Abstract
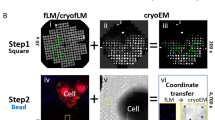
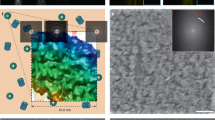
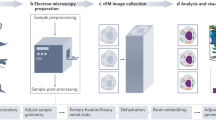
Correlative light and electron microscopy (CLEM) combines spatiotemporal information from fluorescence light microscopy (fLM) with high-resolution structural data from cryo-electron tomography (cryo-ET). These technologies provide opportunities to bridge knowledge gaps between cell and structural biology. Here we describe our protocol for correlated cryo-fLM, cryo-electron microscopy (cryo-EM), and cryo-ET (i.e., cryo-CLEM) of virus-infected or transfected mammalian cells. Mammalian-derived cells are cultured on EM substrates, using optimized conditions that ensure that the cells are spread thinly across the substrate and are not physically disrupted. The cells are then screened by fLM and vitrified before acquisition of cryo-fLM and cryo-ET images, which is followed by data processing. A complete session from grid preparation through data collection and processing takes 5–15 d for an individual experienced in cryo-EM.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout













Similar content being viewed by others
References
Morgan, C., Godman, G.C., Breitenfeld, P.M. & Rose, H.M. A correlative study by electron and light microscopy of the development of type 5 adenovirus. I. Electron microscopy. J. Exp. Med. 112, 373–382 (1960).
Godman, G.C., Morgan, C., Breitenfeld, P.M. & Rose, H.M. A correlative study by electron and light microscopy of the development of type 5 adenovirus. II. Light microscopy. J. Exp. Med. 112, 383–402 (1960).
Dubochet, J. et al. Cryo-electron microscopy of vitrified specimens. Q. Rev. Biophys. 21, 129–228 (1988).
Mancini, E.J., de Haas, F. & Fuller, S.D. High-resolution icosahedral reconstruction: fulfilling the promise of cryo-electron microscopy. Structure 5, 741–750 (1997).
Steven, A.C. & Aebi, U. The next ice age: cryo-electron tomography of intact cells. Trends Cell Biol. 13, 107–110 (2003).
Risco, C. & Carrascosa, J.L. Visualization of viral assembly in the infected cell. Histol. Histopathol. 14, 905–926 (1999).
Schroder, R.R. Advances in electron microscopy: a qualitative view of instrumentation development for macromolecular imaging and tomography. Arch. Biochem. Biophys. 581, 25–38 (2015).
McDonald, K.L. A review of high-pressure freezing preparation techniques for correlative light and electron microscopy of the same cells and tissues. J. Microsc. 235, 273–281 (2009).
Carroni, M. & Saibil, H.R. Cryo electron microscopy to determine the structure of macromolecular complexes. Methods 95, 78–85 (2016).
Giepmans, B.N., Adams, S.R., Ellisman, M.H. & Tsien, R.Y. The fluorescent toolbox for assessing protein location and function. Science 312, 217–224 (2006).
Desai, T.M. et al. Fluorescent protein-tagged Vpr dissociates from HIV-1 core after viral fusion and rapidly enters the cell nucleus. Retrovirology 12, 88 (2015).
Padilla-Parra, S. et al. Fusion of mature HIV-1 particles leads to complete release of a gag-GFP-based content marker and raises the intraviral pH. PLoS One 8, e71002 (2013).
Briegel, A. et al. Correlated light and electron cryo-microscopy. Methods Enzymol. 481, 317–341 (2010).
Strauss, J.D. et al. Three-dimensional structural characterization of HIV-1 tethered to human cells. J. Virol. 90, 1507–1521 (2015).
Yi, H. et al. Native immunogold labeling of cell surface proteins and viral glycoproteins for cryo-electron microscopy and cryo-electron tomography applications. J. Histochem. Cytochem. 63, 780–792 (2015).
Mastronarde, D.N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Bharat, T.A., Russo, C.J., Lowe, J., Passmore, L.A. & Scheres, S.H. Advances in single-particle electron cryomicroscopy structure determination applied to sub-tomogram averaging. Structure 23, 1743–1753 (2015).
Hrabe, T. Localize.pytom: a modern webserver for cryo-electron tomography. Nucleic Acids Res. 43, W231–236 (2015).
Huiskonen, J.T. et al. Averaging of viral envelope glycoprotein spikes from electron cryotomography reconstructions using Jsubtomo. J. Vis. Exp. e51714 (2014).
Heymann, J.B., Cardone, G., Winkler, D.C. & Steven, A.C. Computational resources for cryo-electron tomography in Bsoft. J. Struct. Biol. 161, 232–242 (2008).
Castano-Diez, D., Kudryashev, M., Arheit, M. & Stahlberg, H. Dynamo: a flexible, user-friendly development tool for subtomogram averaging of cryo-EM data in high-performance computing environments. J. Struct. Biol. 178, 139–151 (2012).
Koning, R.I. et al. Correlative cryo-fluorescence light microscopy and cryo-electron tomography of Streptomyces. Methods Cell Biol. 124, 217–239 (2014).
van Driel, L.F., Valentijn, J.A., Valentijn, K.M., Koning, R.I. & Koster, A.J. Tools for correlative cryo-fluorescence microscopy and cryo-electron tomography applied to whole mitochondria in human endothelial cells. Eur. J. Cell Biol. 88, 669–684 (2009).
Jun, S. et al. Direct visualization of HIV-1 with correlative live-cell microscopy and cryo-electron tomography. Structure 19, 1573–1581 (2011).
Ibiricu, I. et al. Cryo electron tomography of herpes simplex virus during axonal transport and secondary envelopment in primary neurons. PLoS Pathog. 7, e1002406 (2011).
Schorb, M. & Briggs, J.A. Correlated cryo-fluorescence and cryo-electron microscopy with high spatial precision and improved sensitivity. Ultramicroscopy 143, 24–32 (2014).
Briegel, A. et al. Location and architecture of the Caulobacter crescentus chemoreceptor array. Mol. Microbiol. 69, 30–41 (2008).
Swulius, M.T. et al. Long helical filaments are not seen encircling cells in electron cryotomograms of rod-shaped bacteria. Biochem. Biophys. Res. Commun. 407, 650–655 (2011).
Chang, Y.W. et al. Correlated cryogenic photoactivated localization microscopy and cryo-electron tomography. Nat. Methods 11, 737–739 (2014).
Wang, Q., Mercogliano, C.P. & Lowe, J. A ferritin-based label for cellular electron cryotomography. Structure 19, 147–154 (2011).
Gold, V.A. et al. Visualizing active membrane protein complexes by electron cryotomography. Nat. Commun. 5, 4129 (2014).
Mercogliano, C.P. & DeRosier, D.J. Concatenated metallothionein as a clonable gold label for electron microscopy. J. Struct. Biol. 160, 70–82 (2007).
Al-Amoudi, A. et al. Cryo-electron microscopy of vitreous sections. EMBO J. 23, 3583–3588 (2004).
Gruska, M., Medalia, O., Baumeister, W. & Leis, A. Electron tomography of vitreous sections from cultured mammalian cells. J. Struct. Biol. 161, 384–392 (2008).
Engel, B.D. et al. Native architecture of the Chlamydomonas chloroplast revealed by in situ cryo-electron tomography. Elife 4, e04889 (2015).
Marko, M., Hsieh, C., Schalek, R., Frank, J. & Mannella, C. Focused-ion-beam thinning of frozen-hydrated biological specimens for cryo-electron microscopy. Nat. Methods 4, 215–217 (2007).
Wolf, S.G., Houben, L. & Elbaum, M. Cryo-scanning transmission electron tomography of vitrified cells. Nat. Methods 11, 423–428 (2014).
Bykov, Y.S., Cortese, M., Briggs, J.A. & Bartenschlager, R. Correlative light and electron microscopy methods for the study of virus-cell interactions. FEBS Lett. 590, 1877–1895 (2016).
Schellenberger, P. et al. High-precision correlative fluorescence and electron cryo microscopy using two independent alignment markers. Ultramicroscopy 143, 41–51 (2014).
Liu, B. et al. Three-dimensional super-resolution protein localization correlated with vitrified cellular context. Sci. Rep. 5, 13017 (2015).
Wolff, G., Hagen, C., Grunewald, K. & Kaufmann, R. Towards correlative super-resolution fluorescence and electron cryo-microscopy. Biol. Cell 108, 245–258 (2016).
Schwartz, C.L., Sarbash, V.I., Ataullakhanov, F.I., McIntosh, J.R. & Nicastro, D. Cryo-fluorescence microscopy facilitates correlations between light and cryo-electron microscopy and reduces the rate of photobleaching. J. Microsc. 227, 98–109 (2007).
Sartori, A. et al. Correlative microscopy: bridging the gap between fluorescence light microscopy and cryo-electron tomography. J. Struct. Biol. 160, 135–145 (2007).
Rigort, A. et al. Micromachining tools and correlative approaches for cellular cryo-electron tomography. J. Struct. Biol. 172, 169–179 (2010).
Faas, F.G. et al. Localization of fluorescently labeled structures in frozen-hydrated samples using integrated light electron microscopy. J. Struct. Biol. 181, 283–290 (2013).
Schorb, M. et al. New hardware and workflows for semi-automated correlative cryo-fluorescence and cryo-electron microscopy/tomography. J. Struct. Biol. https://dx.doi.org/10.1016/j.jsb.2016.06.020 (2016).
Galaz-Montoya, J.G., Flanagan, J., Schmid, M.F. & Ludtke, S.J. Single particle tomography in EMAN2. J. Struct. Biol. 190, 279–290 (2015).
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007).
Nicastro, D. et al. The molecular architecture of axonemes revealed by cryoelectron tomography. Science 313, 944–948 (2006).
Heumann, J.M., Hoenger, A. & Mastronarde, D.N. Clustering and variance maps for cryo-electron tomography using wedge-masked differences. J. Struct. Biol. 175, 288–299 (2011).
Winkler, H. 3D reconstruction and processing of volumetric data in cryo-electron tomography. J. Struct. Biol. 157, 126–137 (2007).
Huiskonen, J.T. et al. Electron cryotomography of Tula hantavirus suggests a unique assembly paradigm for enveloped viruses. J. Virol. 84, 4889–4897 (2010).
Kremer, J.R., Mastronarde, D.N. & McIntosh, J.R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Hagen, W.J., Wan, W. & Briggs, J.A. Implementation of a cryo-electron tomography tilt-scheme optimized for high resolution subtomogram averaging. J. Struct. Biol. http://dx.doi/org/10.1016/j.jsb.2016.06.007 (2016).
Moerner, W.E. & Orrit, M. Illuminating single molecules in condensed matter. Science 283, 1670–1676 (1999).
Kaufmann, R., Hagen, C. & Grunewald, K. Fluorescence cryo-microscopy: current challenges and prospects. Curr. Opin. Chem. Biol. 20, 86–91 (2014).
Xiong, Q., Morphew, M.K., Schwartz, C.L., Hoenger, A.H. & Mastronarde, D.N. CTF determination and correction for low dose tomographic tilt series. J. Struct. Biol. 168, 378–387 (2009).
Heymann, J.B. & Belnap, D.M. Bsoft: image processing and molecular modeling for electron microscopy. J. Struct. Biol. 157, 3–18 (2007).
Heymann, J.B. Bsoft: image and molecular processing in electron microscopy. J. Struct. Biol. 133, 156–169 (2001).
Acknowledgements
We thank the Robert P. Apkarian Integrated Electron Microscopy Core of Emory University for microscopy services and support. This work was supported in part by grants from Emory University, Children's Healthcare of Atlanta, and the Georgia Research Alliance to E.R.W.; a grant from the Center for AIDS Research at Emory University (P30 AI050409); a grant from the James B. Pendleton Charitable Trust to E.R.W. and P.W.S.; public health service grants R01GM104540, R21AI101775, and R01GM104540-03S1 from the NIH to E.R.W.; NSF grant 0923395 to E.R.W.; public health service grant R01GM114561 from the NIH to E.R.W. and P.J.S.; public health service grant R01AI058828 from the NIH to P.W.S.; public health service grants R01GM054787 and R01AI053668 from the NIH to G.B.M.; public health service grant R01GM094198 from the NIH to P.J.S.; and public health service grant F32GM112517 from the NIH to J.D.S. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
C.M.H. and E.R.W. wrote the manuscript. C.M.H., J.D.S., Z.K., R.S.D., J.E.H., E.A., T.M.D., M.M., G.B.M., P.J.S., P.W.S., and E.R.W. designed and performed the experiments, and edited the manuscript. R.E.S. and F.L. processed and analyzed data. All authors read and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–3 (PDF 238 kb)
Rights and permissions
About this article
Cite this article
Hampton, C., Strauss, J., Ke, Z. et al. Correlated fluorescence microscopy and cryo-electron tomography of virus-infected or transfected mammalian cells. Nat Protoc 12, 150–167 (2017). https://doi.org/10.1038/nprot.2016.168
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.168
This article is cited by
-
HOPE-SIM, a cryo-structured illumination fluorescence microscopy system for accurately targeted cryo-electron tomography
Communications Biology (2023)
-
Correlative montage parallel array cryo-tomography for in situ structural cell biology
Nature Methods (2023)
-
ELI trifocal microscope: a precise system to prepare target cryo-lamellae for in situ cryo-ET study
Nature Methods (2023)
-
Sample preparation strategies for efficient correlation of 3D SIM and soft X-ray tomography data at cryogenic temperatures
Nature Protocols (2021)
-
Post-correlation on-lamella cryo-CLEM reveals the membrane architecture of lamellar bodies
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.