Abstract
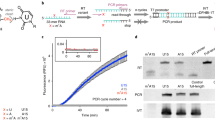
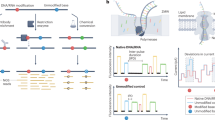
Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE) chemistries exploit small electrophilic reagents that react with 2′-hydroxyl groups to interrogate RNA structure at single-nucleotide resolution. Mutational profiling (MaP) identifies modified residues by using reverse transcriptase to misread a SHAPE-modified nucleotide and then counting the resulting mutations by massively parallel sequencing. The SHAPE-MaP approach measures the structure of large and transcriptome-wide systems as accurately as can be done for simple model RNAs. This protocol describes the experimental steps, implemented over 3 d, that are required to perform SHAPE probing and to construct multiplexed SHAPE-MaP libraries suitable for deep sequencing. Automated processing of MaP sequencing data is accomplished using two software packages. ShapeMapper converts raw sequencing files into mutational profiles, creates SHAPE reactivity plots and provides useful troubleshooting information. SuperFold uses these data to model RNA secondary structures, identify regions with well-defined structures and visualize probable and alternative helices, often in under 1 d. SHAPE-MaP can be used to make nucleotide-resolution biophysical measurements of individual RNA motifs, rare components of complex RNA ensembles and entire transcriptomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout













Similar content being viewed by others
References
Sharp, P.A. The centrality of RNA. Cell 136, 577–580 (2009).
Dethoff, E.A., Chugh, J., Mustoe, A.M. & Al-Hashimi, H.M. Functional complexity and regulation through RNA dynamics. Nature 482, 322–330 (2012).
Mauger, D.M., Siegfried, N.A. & Weeks, K.M. The genetic code as expressed through relationships between mRNA structure and protein function. FEBS Lett. 587, 1180–1188 (2013).
Weeks, K.M. Advances in RNA structure analysis by chemical probing. Curr. Opin. Struct. Biol. 20, 295–304 (2010).
Mortimer, S.A., Kidwell, M.A. & Doudna, J.A. Insights into RNA structure and function from genome-wide studies. Nat Rev Genet. 15, 469–479 (2014).
Kwok, C.K., Tang, Y., Assmann, S.M. & Bevilacqua, P.C. The RNA structurome: transcriptome-wide structure probing with next-generation sequencing. Trends Biochem. Sci. 40, 221–232 (2015).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Gherghe, C.M., Shajani, Z., Wilkinson, K.A., Varani, G. & Weeks, K.M. Strong correlation between SHAPE chemistry and the generalized NMR order parameter (S2) in RNA. J. Am. Chem. Soc. 130, 12244–12245 (2008).
Wilkinson, K.A. et al. Influence of nucleotide identity on ribose 2′-hydroxyl reactivity in RNA. RNA 15, 1314–1321 (2009).
McGinnis, J.L., Dunkle, J.A., Cate, J.H.D. & Weeks, K.M. The mechanisms of RNA SHAPE chemistry. J. Am. Chem. Soc. 134, 6617–6624 (2012).
Wilkinson, K.A. et al. High-throughput SHAPE analysis reveals structures in HIV-1 genomic RNA strongly conserved across distinct biological states. PLoS Biol. 6, e96 (2008).
Gherghe, C. et al. Definition of a high-affinity Gag recognition structure mediating packaging of a retroviral RNA genome. Proc. Natl. Acad. Sci. 107, 19248–19253 (2010).
Archer, E.J. et al. Long-range architecture in a viral RNA genome. Biochemistry 52, 3182–3190 (2013).
Spitale, R.C. et al. RNA SHAPE analysis in living cells. Nat. Chem. Biol. 9, 18–20 (2013).
Tyrrell, J., McGinnis, J.L., Weeks, K.M. & Pielak, G.J. The cellular environment stabilizes adenine riboswitch RNA structure. Biochemistry 52, 8777–8785 (2013).
McGinnis, J.L. & Weeks, K.M. Ribosome RNA assembly intermediates visualized in living cells. Biochemistry 53, 3237–3247 (2014).
McGinnis, J.L. et al. In-cell SHAPE reveals that free 30S ribosome subunits are in the inactive state. Proc. Natl. Acad. Sci. 112, 2425–2430 (2015).
Mortimer, S.A. & Weeks, K.M. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J. Am. Chem. Soc. 129, 4144–4145 (2007).
Deigan, K.E., Li, T.W., Mathews, D.H. & Weeks, K.M. Accurate SHAPE-directed RNA structure determination. Proc. Natl. Acad. Sci. 106, 97–102 (2009).
Hajdin, C.E. et al. Accurate SHAPE-directed RNA secondary structure modeling, including pseudoknots. Proc. Natl. Acad. Sci. 110, 5498–5503 (2013).
Rice, G.M., Leonard, C.W. & Weeks, K.M. RNA secondary structure modeling at consistent high accuracy using differential SHAPE. RNA 20, 846–854 (2014).
Underwood, J.G. et al. FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat. Methods 7, 995–1001 (2010).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Lucks, J.B. et al. Multiplexed RNA structure characterization with selective 2′-hydroxyl acylation analyzed by primer extension sequencing (SHAPE-seq). Proc. Natl. Acad. Sci. 108, 11063–11068 (2011).
Incarnato, D., Neri, F., Anselmi, F. & Oliviero, S. Genome-wide profiling of mouse RNA secondary structures reveals key features of the mammalian transcriptome. Genome Biol. 15, 491 (2014).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J.S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Siegfried, N.A., Busan, S., Rice, G.M., Nelson, J.A.E. & Weeks, K.M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Vogel, J., Hess, W.R. & Börner, T. Precise branch point mapping and quantification of splicing intermediates. Nucleic Acids Res. 25, 2030–2031 (1997).
Ule, J., Jensen, K., Mele, A. & Darnell, R.B. CLIP: a method for identifying protein-RNA interaction sites in living cells. Methods 37, 376–386 (2005).
Mauger, D.M. et al. Functionally conserved architecture of hepatitis C virus RNA genomes. Proc. Natl. Acad. Sci. 112, 3692–3697 (2015).
Lavender, C.A., Gorelick, R.J. & Weeks, K.M. Structure-based alignment and consensus secondary structures for three HIV-related RNA genomes. PLoS Comput. Biol. 11, e1004230 (2015).
Reuter, J.S. & Mathews, D.H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
McCaskill, J.S. The equilibrium partition function and base pair binding probabilities for RNA secondary structure. Biopolymers 29, 1105–1119 (1990).
Mathews, D.H. Using an RNA secondary structure partition function to determine confidence in base pairs predicted by free energy minimization. RNA 10, 1178–1190 (2004).
Shannon, E. A mathematical theory of communication. Bell Syst. Tech. J. 27, 379–423 (1948).
Talkish, J., May, G., Lin, Y., Woolford, J.L. & McManus, C.J. Mod-seq: high-throughput sequencing for chemical probing of RNA structure. RNA 20, 713–720 (2014).
Staple, D.W. et al. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Taylor, A.I. et al. Catalysts from synthetic genetic polymers. Nature 518, 427–430 (2015).
Wang, B., Wilkinson, K.A. & Weeks, K.M. Complex ligand-induced conformational changes in tRNA(Asp) revealed by single-nucleotide resolution SHAPE chemistry. Biochemistry 47, 3454–3461 (2008).
Warner, K.D. et al. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Mortimer, S.A. & Weeks, K.M. C2′-endo nucleotides as molecular timers suggested by the folding of an RNA domain. Proc. Natl. Acad. Sci. 106, 15622–15627 (2009).
Duncan, C.D.S. & Weeks, K.M. Nonhierarchical ribonucleoprotein assembly suggests a strain-propagation model for protein-facilitated RNA folding. Biochemistry 49, 5418–5425 (2010).
Grohman, J.K. et al. A guanosine-centric mechanism for RNA chaperone function. Science 340, 190–195 (2013).
Raabe, C.A., Tang, T.-H., Brosius, J. & Rozhdestvensky, T.S. Biases in small RNA deep sequencing data. Nucleic Acids Res. 42, 1414–1426 (2014).
Weeks, K.M. Review toward all RNA structures, concisely. Biopolymers 103, 438–448 (2015).
Loughrey, D., Watters, K.E., Settle, A.H. & Lucks, J.B. SHAPE-Seq 2.0: systematic optimization and extension of high-throughput chemical probing of RNA secondary structure with next generation sequencing. Nucleic Acids Res. 42, e165 (2014).
Homan, P.J. et al. Single-molecule correlated chemical probing of RNA. Proc. Natl. Acad. Sci. 111, 13858–13863 (2014).
Nicholson, B.L. & White, K.A. Functional long-range RNA-RNA interactions in positive-strand RNA viruses. Nat. Rev. Microbiol. 12, 493–504 (2014).
Duncan, C.D.S. & Weeks, K.M. SHAPE analysis of long-range interactions reveals extensive and thermodynamically preferred misfolding in a fragile group I intron RNA. Biochemistry 47, 8504–8513 (2008).
McGinnis, J.L., Duncan, C.D. & Weeks, K.M. High-throughput SHAPE and hydroxyl radical analysis of RNA structure and ribonucleoprotein assembly. Methods Enzymol. 468, 67–89 (2009).
Watts, J.M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
Steen, K.-A., Rice, G.M. & Weeks, K.M. Fingerprinting noncanonical and tertiary RNA structures by differential SHAPE reactivity. J. Am. Chem. Soc. 134, 13160–13163 (2012).
Mortimer, S.A. & Weeks, K.M. Time-resolved RNA SHAPE chemistry. J. Am. Chem. Soc. 130, 16178–16180 (2008).
Bai, Y., Tambe, A., Zhou, K. & Doudna, J.A. RNA-guided assembly of Rev-RRE nuclear export complexes. Elife 3, e03656 (2014).
Steen, K.-A., Siegfried, N.A. & Weeks, K.M. Synthesis of 1-methyl-7-nitroisatoic anhydride (1M7). Protoc. Exch. 10.1038/protex.2011.255 (2011).
Turner, R., Shefer, K. & Ares, M. Safer one-pot synthesis of the 'SHAPE' reagent 1-methyl-7-nitroisatoic anhydride (1m7). RNA 19, 1857–1863 (2013).
Ye, J. et al. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13, 134 (2012).
Byun, Y. & Han, K. PseudoViewer: web application and web service for visualizing RNA pseudoknots and secondary structures. Nucleic Acids Res. 34, W416–W422 (2006).
Serganov, A., Polonskaia, A., Phan, A.T., Breaker, R.R. & Patel, D.J. Structural basis for gene regulation by a thiamine pyrophosphate-sensing riboswitch. Nature 441, 1167–1171 (2006).
Acknowledgements
This work was supported by grants from the US National Institutes of Health (NIH) (AI068462) and the National Science Foundation (MCB-1121024) to K.M.W. M.J.S. is a Graduate Research Fellow of the National Science Foundation (DGE-1144081). G.M.R. and M.J.S. were supported in part by an NIH training grant in molecular and cellular biophysics (T32 GM08570). N.A.S. was a Lineberger Postdoctoral Fellow in the Basic Sciences and a Ruth L. Kirschstein NRSA Fellow (F32 GM010169). We are indebted to many members of the Weeks laboratory who provided continuous and thoughtful feedback on the strategies and algorithms described here.
Author information
Authors and Affiliations
Contributions
N.A.S., S.B., G.M.R. and K.M.W. conceived and developed the original SHAPE-MaP strategy. M.J.S., S.B. and G.M.R. modified and updated the technology to its current state. All authors wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
N.S., S.B. and K.M.W. are listed on a pending PCT patent application that covers concepts in this manuscript.
Integrated supplementary information
Supplementary Figure 1 Ambiguously aligned deletion detection algorithm.
Example shows the ambiguous deletion detection algorithm applied to a single short read. (a) Reference sequence and aligned read. (b) The deletion is “collapsed” and the surrounding sequence stored. (c) The deletion is repositioned upstream and downstream of its Bowtie2-reported location, up to as many nucleotides as it is long. (d) The repositioned deletions are “collapsed” and the surrounding sequences compared to the original surrounding sequence. In any reposition, the absence of mismatches with the original surrounding sequence indicates an alternate valid deletion placement.
Supplementary Figure 2 Ambiguously aligned deletion removal.
(a) SHAPE reactivities from a portion of the bacterial 16S rRNA, identified as stops to primer extension by reverse transcription, read out by capillary electrophoresis using fluorescently-label primers. (b) SHAPE-MaP reactivities from the same region of 16S rRNA, obtained using the reverse transcription read-through SHAPE-MaP strategy, including the ambiguous deletion removal algorithm. (c) Reactivities from the same experiment, but analysed omitting ambiguous deletion removal. The orange arrow highlights a nucleotide reactivity inconsistent with the capillary electrophoresis experiment.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Methods 1 and 2 (PDF 587 kb)
Rights and permissions
About this article
Cite this article
Smola, M., Rice, G., Busan, S. et al. Selective 2′-hydroxyl acylation analyzed by primer extension and mutational profiling (SHAPE-MaP) for direct, versatile and accurate RNA structure analysis. Nat Protoc 10, 1643–1669 (2015). https://doi.org/10.1038/nprot.2015.103
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.103
This article is cited by
-
NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
Nature Communications (2024)
-
The SMN complex drives structural changes in human snRNAs to enable snRNP assembly
Nature Communications (2023)
-
Dynamical control enables the formation of demixed biomolecular condensates
Nature Communications (2023)
-
The p53 endoplasmic reticulum stress-response pathway evolved in humans but not in mice via PERK-regulated p53 mRNA structures
Cell Death & Differentiation (2023)
-
Systematic detection of tertiary structural modules in large RNAs and RNP interfaces by Tb-seq
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.