Abstract
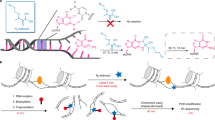
Many features of how gene transcription occurs in human cells remain unclear, mainly because of a lack of quantitative approaches to follow genome transcription with nucleotide precision in vivo. Here we present a robust genome-wide approach for studying RNA polymerase II (Pol II)–mediated transcription in human cells at single-nucleotide resolution by native elongating transcript sequencing (NET-seq). Elongating RNA polymerase and the associated nascent RNA are prepared by cell fractionation, avoiding immunoprecipitation or RNA labeling. The 3′ ends of nascent RNAs are captured through barcode linker ligation and converted into a DNA sequencing library. The identity and abundance of the 3′ ends are determined by high-throughput sequencing, which reveals the exact genomic locations of Pol II. Human NET-seq can be applied to the study of the full spectrum of Pol II transcriptional activities, including the production of unstable RNAs and transcriptional pausing. By using the protocol described here, a NET-seq library can be obtained from human cells in 5 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Margaritis, T. & Holstege, F.C.P. Poised RNA polymerase II gives pause for thought. Cell 133, 581–584 (2008).
Jonkers, I. & Lis, J.T. Getting up to speed with transcription elongation by RNA polymerase II. Nat. Rev. Mol. Cell Biol. 16, 167–177 (2015).
Adelman, K. & Lis, J.T. Promoter-proximal pausing of RNA polymerase II: emerging roles in metazoans. Nat. Rev. Genet. 13, 720–731 (2012).
Ptashne, M. & Gann, A. Transcriptional activation by recruitment. Nature 386, 599–577 (1997).
Rahl, P.B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Jonkers, I., Kwak, H. & Lis, J.T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. Elife 3, e02407 (2014).
Preker, P. et al. RNA exosome depletion reveals transcription upstream of active human promoters. Science 322, 1851–1854 (2008).
Core, L.J., Waterfall, J.J. & Lis, J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Seila, A.C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008).
Sigova, A.A. et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proc. Natl. Acad. Sci. USA 110, 2876–2881 (2013).
Seila, A.C., Core, L.J., Lis, J.T. & Sharp, P.A. Divergent transcription: a new feature of active promoters. Cell Cycle 8, 2557–2564 (2009).
Djebali, S. et al. Landscape of transcription in human cells. Nature 488, 101–108 (2013).
Pelechano, V. & Steinmetz, L.M. Gene regulation by antisense transcription. Nat. Rev. Genet. 14, 880–893 (2013).
Jensen, T.H., Jacquier, A. & Libri, D. Dealing with pervasive transcription. Mol. Cell 52, 473–484 (2013).
Buck, M.J. & Lieb, J.D. ChIP-chip: considerations for the design, analysis, and application of genome-wide chromatin immunoprecipitation experiments. Genomics 83, 349–360 (2004).
Guenther, M.G., Levine, S.S., Boyer, L.A., Jaenisch, R. & Young, R.A. A chromatin landmark and transcription initiation at most promoters in human cells. Cell 130, 77–88 (2007).
Raha, D., Hong, M. & Snyder, M. ChIP-Seq: a method for global identification of regulatory elements in the genome. Curr. Protoc. Mol. Biol. 91, 21.19.1–21.19.14 (2010).
Descostes, N. et al. Tyrosine phosphorylation of RNA polymerase II CTD is associated with antisense promoter transcription and active enhancers in mammalian cells. Elife 3, e02105 (2014).
Rhee, H.S. & Pugh, B.F. ChIP-exo method for identifying genomic location of DNA-binding proteins with near-single-nucleotide accuracy. Curr. Protoc. Mol. Biol. 100, 21.24.1–21.24.14 (2012).
He, Q., Johnston, J. & Zeitlinger, J. ChIP-nexus enables improved detection of in vivo transcription factor binding footprints. Nat. Biotechnol. 33, 395–401 (2015).
Kwak, H., Fuda, N.J., Core, L.J. & Lis, J.T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Wang, I.X. et al. RNA-DNA differences are generated in human cells within seconds after RNA exits polymerase II. Cell Rep. 6, 906–915 (2014).
Churchman, L.S. & Weissman, J.S. Nascent transcript sequencing visualizes transcription at nucleotide resolution. Nature 469, 368–373 (2012).
Churchman, L.S. & Weissman, J.S. Native elongating transcript sequencing (NET-seq). Curr. Protoc. Mol. Biol. 98, 14.4.1–14.4.17 (2012).
Larson, M.H. et al. A pause sequence enriched at translation start sites drives transcription dynamics in vivo. Science 344, 1042–1047 (2014).
Vvedenskaya, I.O. et al. Interactions between RNA polymerase and the 'core recognition element' counteract pausing. Science 344, 1285–1289 (2014).
Weber, C.M., Ramachandran, S. & Henikoff, S. Nucleosomes are context-specific, H2A.Z-modulated barriers to RNA polymerase. Mol. Cell 53, 819–830 (2014).
Ferrari, F. et al. 'Jump start and gain' model for dosage compensation in drosophila based on direct sequencing of nascent transcripts. Cell Rep. 5, 629–636 (2013).
Mayer, A. et al. Native elongating transcript sequencing reveals human transcriptional activity at nucleotide resolution. Cell 161, 541–554 (2015).
Martinez-Rucobo, F.W., Sainsbury, S., Cheung, A.C. & Cramer, P. Architecture of the RNA polymerase–Spt4/5 complex and basis of universal transcription processivity. EMBO J. 30, 1302–1310 (2011).
Kireeva, M.L., Komissarova, N., Waugh, D.S. & Kashlev, M. The 8-nucleotide-long RNA:DNA hybrid is a primary stability determinant of the RNA polymerase II elongation complex. J. Biol. Chem. 275, 6530–6536 (2000).
Wuarin, J. & Schibler, U. Physical isolation of nascent RNA chains transcribed by RNA polymerase II: evidence for cotranscriptional splicing. Mol. Cell. Biol. 14, 7219–7225 (1994).
Cai, H. & Luse, D.S. Transcription initiation by RNA polymerase II in vitro. J. Biol. Chem. 262, 298–304 (1987).
Bhatt, D.M. et al. Transcript dynamics of proinflammatory genes revealed by sequence analysis of subcellular RNA fractions. Cell 150, 279–290 (2012).
Khodor, Y.L. et al. Nascent-seq indicates widespread cotranscriptional pre-mRNA splicing in Drosophila. Genes Dev. 25, 2502–2512 (2011).
Pandya-Jones, A. & Black, D.L. Co-transcriptional splicing of constitutive and alternative exons. RNA 15, 1896–1908 (2009).
Lindell, T.J., Weinberg, F., Morris, P.W., Roeder, R.G. & Rutter, W.J. Specific inhibition of nuclear RNA polymerase II by alpha-amanitin. Science 170, 447–449 (1970).
Ingolia, N.T., Brar, G.A., Rouskin, S., McGeachy, A.M. & Weissman, J.S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Flynn, R.A., Almada, A.E., Zamudio, J.R. & Sharp, P.A. Antisense RNA polymerase II divergent transcripts are P-TEFb dependent and substrates for the RNA exosome. Proc. Natl. Acad. Sci. USA 108, 10460–10465 (2011).
Kim, T.K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010).
De Santa, F. et al. A large fraction of extragenic RNA pol II transcription sites overlap enhancers. PLoS Biol. 8, e1000384 (2010).
Almada, A.E., Wu, X., Kriz, A.J., Burge, C.B. & Sharp, P.A. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature 499, 360–363 (2013).
Brannan, K. et al. mRNA decapping factors and the exonuclease Xrn2 function in widespread premature termination of RNA polymerase II transcription. Mol. Cell 46, 311–324 (2012).
Ntini, E. et al. Polyadenylation site–induced decay of upstream transcripts enforces promoter directionality. Nat. Struct. Mol. Biol. 20, 923–928 (2013).
Ozsolak, F. et al. Comprehensive polyadenylation site maps in yeast and human reveal pervasive alternative polyadenylation. Cell 143, 1018–1029 (2010).
Kim, J.B. & Sharp, P.A. Positive transcription elongation factor B phosphorylates hSPT5 and RNA polymerase II carboxyl-terminal domain independently of cyclin-dependent kinase-activating kinase. J. Biol. Chem. 276, 12317–12323 (2001).
Peterlin, B.M. & Price, D.H. Controlling the elongation phase of transcription with P-TEFb. Mol. Cell 23, 297–305 (2006).
Scruggs, B.S. et al. Bidirectional transcription arises from two distinct hubs of transcription factor binding and active chromatin. Mol. Cell 58, 1101–1112 (2015).
Danko, C.G. et al. Signaling pathways differentially affect RNA polymerase II initiation, pausing, and elongation rate in cells. Mol. Cell 50, 212–222 (2013).
Laitem, C. et al. CDK9 inhibitors define elongation checkpoints at both ends of RNA polymerase II–transcribed genes. Nat. Struct. Mol. Biol. 22, 396–403 (2015).
Nojima, T. et al. Mammalian NET-Seq reveals genome-wide nascent transcription coupled to RNA processing. Cell 161, 526–540 (2015).
Mayer, A. et al. Uniform transitions of the general RNA polymerase II transcription complex. Nat. Struct. Mol. Biol. 17, 1272–1278 (2010).
Kim, H. et al. Gene-specific RNA polymerase II phosphorylation and the CTD code. Nat. Struct. Mol. Biol. 17, 1279–1286 (2010).
Zeitlinger, J. et al. RNA polymerase stalling at developmental control genes in the Drosophila melanogaster embryo. Nat. Genet. 39, 1512–1516 (2007).
Muse, G.W. et al. RNA polymerase is poised for activation across the genome. Nat. Genet. 39, 1507–1511 (2007).
Steinmetz, E.J. et al. Genome-wide distribution of yeast RNA polymerase II and its control by Sen1 helicase. Mol. Cell 24, 735–746 (2006).
Bataille, A.R. et al. A universal RNA polymerase II CTD cycle is orchestrated by complex interplays between kinase, phosphatase, and isomerase enzymes along genes. Mol. Cell 45, 158–170 (2012).
Tietjen, J.R. et al. Chemical-genomic dissection of the CTD code. Nat. Struct. Mol. Biol. 17, 1154–1161 (2010).
Mahony, S. & Pugh, B.F. Protein–DNA binding in high-resolution. Crit. Rev. Biochem. Mol. Biol. 50, 269–283 (2015).
Gilmour, D.S. & Fan, R. Detecting transcriptionally engaged RNA polymerase in eukaryotic cells with permanganate genomic footprinting. Methods 48, 368–374 (2009).
Core, L.J. et al. Defining the status of RNA polymerase at promoters. Cell Rep. 2, 1025–1035 (2012).
Brueckner, F. & Cramer, P. Structural basis of transcription inhibition by α-amanitin and implications for RNA polymerase II translocation. Nat. Struct. Mol. Biol. 15, 811–818 (2008).
Chapman, R.D. et al. Transcribing RNA polymerase II is phosphorylated at CTD residue serine-7. Science 318, 1780–1782 (2007).
Schneider, C.A., Rasband, W.S. & Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2012).
Anders, S., Pyl, P.T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, J.T. et al. Integrative Genomics Viewer. Nat. Biotechnol. 29, 24–26 (2011).
Acknowledgements
We thank J. di Iulio and S. Maleri for help developing this approach. We thank J. di Iulio, H. Landry and A. Snavely for critical comments on the manuscript. This work was supported by US National Institutes of Health National Human Genome Research Institute (NHGRI) grant R01HG007173 to L.S.C.; a Damon Runyon Dale F. Frey Award for Breakthrough Scientists (to L.S.C.); and a Burroughs Wellcome Fund Career Award at the Scientific Interface (to L.S.C.). A.M. was supported by Long-Term Postdoctoral Fellowships of the Human Frontier Science Program (HFSP) (LT000314/2013-L) and the European Molecular Biology Organization (EMBO) (ALTF858-2012).
Author information
Authors and Affiliations
Contributions
A.M. performed experiments and prepared figures. A.M. and L.S.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Representative gel to monitor the efficiency of DNA linker ligations.
The RNA control oligo oGAB11 (lane 2) was ligated to the DNA linker (no barcode, lane 3) and to the Barcode DNA linker (lane 4). The ligation was performed as described in Procedure steps 25 to 28. The samples were separated on a 15% (wt/vol) polyacrylamide TBE-Urea gel as described in Procedure steps 38 to 43 of this protocol. The ligation efficiency is >95% for both ligations (lanes 3 and 4) as determined by image quantification using ImageJ 1.47v software.
Supplementary Figure 2 Full-size blots of the western analysis shown in Fig. 2.
For more details please see legend of Fig. 2.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 (PDF 217 kb)
Rights and permissions
About this article
Cite this article
Mayer, A., Churchman, L. Genome-wide profiling of RNA polymerase transcription at nucleotide resolution in human cells with native elongating transcript sequencing. Nat Protoc 11, 813–833 (2016). https://doi.org/10.1038/nprot.2016.047
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.047
This article is cited by
-
Expression of the miR-302/367 microRNA cluster is regulated by a conserved long non-coding host-gene
Scientific Reports (2021)
-
A dual role for H2A.Z.1 in modulating the dynamics of RNA polymerase II initiation and elongation
Nature Structural & Molecular Biology (2021)
-
Variability in mRNA translation: a random matrix theory approach
Scientific Reports (2021)
-
Pho dynamically interacts with Spt5 to facilitate transcriptional switches at the hsp70 locus
Epigenetics & Chromatin (2017)
-
Total RNA-seq to identify pharmacological effects on specific stages of mRNA synthesis
Nature Chemical Biology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.