. & de Rinaldis, E. Cluster stability in the analysis of mass cytometry data. Cytometry A 91, 73–84 (2017).
Pennartz, S., Reiss, S., Biloune, R., Hasselmann, D. & Bosio, A. Generation of single-cell suspensions from mouse neural tissue. J. Vis. Exp. (29), e1267 (2009).
Njie, E.G. et al. Ex vivo cultures of microglia from young and aged rodent brain reveal age-related changes in microglial function. Neurobiol. Aging 33, 195.e1–195.e12 (2012).
Lee, J.-K. & Tansey, M.G. Microglia isolation from adult mouse brain. in Microglia 17–23 (Humana Press, 2013).
Martin, E., El-Behi, M., Fontaine, B. & Delarasse, C. Analysis of microglia and monocyte-derived macrophages from the central nervous system by flow cytometry. J. Vis. Exp. (124), e55781 (2017).
Nikodemova, M. & Watters, J.J. Efficient isolation of live microglia with preserved phenotypes from adult mouse brain. J. Neuroinflammation 9, 147 (2012).
Weigmann, B. et al. Isolation and subsequent analysis of murine lamina propria mononuclear cells from colonic tissue. Nat. Protoc. 2, 2307–2311 (2007).
Autengruber, A., Gereke, M., Hansen, G., Hennig, C. & Bruder, D. Impact of enzymatic tissue disintegration on the level of surface molecule expression and immune cell function. Eur. J. Microbiol. Immunol. 2, 112–120 (2012).
Ford, A.L., Foulcher, E., Goodsall, A.L. & Sedgwick, J.D. Tissue digestion with dispase substantially reduces lymphocyte and macrophage cell-surface antigen expression. J. Immunol. Methods 194, 71–75 (1996).
Derecki, N., Derecki, N. & Kipnis, J. Mouse meninges isolation for FACS. Protoc. Exch. http://dx.doi.org/10.1038/protex.2014.030 (2014).
Ornatsky, O. et al. Highly multiparametric analysis by mass cytometry. J. Immunol. Methods 361, 1–20 (2010).
Ornatsky, O.I. et al. Study of cell antigens and intracellular DNA by identification of element-containing labels and metallointercalators using inductively coupled plasma mass spectrometry. Anal. Chem. 80, 2539–2547 (2008).
Finck, R. et al. Normalization of mass cytometry data with bead standards. Cytometry A 83A, 483–494 (2013).
Leipold, M.D., Newell, E.W. & Maecker, H.T. Multiparameter phenotyping of human PBMCs using mass cytometry. Methods Mol. Biol. 1343, 81–95 (2015).
Kay, A.W., Strauss-Albee, D.M. & Blish, C.A. Application of mass cytometry (CyTOF) for functional and phenotypic analysis of natural killer cells. Methods Mol. Biol. 1441, 13–26 (2016).
Louveau, A. et al. Structural and functional features of central nervous system lymphatic vessels. Nature 523, 337–341 (2015).
Nassar, A.F., Wisnewski, A.V. & Raddassi, K. Automation of sample preparation for mass cytometry barcoding in support of clinical research: protocol optimization. Anal. Bioanal. Chem. 409, 1–10 (2017).
Kaiser, O. et al. Dissociated neurons and glial cells derived from rat inferior colliculi after digestion with papain. PLoS ONE 8, e80490 (2013).
Lehmann, M.L., Cooper, H.A., Maric, D. & Herkenham, M. Social defeat induces depressive-like states and microglial activation without involvement of peripheral macrophages. J. Neuroinflammation 13, 224 (2016).
Acknowledgements
We thank S. Schwarzbaum and S. Urim for editing the paper; M. Schwartz, S. Shen-Orr, T. Ben-Shaanan and M. Schiller for their advice and helpful remarks; and A. Grau, Y. Sakuory and the Biomedical Core Facility at the Technion Faculty of Medicine for technical support and comments. This study was supported by the Israeli Ministry of Science, Technology & Space (MOST; 3-12070), the Prince Center for Neurodegenerative Diseases, Israeli Society for Science (ISF; 1862/15), an FP-7-CIG grant (618654) and the ADELIS Foundation. A.R. is an International Howard Hughes Medical Institute (HHMI)–Wellcome Trust researcher.
Author information
Authors and Affiliations
Contributions
B.K. designed the protocol, acquired the data and wrote the manuscript; T.D. contributed to the development of the protocol and to writing of the manuscript; and A.R. led the experimental design and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Gating strategy for flow cytometry analysis.
Cells from the (a) blood and (b) brain compartment are selected according to the forward scatter (FSC) and side scatter (SSC) area (A) parameters. Then, doublet exclusion is performed using height (H) versus width (W) parameters of FSC and SSC accordingly, and live/dead discrimination is applied using Zombie dye.
Supplementary information
Supplementary Figures and Text
Supplementary Figure 1 and Supplementary Table 1. (PDF 592 kb)
Rights and permissions
About this article
Cite this article
Korin, B., Dubovik, T. & Rolls, A. Mass cytometry analysis of immune cells in the brain. Nat Protoc 13, 377–391 (2018). https://doi.org/10.1038/nprot.2017.155
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2017.155
This article is cited by
-
Meningeal immunity and neurological diseases: new approaches, new insights
Journal of Neuroinflammation (2023)
-
Neuroinflammation, memory, and depression: new approaches to hippocampal neurogenesis
Journal of Neuroinflammation (2023)
-
Inductively coupled plasma mass spectrometry
Nature Reviews Methods Primers (2023)
-
Clonally expanded CD8 T cells characterize amyotrophic lateral sclerosis-4
Nature (2022)
-
Immune cell compartmentalization for brain surveillance and protection
Nature Immunology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.