Abstract
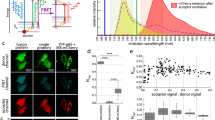
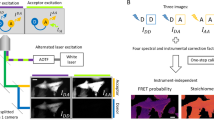
Förster resonance energy transfer (FRET) is a versatile method for analyzing protein–protein interactions within living cells. This protocol describes a nondestructive live-cell FRET assay for robust quantification of relative binding affinities for protein–protein interactions. Unlike other approaches, our method correlates the measured FRET efficiencies to relative concentration of interacting proteins to determine binding isotherms while including collisional FRET corrections. We detail how to assemble and calibrate the equipment using experimental and theoretical procedures. A step-by-step protocol is given for sample preparation, data acquisition and analysis. The method uses relatively inexpensive and widely available equipment and can be performed with minimal training. Implementation of the imaging setup requires up to 1 week, and sample preparation takes ∼1–3 d. An individual FRET experiment, including control measurements, can be completed within 4–6 h, with data analysis requiring an additional 1–3 h.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout











Similar content being viewed by others
Change history
01 December 2016
In the original version that appeared online, equal contribution of authors was incorrectly attributed; this has been corrected to indicate that Elisabeth Butz and Manu Ben Johny contributed equally to the work. The error has been corrected for the print, PDF and HTML versions of this article.
References
Clegg, R.M. Fluorescence resonance energy transfer and nucleic acids. Methods Enzymol. 211, 353–388 (1992).
Jares-Erijman, E.A. & Jovin, T.M. FRET imaging. Nat. Biotechnol. 21, 1387–1395 (2003).
Shaner, N.C., Steinbach, P.A. & Tsien, R.Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Berney, C. & Danuser, G. FRET or no FRET: a quantitative comparison. Biophys. J. 84, 3992–4010 (2003).
Gordon, G.W., Berry, G., Liang, X.H., Levine, B. & Herman, B. Quantitative fluorescence resonance energy transfer measurements using fluorescence microscopy. Biophys. J. 74, 2702–2713 (1998).
Xia, Z. & Liu, Y. Reliable and global measurement of fluorescence resonance energy transfer using fluorescence microscopes. Biophys. J. 81, 2395–2402 (2001).
Youvan, D.C. & Marrs, B.L. Molecular genetics and the light reactions of photosynthesis. Cell 39, 1–3 (1984).
Zal, T. & Gascoigne, N.R. Photobleaching-corrected FRET efficiency imaging of live cells. Biophys. J. 86, 3923–3939 (2004).
Erickson, M.G., Alseikhan, B.A., Peterson, B.Z. & Yue, D.T. Preassociation of calmodulin with voltage-gated Ca(2+) channels revealed by FRET in single living cells. Neuron 31, 973–985 (2001).
Erickson, M.G., Liang, H., Mori, M.X. & Yue, D.T. FRET two-hybrid mapping reveals function and location of L-type Ca2+ channel CaM preassociation. Neuron 39, 97–107 (2003).
Erickson, M.G., Moon, D.L. & Yue, D.T. DsRed as a potential FRET partner with CFP and GFP. Biophys. J. 85, 599–611 (2003).
Miyawaki, A. et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388, 882–887 (1997).
Chen, H., Puhl, H.L. III, Koushik, S.V., Vogel, S.S. & Ikeda, S.R. Measurement of FRET efficiency and ratio of donor to acceptor concentration in living cells. Biophys. J. 91, L39–L41 (2006).
Yang, P.S., Johny, M.B. & Yue, D.T. Allostery in Ca(2)(+) channel modulation by calcium-binding proteins. Nat. Chem. Biol. 10, 231–238 (2014).
Broussard, J.A., Rappaz, B., Webb, D.J. & Brown, C.M. Fluorescence resonance energy transfer microscopy as demonstrated by measuring the activation of the serine/threonine kinase Akt. Nat. Protoc. 8, 265–281 (2013).
Ormo, M. et al. Crystal structure of the Aequorea victoria green fluorescent protein. Science 273, 1392–1395 (1996).
Lohse, M.J., Nuber, S. & Hoffmann, C. Fluorescence/bioluminescence resonance energy transfer techniques to study G-protein-coupled receptor activation and signaling. Pharmacol. Rev. 64, 299–336 (2012).
Zhang, J., Ma, Y., Taylor, S.S. & Tsien, R.Y. Genetically encoded reporters of protein kinase A activity reveal impact of substrate tethering. Proc. Natl. Acad. Sci. USA 98, 14997–15002 (2001).
Takao, K. et al. Visualization of synaptic Ca2+/calmodulin-dependent protein kinase II activity in living neurons. J. Neurosci. 25, 3107–3112 (2005).
Bazzazi, H., Ben Johny, M., Adams, P.J., Soong, T.W. & Yue, D.T. Continuously tunable Ca(2+) regulation of RNA-edited CaV1.3 channels. Cell Rep. 5, 367–377 (2013).
Ben Johny, M., Yang, P.S., Bazzazi, H. & Yue, D.T. Dynamic switching of calmodulin interactions underlies Ca2+ regulation of CaV1.3 channels. Nat. Commun. 4, 1717 (2013).
Takahashi, S.X., Miriyala, J., Tay, L.H., Yue, D.T. & Colecraft, H.M. A CaVbeta SH3/guanylate kinase domain interaction regulates multiple properties of voltage-gated Ca2+ channels. J. Gen. Physiol. 126, 365–377 (2005).
Agler, H.L. et al. G protein-gated inhibitory module of N-type (ca(v)2.2) ca2+ channels. Neuron 46, 891–904 (2005).
Karpenko, I.A. et al. Selective nonpeptidic fluorescent ligands for oxytocin receptor: design, synthesis, and application to time-resolved FRET binding assay. J. Med. Chem. 58, 2547–2552 (2015).
Chang, D.D. & Colecraft, H.M. Rad and Rem are non-canonical G-proteins with respect to the regulatory role of guanine nucleotide binding in Ca 1.2 regulation. J. Physiol. 593, 5075–5090 (2015).
Yang, T., Xu, X., Kernan, T., Wu, V. & Colecraft, H.M. Rem, a member of the RGK GTPases, inhibits recombinant CaV1.2 channels using multiple mechanisms that require distinct conformations of the GTPase. J. Physiol. 588, 1665–1681 (2010).
Yamashita, M., Somasundaram, A. & Prakriya, M. Competitive modulation of Ca2+ release-activated Ca2+ channel gating by STIM1 and 2-aminoethyldiphenyl borate. J. Biol. Chem. 286, 9429–9442 (2011).
Muik, M. et al. Dynamic coupling of the putative coiled-coil domain of ORAI1 with STIM1 mediates ORAI1 channel activation. J. Biol. Chem. 283, 8014–8022 (2008).
Murphy, J.G. et al. AKAP-anchored PKA maintains neuronal L-type calcium channel activity and NFAT transcriptional signaling. Cell Rep. 7, 1577–1588 (2014).
Ruiz-Velasco, V. & Ikeda, S.R. Functional expression and FRET analysis of green fluorescent proteins fused to G-protein subunits in rat sympathetic neurons. J. Physiol. 537, 679–692 (2001).
Kern, A., Albarran-Zeckler, R., Walsh, H.E. & Smith, R.G. Apo-ghrelin receptor forms heteromers with DRD2 in hypothalamic neurons and is essential for anorexigenic effects of DRD2 agonism. Neuron 73, 317–332 (2012).
Albizu, L. et al. Time-resolved FRET between GPCR ligands reveals oligomers in native tissues. Nat. Chem. Biol. 6, 587–594 (2010).
Bazzazi, H. et al. Novel fluorescence resonance energy transfer-based reporter reveals differential calcineurin activation in neonatal and adult cardiomyocytes. J. Physiol. 593, 3865–3884 (2015).
Lee, S.R., Sang, L. & Yue, D.T. Uncovering aberrant mutant PKA function with flow cytometric FRET. Cell Rep. 14, 3019–3029 (2016).
Tadross, M.R., Park, S.A., Veeramani, B. & Yue, D.T. Robust approaches to quantitative ratiometric FRET imaging of CFP/YFP fluorophores under confocal microscopy. J. Microsc. 233, 192–204 (2009).
Shrestha, D., Jenei, A., Nagy, P., Vereb, G. & Szollosi, J. Understanding FRET as a research tool for cellular studies. Int. J. Mol. Sci. 16, 6718–6756 (2015).
Ben-Johny, M., Yue, D.N. & Yue, D.T. A novel FRET-based assay reveals 1:1 stoichiometry of apocalmodulin binding across voltage-gated Ca and Na ion channels. Biophys. J. 102, 125a–126a (2012).
Koushik, S.V., Chen, H., Thaler, C., Puhl, H.L. III & Vogel, S.S. Cerulean, venus, and venusY67C FRET reference standards. Biophys. J. 91, L99–L101 (2006).
Sun, Y., Day, R.N. & Periasamy, A. Investigating protein-protein interactions in living cells using fluorescence lifetime imaging microscopy. Nat. Protoc. 6, 1324–1340 (2011).
Yasuda, R. Imaging spatiotemporal dynamics of neuronal signaling using fluorescence resonance energy transfer and fluorescence lifetime imaging microscopy. Curr. Opin. Neurobiol. 16, 551–561 (2006).
Thaler, C., Koushik, S.V., Blank, P.S. & Vogel, S.S. Quantitative multiphoton spectral imaging and its use for measuring resonance energy transfer. Biophys. J. 89, 2736–2749 (2005).
Zeug, A., Woehler, A., Neher, E. & Ponimaskin, E.G. Quantitative intensity-based FRET approaches—a comparative snapshot. Biophys. J. 103, 1821–1827 (2012).
Leavesley, S.J., Britain, A.L., Cichon, L.K., Nikolaev, V.O. & Rich, T.C. Assessing FRET using spectral techniques. Cytometry A 83, 898–912 (2013).
Mustafa, S., Hannagan, J., Rigby, P., Pfleger, K. & Corry, B. Quantitative Forster resonance energy transfer efficiency measurements using simultaneous spectral unmixing of excitation and emission spectra. J. Biomed. Opt. 18, 26024 (2013).
Bykova, E.A., Zhang, X.D., Chen, T.Y. & Zheng, J. Large movement in the C terminus of CLC-0 chloride channel during slow gating. Nat. Struct. Mol. Biol. 13, 1115–1119 (2006).
Takanishi, C.L., Bykova, E.A., Cheng, W. & Zheng, J. GFP-based FRET analysis in live cells. Brain Res. 1091, 132–139 (2006).
Hoppe, A.D. & Swanson, J.A. Cdc42, Rac1, and Rac2 display distinct patterns of activation during phagocytosis. Mol. Biol. Cell 15, 3509–3519 (2004).
Chen, H., Puhl, H.L. III & Ikeda, S.R. Estimating protein-protein interaction affinity in living cells using quantitative Forster resonance energy transfer measurements. J. Biomed. Opt. 12, 054011 (2007).
Hoppe, A., Christensen, K. & Swanson, J.A. Fluorescence resonance energy transfer-based stoichiometry in living cells. Biophys. J. 83, 3652–3664 (2002).
Chen, Y., Mauldin, J.P., Day, R.N. & Periasamy, A. Characterization of spectral FRET imaging microscopy for monitoring nuclear protein interactions. J. Microsc. 228, 139–152 (2007).
Levy, S. et al. SpRET: highly sensitive and reliable spectral measurement of absolute FRET efficiency. Microsc. Microanal. 17, 176–190 (2011).
Kemmer, G. & Keller, S. Nonlinear least-squares data fitting in Excel spreadsheets. Nat. Protoc. 5, 267–281 (2010).
Findeisen, F. & Minor, D.L. Jr. Structural basis for the differential effects of CaBP1 and calmodulin on Ca(V)1.2 calcium-dependent inactivation. Structure 18, 1617–1631 (2010).
Martin, S.R. & Bayley, P.M. Calmodulin bridging of IQ motifs in myosin-V. FEBS Lett. 567, 166–170 (2004).
Acknowledgements
We thank R. Rötzer, A. Rößing, T. Kiwitt and W. Yang for excellent technical support. This work was supported by funding from the German Research Foundation (SFB/TRR 152 TP12 to M.B., SFB/TRR 152 TP06 to C.W.-S. and SFB 870 TP B05 to C.W.-S.) and the National Institute of Mental Health (R01 MH065531 to D.T.Y. and M.B.-J.). D.T.Y. passed away on December 23, 2014. He was a remarkable teacher, inspirational scientist and a generous colleague. He pioneered the development and the usage of FRET two-hybrid assays to elucidate the mechanistic underpinnings of voltage-gated Ca2+ channels. His scientific brilliance and personal charm are greatly missed in the scientific community.
Author information
Authors and Affiliations
Contributions
E.S.B., M.B.-J., D.T.Y. and C.W.-S. developed and/or designed the study. E.S.B. and M.B.-J. collected and analyzed the data. M.S. and L.S. furnished initial experimental data for dimer calibrations. M.B.-J. and D.T.Y. provided theoretical derivations. M.S., M.B.-J. and P.S.Y. constructed YFP- and CFP-tagged constructs and dimers. E.S.B., M.B.-J. and C.W.-S. wrote the manuscript and created the figures. M.B. discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Linearity of fluorescence detection subsystem
(a) Fluorescence intensity measured through the YFP cube is linearly proportional to the concentration of Alexa Fluor 514 dye. This result illustrates the linearity of the PMT detector enabling robust estimation of number of acceptor molecules. (b) Fluorescence intensity measured through the CFP cube is also linearly proportional to the concentration of Proflavin dye.
Supplementary Figure 2 Main graphical user interface of Felix GX
(a) The tab “Delta Ram Analog” (right upper corner) is clicked to open the “Hardware Configuration” window (Supplementary Fig. 2a). (b) The setup tab is selected (left lower corner) to open the “Setup” window (Supplementary Fig. 2b). (c) The “Action” tab opens the “Macro” window from which the “Macro Editor” window is opened (Supplementary Fig. 4).
Supplementary Figure 3 The “Hardware Configuration” window and the “Setup” window.
(a) Settings for the “Hardware Configuration” window. (b) Settings for the “Setup” window. From the 10 tabs in the “Setup” window select “Acquisition Type” and click the option “Timebased”.
Supplementary Figure 4 Parameter settings specific for Timebased acquisition in the “Setup window”.
(a) Acquisition settings; (b) PMTs; (c) Backgrounds; (d) Traces tab.
Supplementary Figure 5 Setup of Macros using the “Macro” and the “Macro Editor” window.
(a) Open the “Macro” window by clicking the “Action“ tab in the main graphical user interface (Supplemental Fig. 1). Click the green “+” button to open the “Macro Editor” window (b and c). In this window, different actions (listed on the left hand side) can be added to the macro (list on the black panel on the right hand side). In this example, a pause step (b) and a protocol (Run Acquisition; c) is added to the macro by clicking the Add tab.
Supplementary Figure 6 Data input field of the Background sheet.
(a) Raw data pasted into the respective columns from measurements using CFP- FRET- and YFP-cubes at a distinct HV gain. (b) The excel sheet contains 9 individual sub sheets (labeled from left to right: Background, RD-value (CFP), RA-value (YFP), Dimer sheet, Dimer Analysis, Spurious FRET, Samples (FRET sample data), BC Analysis and HV lookup.
Supplementary Figure 7 Data input field of the RD-value (CFP) and the RA-value (YFP) sheets.
Raw data are pasted into the respective columns on the left hand side from measurements in (a) CFP-only or (b) YFP-only expressing cells using CFP- FRET- and YFP-cubes at a distinct HV gain. Background subtraction and RA value calculation is performed automatically. (c) The mean RD and RA value, resp., is calculated on the right hand side (gray panel). Statistical values (Standard deviation of the mean (SEM), Minimum (Min) and Maximum (Max)) provide a useful tool to assess the variability of the values. Note that only the mean RD1 and RA1 values are used for subsequent analysis.
Supplementary Figure 8 Data input field of the Dimer sheet.
(a) Raw data are pasted into the respective columns on the left hand side from measurements in CFP-YFP dimer expressing cells using CFP- FRET- and YFP-cubes at a distinct HV gain. Raw data for individual cells are collected separately for a given dimer as indicated. (b) Background subtracted values are shown on the right. RA1 and RD1 values are automatically picked from RD-value (CFP) and the RA-value (YFP) sheets. Statistics are shown on the right hand side (Mean, SEM).
Supplementary Figure 9 Dimer analysis sheet.
(a) Columns D-G list background subtracted raw data from the “Dimer” sheet. Data for individual cells are collected separately for a given dimer as indicated in Column B. RA1 and RD1 values are automatically transferred from “RD-value (CFP)” and the “RA-value (YFP)” sheets (grey panel on the top, left). Columns H-P comprise calculations as outlined in Procedure Step 97. (b) The graph (column K plotted against column L) depicts data points for individual cells. For each dimer, cells cluster together. In the table below, the mean values of EA and ED for each dimer as well as mean values of the x- and y- coordinates (from columns M and N) are indicated. (c) The latter can be plotted in a new graph and analyzed by a linear regression line. In this template, the regression line is automatically computed and the resulting equation displayed in the graph. Note, that the y-intercept and the slope of this line is copied to cell T56 and T57 respectively for further analysis.
Supplementary Figure 10 Data input field of the “Spurious FRET” sheet.
(a) Raw data from cells expressing untagged eCFP and eYFP are transferred to respective cells in Columns C-G. (b) Background-subtraction as well as subsequent calculation of Fc/(RA1*SYFP), CFPEST and EA (columns K-M) is automatically performed referring to the constants in the gray panel at the upper left. (c) The graph at the right hand side, illustrates CFPEST as a function of EA. The slope of the regression line is used for spurious FRET correction in the “BC analysis sheet”. The equation of the regression line is automatically displayed in the graph. The slope is subsequently entered into cell “P7” linked to the “BC analysis” sheet.
Supplementary Figure 11 Data input field of the binding curve analysis (“BC analysis”) sheet.
(a) Background subtracted raw data (columns D-H) are automatically adopted from the “Samples” sheet. The principal calculation steps required in order to solve binding curves are automatically performed in columns I-W for each (biological) cell according Steps 117-130 outlined in the Procedure section. (b) The table in the upper left of the worksheet contains all parameters (columns F and I) necessary to correct for spectral properties and spurious FRET. Thereby, all values except the excitation ratio ((ƐYFP(440)/ƐCFP(440)) are automatically transferred from previously analyzed sheets. Values for KdEff, EAmax and EDmax (Column K) are place holders that are adjusted during the fitting procedure. (c) Graphs shown displays data points for individual biological cells (blue dots) and the fitted binding isotherme (black lines). The graphs represent data for 33-FRET (Dfree plotted against EA) and E-FRET (Afree plotted against ED) and allow a control of the fitting procedure.
Supplementary Figure 12 Setting up Excel Solver for optimizing 33-FRET and E-FRET binding parameters.
(a) Least squares optimization of Cell K8 (Sum_Err_3Cube) by changing both KD_eff (KD,EFF, relative affinity) and EAmax (EA,max, maximum 33-FRET efficiency). In this scenario, two constraints are EAmax>0, and Kd_eff>0 in accordance with physical principles. (b) Least squares optimization of Cell K10 (Total Error) by changing both KD_eff (KD,EFF, relative affinity), EAmax (EA,max, maximum 33-FRET efficiency), and EDmax (ED,max, maximum E-FRET efficiency). Three constraints are imposed: EDmax>0, EAmax>0, and Kd_eff>0.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Notes 1–4 (PDF 4424 kb)
Supplementary Data 1
MS Excel analysis spreadsheet for theoretical calculation of MD and MA containing sample data. (XLSX 270 kb)
Supplementary Data 2
MATLAB import/export script source file (M). (XLSX 46 kb)
Supplementary Data 3
MS Excel analysis spreadsheet containing sample data. (XLSX 133 kb)
Rights and permissions
About this article
Cite this article
Butz, E., Ben-Johny, M., Shen, M. et al. Quantifying macromolecular interactions in living cells using FRET two-hybrid assays. Nat Protoc 11, 2470–2498 (2016). https://doi.org/10.1038/nprot.2016.128
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.128
This article is cited by
-
Interaction of human CRX and NRL in live HEK293T cells measured using fluorescence resonance energy transfer (FRET)
Scientific Reports (2022)
-
Cytosolic peptides encoding CaV1 C-termini downregulate the calcium channel activity-neuritogenesis coupling
Communications Biology (2022)
-
Critical contributions of pre-S1 shoulder and distal TRP box in DAG-activated TRPC6 channel by PIP2 regulation
Scientific Reports (2022)
-
An epilepsy-causing mutation leads to co-translational misfolding of the Kv7.2 channel
BMC Biology (2021)
-
Bak instead of Bax plays a key role in metformin-induced apoptosis s in HCT116 cells
Cell Death Discovery (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.