Abstract
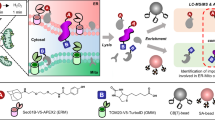
This protocol describes a method to obtain spatially resolved proteomic maps of specific compartments within living mammalian cells. An engineered peroxidase, APEX2, is genetically targeted to a cellular region of interest. Upon the addition of hydrogen peroxide for 1 min to cells preloaded with a biotin-phenol substrate, APEX2 generates biotin-phenoxyl radicals that covalently tag proximal endogenous proteins. Cells are then lysed, and biotinylated proteins are enriched with streptavidin beads and identified by mass spectrometry. We describe the generation of an appropriate APEX2 fusion construct, proteomic sample preparation, and mass spectrometric data acquisition and analysis. A two-state stable isotope labeling by amino acids in cell culture (SILAC) protocol is used for proteomic mapping of membrane-enclosed cellular compartments from which APEX2-generated biotin-phenoxyl radicals cannot escape. For mapping of open cellular regions, we instead use a 'ratiometric' three-state SILAC protocol for high spatial specificity. Isotopic labeling of proteins takes 5–7 cell doublings. Generation of the biotinylated proteomic sample takes 1 d, acquiring the mass spectrometric data takes 2–5 d and analysis of the data to obtain the final proteomic list takes 1 week.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Andersen, J.S. & Mann, M. Organellar proteomics: turning inventories into insights. EMBO Rep. 7, 874–879 (2006).
ten Have, S., Boulon, S., Ahmad, Y. & Lamond, A.I. Mass spectrometry-based immuno-precipitation proteomics—the user's guide. Proteomics 11, 1153–1159 (2011).
Brunner, Y., Schvartz, D., Couté, Y. & Sanchez, J.-C. Proteomics of regulated secretory organelles. Mass Spectrom. Rev. 28, 844–867 (2009).
Rhee, H.-W. et al. Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339, 1328–1331 (2013).
Hung, V. et al. Proteomic mapping of the human mitochondrial intermembrane space in live cells via ratiometric APEX tagging. Mol. Cell 55, 332–341 (2014).
Ong, S.-E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell. Proteomics 3, 1154–1169 (2004).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Lam, S.S. et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 12, 51–54 (2015).
Martell, J.D. et al. Engineered ascorbate peroxidase as a genetically encoded reporter for electron microscopy. Nat. Biotechnol. 30, 1143–1148 (2012).
Wishart, J.F. & Rao, B.S.M. Recent Trends in Radiation Chemistry (World Scientific, 2010).
Mortensen, A. & Skibsted, L.H. Importance of carotenoid structure in radical-scavenging reactions. J. Agric. Food Chem. 45, 2970–2977 (1997).
Bhaskar, B. et al. A novel heme and peroxide-dependent tryptophan–tyrosine cross-link in a mutant of cytochrome c peroxidase. J. Mol. Biol. 328, 157–166 (2003).
Rogers, M.S. et al. Cross-link formation of the cysteine 228–tyrosine 272 catalytic cofactor of galactose oxidase does not require dioxygen. Biochemistry 47, 10428–10439 (2008).
Amini, F., Kodadek, T. & Brown, K.C. Protein affinity labeling mediated by genetically encoded peptide tags. Angew. Chem. Int. Ed. 41, 356–359 (2002).
Colombini, M. A candidate for the permeability pathway of the outer mitochondrial membrane. Nature 279, 643–645 (1979).
Chen, C.L. et al. Proteomic mapping in live Drosophila tissues using an engineered ascorbate peroxidase. Proc. Natl. Acad. Sci. USA 112, 12093–12098 (2015).
Jing, J. et al. Proteomic mapping of ER-PM junctions identifies STIMATE as a regulator of Ca2+ influx. Nat. Cell Biol. 17, 1339–1347 (2015).
Mick, D.U. et al. Proteomics of primary cilia by proximity labeling. Dev. Cell 35, 497–512 (2015).
Polianskyte, Z. et al. LACTB is a filament-forming protein localized in mitochondria. Proc. Natl. Acad. Sci. USA 106, 18960–18965 (2009).
Chapman-Smith, A. & Cronan, J.E. Molecular biology of biotin attachment to proteins. J. Nutr. 129, 477S–484S (1999).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Choi-Rhee, E., Schulman, H. & Cronan, J.E. Promiscuous protein biotinylation by Escherichia coli biotin protein ligase. Protein Sci. 13, 3043–3050 (2004).
Roux, K.J., Kim, D.I., Raida, M. & Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 196, 801–810 (2012).
Morriswood, B. et al. Novel bilobe components in Trypanosoma brucei identified using proximity-dependent biotinylation. Eukaryot. Cell 12, 356–367 (2013).
Kim, D.I. et al. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc. Natl. Acad. Sci. USA 111, E2453–E2461 (2014).
Firat-Karalar, E.N., Rauniyar, N., Yates, J.R. III & Stearns, T. Proximity interactions among centrosome components identify regulators of centriole duplication. Curr. Biol. 24, 664–670 (2014).
DeMoss, J.A., Genuth, S.M. & Novelli, G.D. The enzymatic activation of amino acids via their acyl-adenylate derivatives. Proc. Natl. Acad. Sci. USA 42, 325–332 (1956).
Ryan, B.J., Carolan, N. & Ó'Fágáin, C. Horseradish and soybean peroxidases: comparable tools for alternative niches? Trends Biotechnol. 24, 355–363 (2006).
Ellisman, M.H., Deerinck, T.J., Shu, X. & Sosinsky, G.E. Picking faces out of a crowd: genetic labels for identification of proteins in correlated light and electron microscopy imaging. Methods Cell Biol. 111, 139–155 (2012).
Hopkins, C., Gibson, A., Stinchcombe, J. & Futter, C. Methods Enzymol. 327, 35–45 (2000).
Jiang, S. et al. A proteomics approach to the cell-surface interactome using the enzyme-mediated activation of radical sources reaction. Proteomics 12, 54–62 (2012).
Hubner, N.C. et al. Quantitative proteomics combined with BAC TransgeneOmics reveals in vivo protein interactions. J. Cell Biol. 189, 739–754 (2010).
Sancak, Y. et al. EMRE is an essential component of the mitochondrial calcium uniporter complex. Science 342, 1379–1382 (2013).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Pagliarini, D.J. et al. A mitochondrial protein compendium elucidates complex I disease biology. Cell 134, 112–123 (2008).
Acknowledgements
We thank H.-W. Rhee, P. Zou and J. Martell for help with EM samples. E. Vasile (Koch Institute Microscopy Core Facility) acquired the EM images. We thank O. Aygun and S. Han for their edits to the manuscript. V.H., S.S.L. and K.J.C. were supported by National Science Foundation Graduate Research Fellowships. S.S.L. was also supported by a National Defense Science and Engineering Graduate Fellowship. Funding was provided by the US National Institutes of Health (NIH R01 CA186568 to A.Y.T.) and the Howard Hughes Medical Institute Collaborative Initiative Award (to A.Y.T. and S.A.C.).
Author information
Authors and Affiliations
Contributions
V.H. performed the fluorescence characterization of matrix-APEX labeling and the proteomic data analysis. V.H., N.D.U. and A.Y.T. wrote the paper. All authors edited the paper.
Corresponding author
Ethics declarations
Competing interests
The Massachusetts Institute of Technology has submitted a patent application on this work.
Rights and permissions
About this article
Cite this article
Hung, V., Udeshi, N., Lam, S. et al. Spatially resolved proteomic mapping in living cells with the engineered peroxidase APEX2. Nat Protoc 11, 456–475 (2016). https://doi.org/10.1038/nprot.2016.018
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2016.018
This article is cited by
-
Profiling phagosome proteins identifies PD-L1 as a fungal-binding receptor
Nature (2024)
-
Streamlined Biotinylation, Enrichment and Analysis for Enhanced Plasma Membrane Protein Identification Using TurboID and TurboID-Start Biotin Ligases
The Journal of Membrane Biology (2024)
-
Proximity proteome mapping reveals PD-L1-dependent pathways disrupted by anti-PD-L1 antibody specifically in EGFR-mutant lung cancer cells
Cell Communication and Signaling (2023)
-
The development of proximity labeling technology and its applications in mammals, plants, and microorganisms
Cell Communication and Signaling (2023)
-
Adaptive preservation of orphan ribosomal proteins in chaperone-dispersed condensates
Nature Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.