Abstract
Antibodies (or immunoglobulins) are crucial for defending organisms from pathogens, but they are also key players in many medical, diagnostic and biotechnological applications. The ability to predict their structure and the specific residues involved in antigen recognition has several useful applications in all of these areas. Over the years, we have developed or collaborated in developing a strategy that enables researchers to predict the 3D structure of antibodies with a very satisfactory accuracy. The strategy is completely automated and extremely fast, requiring only a few minutes (∼10 min on average) to build a structural model of an antibody. It is based on the concept of canonical structures of antibody loops and on our understanding of the way light and heavy chains pack together.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
21 November 2014
In the version of this article initially published online, the title was incorrect and was changed to read as follows: ‘Antibody modeling using the Prediction of ImmunoGlobulin Structure (PIGS) web server’. The error has been corrected for the PDF and HTML versions of this article.
References
Bassing, C.H., Swat, W. & Alt, F.W. The mechanism and regulation of chromosomal V(D)J recombination. Cell 109 (suppl.): S45–S55 (2002).
Li, Z., Woo, C.J., Iglesias-Ussel, M.D., Ronai, D. & Scharff, M.D. The generation of antibody diversity through somatic hypermutation and class switch recombination. Genes Dev. 18, 1–11 (2004).
Schatz, D.G. & Ji, Y. Recombination centres and the orchestration of V(D)J recombination. Nat. Rev. Immunol. 11, 251–263 (2011).
Padlan, E.A. Anatomy of the antibody molecule. Mol. Immunol. 31, 169–217 (1994).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Mariuzza, R.A., Phillips, S.E. & Poljak, R.J. The structural basis of antigen-antibody recognition. Annu. Rev. Biophys. Biophys. Chem. 16, 139–159 (1987).
Wu, T.T. & Kabat, E.A. An analysis of the sequences of the variable regions of Bence Jones proteins and myeloma light chains and their implications for antibody complementarity. J. Exp. Med. 132, 211–250 (1970).
Collis, A.V., Brouwer, A.P. & Martin, A.C. Analysis of the antigen combining site: correlations between length and sequence composition of the hypervariable loops and the nature of the antigen. J. Mol. Biol. 325, 337–354 (2003).
Kuroda, D., Shirai, H., Jacobson, M.P. & Nakamura, H. Computer-aided antibody design. Protein Eng. Des. Sel. 25, 507–521 (2012).
Pedotti, M., Simonelli, L., Livoti, E. & Varani, L. Computational docking of antibody-antigen complexes, opportunities and pitfalls illustrated by influenza hemagglutinin. Int. J. Mol. Sci. 12, 226–251 (2011).
Lippow, S.M., Wittrup, K.D. & Tidor, B. Computational design of antibody-affinity improvement beyond in vivo maturation. Nat. Biotechnol. 25, 1171–1176 (2007).
Kettleborough, C.A., Saldanha, J., Heath, V.J., Morrison, C.J. & Bendig, M.M. Humanization of a mouse monoclonal antibody by CDR-grafting: the importance of framework residues on loop conformation. Protein Eng. 4, 773–783 (1991).
Almagro, J.C. & Fransson, J. Humanization of antibodies. Front. Biosci. 13, 1619–1633 (2008).
Kuramochi, T., Igawa, T., Tsunoda, H. & Hattori, K. Humanization and simultaneous optimization of monoclonal antibody. Methods Mol. Biol. 1060, 123–137 (2014).
Lauer, T.M. et al. Developability index: a rapid in silico tool for the screening of antibody aggregation propensity. J. Pharm. Sci. 101, 102–115 (2012).
Marcatili, P. et al. Igs expressed by chronic lymphocytic leukemia B cells show limited binding-site structure variability. J. Immunol. 190, 5771–5778 (2013).
Zibellini, S. et al. Stereotyped patterns of B cell receptor in splenic marginal zone lymphoma. Haematologica 95, 1792–1796 (2010).
Padlan, E.A. & Davies, D.R. Variability of three-dimensional structure in immunoglobulins. Proc. Natl. Acad. Sci. USA 72, 819–823 (1975).
Poljak, R.J. et al. Three-dimensional structure and diversity of immunoglobulins. Cold Spring Harb. Symp. Quant. Biol. 41 Pt 2: 639–645 (1977).
Schroeder, H.W. Jr. & Cavacini, L. Structure and function of immunoglobulins. J. Allergy Clin. Immunol. 125, S41–S52 (2010).
Chothia, C., Novotny, J., Bruccoleri, R. & Karplus, M. Domain association in immunoglobulin molecules. The packing of variable domains. J. Mol. Biol. 186, 651–663 (1985).
Fiser, A., Do, R.K. & Sali, A. Modeling of loops in protein structures. Protein Sci. 9, 1753–1773 (2000).
Soto, C.S., Fasnacht, M., Zhu, J., Forrest, L. & Honig, B. Loop modeling: sampling, filtering, and scoring. Proteins 70, 834–843 (2008).
Chothia, C. & Lesk, A.M. Canonical structures for the hypervariable regions of immunoglobulins. J. Mol. Biol. 196, 901–917 (1987).
Chothia, C. et al. Conformations of immunoglobulin hypervariable regions. Nature 342, 877–883 (1989).
Shirai, H., Kidera, A. & Nakamura, H. Structural classification of CDR-H3 in antibodies. FEBS Lett. 399, 1–8 (1996).
Morea, V., Tramontano, A., Rustici, M., Chothia, C. & Lesk, A.M. Conformations of the third hypervariable region in the VH domain of immunoglobulins. J. Mol. Biol. 275, 269–294 (1998).
Oliva, B., Bates, P.A., Querol, E., Aviles, F.X. & Sternberg, M.J. Automated classification of antibody complementarity determining region 3 of the heavy chain (H3) loops into canonical forms and its application to protein structure prediction. J. Mol. Biol. 279, 1193–1210 (1998).
Shirai, H., Nakajima, N., Higo, J., Kidera, A. & Nakamura, H. Conformational sampling of CDR-H3 in antibodies by multicanonical molecular dynamics simulation. J. Mol. Biol. 278, 481–496 (1998).
Kim, S.T., Shirai, H., Nakajima, N., Higo, J. & Nakamura, H. Enhanced conformational diversity search of CDR-H3 in antibodies: role of the first CDR-H3 residue. Proteins 37, 683–696 (1999).
Shirai, H., Kidera, A. & Nakamura, H. H3-rules: identification of CDR-H3 structures in antibodies. FEBS Lett. 455, 188–197 (1999).
Kuroda, D., Shirai, H., Kobori, M. & Nakamura, H. Systematic classification of CDR-L3 in antibodies: implications of the light chain subtypes and the VL-VH interface. Proteins 75, 139–146 (2009).
Chothia, C. et al. Structural repertoire of the human VH segments. J. Mol. Biol. 227, 799–817 (1992).
Tomlinson, I.M., Cox, J.P., Gherardi, E., Lesk, A.M. & Chothia, C. The structural repertoire of the human V kappa domain. EMBO J. 14, 4628–4638 (1995).
Chailyan, A., Marcatili, P., Cirillo, D. & Tramontano, A. Structural repertoire of immunoglobulin λ light chains. Proteins 79, 1513–1524 (2011).
Tramontano, A., Chothia, C. & Lesk, A.M. Framework residue 71 is a major determinant of the position and conformation of the second hypervariable region in the VH domains of immunoglobulins. J. Mol. Biol. 215, 175–182 (1990).
Foote, J. & Winter, G. Antibody framework residues affecting the conformation of the hypervariable loops. J. Mol. Biol. 224, 487–499 (1992).
Martin, A.C. & Thornton, J.M. Structural families in loops of homologous proteins: automatic classification, modelling and application to antibodies. J. Mol. Biol. 263, 800–815 (1996).
Chothia, C., Gelfand, I. & Kister, A. Structural determinants in the sequences of immunoglobulin variable domain. J. Mol. Biol. 278, 457–479 (1998).
Decanniere, K., Muyldermans, S. & Wyns, L. Canonical antigen-binding loop structures in immunoglobulins: more structures, more canonical classes? J. Mol. Biol. 300, 83–91 (2000).
Vargas-Madrazo, E. & Paz-Garcia, E. Modifications to canonical structure sequence patterns: analysis for L1 and L3. Proteins 47, 250–254 (2002).
North, B., Lehmann, A. & Dunbrack, R.L. Jr. A new clustering of antibody CDR loop conformations. J. Mol. Biol. 406, 228–256 (2011).
De Wildt, R.M., Hoet, R.M., van Venrooij, W.J., Tomlinson, I.M. & Winter, G. Analysis of heavy and light chain pairings indicates that receptor editing shapes the human antibody repertoire. J. Mol. Biol. 285, 895–901 (1999).
Abhinandan, K.R. & Martin, A.C. Analysis and prediction of VH/VL packing in antibodies. Protein Eng. Des. Sel. 23, 689–697 (2010).
Jayaram, N., Bhowmick, P. & Martin, A.C. Germline VH/VL pairing in antibodies. Protein Eng. Des. Sel. 25, 523–529 (2012).
Dunbar, J., Fuchs, A., Shi, J. & Deane, C.M. ABangle: characterising the VH-VL orientation in antibodies. Protein Eng. Des. Sel. 26, 611–620 (2013).
Almagro, J.C. et al. Antibody modeling assessment. Proteins 79, 3050–3066 (2011).
Zhao, Z., Worthylake, D., LeCour, L. Jr., Maresh, G.A. & Pincus, S.H. Crystal structure and computational modeling of the Fab fragment from a protective anti-ricin monoclonal antibody. PLoS ONE 7, e52613 (2012).
Simonelli, L. et al. Rational engineering of a human anti-dengue antibody through experimentally validated computational docking. PLoS ONE 8, e55561 (2013).
Chemical Computing Group. Molecular Operating Environment http://www.chemcomp.com/MOE-Protein_and_Antibody_Modeling.html (2012).
Accelrys Software. Discovery Studio Modeling Environment, Release 3.5. http://accelrys.com/products/discovery-studio/antibody-modeling.html (2012).
Sircar, A., Kim, E.T. & Gray, J.J. RosettaAntibody: antibody variable region homology modeling server. Nucleic Acids Res. 37, W474–W479 (2009).
Almagro, J.C. et al. Antibody Engineering and Therapeutics Conference: the Annual Meeting of the Antibody Society. MAbs 817–825 (2013).
Wang, F. et al. Reshaping antibody diversity. Cell 153, 1379–1393 (2013).
Corti, D. & Lanzavecchia, A. Broadly neutralizing antiviral antibodies. Annu. Rev. Immunol. 31, 705–742 (2013).
Pellequer, J.L., Chen, S., Roberts, V.A., Tainer, J.A. & Getzoff, E.D. Unraveling the effect of changes in conformation and compactness at the antibody VL-VH interface upon antigen binding. J. Mol. Recognit. 12, 267–275 (1999).
Silverman, B.D. Using molecular principal axes for structural comparison: determining the tertiary changes of a Fab antibody domain induced by antigenic binding. BMC Struct. Biol. 7, 77 (2007).
Babor, M. & Kortemme, T. Multi-constraint computational design suggests that native sequences of germline antibody H3 loops are nearly optimal for conformational flexibility. Proteins 75, 846–858 (2009).
Sela-Culang, I., Alon, S. & Ofran, Y. A systematic comparison of free and bound antibodies reveals binding-related conformational changes. J. Immunol. 189, 4890–4899 (2012).
Marcatili, P., Rosi, A. & Tramontano, A. PIGS: automatic prediction of antibody structures. Bioinformatics 24, 1953–1954 (2008).
Whitelegg, N.R. & Rees, A.R. WAM: an improved algorithm for modelling antibodies on the web. Protein Eng. 13, 819–824 (2000).
Chailyan, A., Tramontano, A. & Marcatili, P. A database of immunoglobulins with integrated tools: DIGIT. Nucleic Acids Res. 40, D1230–D1234 (2012).
Olimpieri, P.P., Chailyan, A., Tramontano, A. & Marcatili, P. Prediction of site-specific interactions in antibody-antigen complexes: the proABC method and server. Bioinformatics 29, 2285–2291 (2013).
Ye, J., Ma, N., Madden, T.L. & Ostell, J.M. IgBLAST: an immunoglobulin variable domain sequence analysis tool. Nucleic Acids Res. 41, W34–W40 (2013).
Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Putnam, F.W., Shinoda, T., Titani, K. & Wikler, M. Immunoglobulin structure: variation in amino acid sequence and length of human λ light chains. Science 157, 1050–1053 (1967).
Al-Lazikani, B., Lesk, A.M. & Chothia, C. Standard conformations for the canonical structures of immunoglobulins. J. Mol. Biol. 273, 927–948 (1997).
Lefranc, M.P. et al. IMGT unique numbering for immunoglobulin and T cell receptor variable domains and Ig superfamily V-like domains. Dev. Comp. Immunol. 27, 55–77 (2003).
Almagro, J.C. et al. Second antibody modeling assessment (AMA-II). Proteins 82, 1553–1562 (2014).
Messih, M.A., Lepore, R., Marcatili, P. & Tramontano, A. Improving the accuracy of the structure prediction of the third hypervariable loop of the heavy chains of antibodies. Bioinformatics 30, 2733–2740 (2014).
Shirai, H. et al. High-resolution modeling of antibody structures by a combination of bioinformatics, expert knowledge, and molecular simulations. Proteins 82, 1624–1635 (2014).
Weitzner, B.D., Kuroda, D., Marze, N., Xu, J. & Gray, J.J. Blind prediction performance of RosettaAntibody 3.0: grafting, relaxation, kinematic loop modeling, and full CDR optimization. Proteins 82, 1611–1623 (2014).
Chailyan, A., Marcatili, P. & Tramontano, A. The association of heavy and light chain variable domains in antibodies: implications for antigen specificity. FEBS J. 278, 2858–2866 (2011).
Krivov, G.G., Shapovalov, M.V. & Dunbrack, R.L. Jr. Improved prediction of protein side-chain conformations with SCWRL4. Proteins 77, 778–795 (2009).
Ghiotto, F. et al. Remarkably similar antigen receptors among a subset of patients with chronic lymphocytic leukemia. J. Clin. Invest. 113, 1008–1016 (2004).
Zemla, A. LGA: a method for finding 3D similarities in protein structures. Nucleic Acids Res. 31, 3370–3374 (2003).
Zhang, Y. & Skolnick, J. Scoring function for automated assessment of protein structure template quality. Proteins 57, 702–710 (2004).
Damle, R.N. et al. Ig V gene mutation status and CD38 expression as novel prognostic indicators in chronic lymphocytic leukemia. Blood 94, 1840–1847 (1999).
Hamblin, T.J., Davis, Z., Gardiner, A., Oscier, D.G. & Stevenson, F.K. Unmutated Ig VH genes are associated with a more aggressive form of chronic lymphocytic leukemia. Blood 94, 1848–1854 (1999).
Grover, R.K. et al. A structurally distinct human mycoplasma protein that generically blocks antigen-antibody union. Science 343, 656–661 (2014).
Sanguineti, S. et al. Specific recognition of a DNA immunogen by its elicited antibody. J. Mol. Biol. 370, 183–195 (2007).
Acknowledgements
The authors are grateful to all the members of the Sapienza University of Rome Biocomputing Group for their help and assistance. We are thankful to the King Abdullah University of Science and Technology (KAUST) for financial support.
Author information
Authors and Affiliations
Contributions
A.T. conceived the study, participated in its design and coordination, and helped to draft the manuscript. P.M. developed the software. A.C. and P.P.O. contributed to the analysis and interpretation of data and to test the system. All authors participated in drafting the article and revising it critically.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
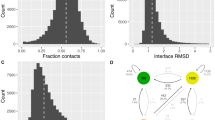
Supplementary Figure 1 RMSD value versus sequence similarity after cross-validation of the PIGS dataset.
The Cα atoms RMSD value for each of the 689 antibody structures in the PIGS dataset with the corresponding models obtained using a leave-one-out procedure is plotted against target–template sequence similarity.
Supplementary Figure 2 RMSD values for all the hypervariable loops except H3 after cross-validation of the PIGS dataset.
Distribution of the Cα RMSD calculated on all the hypervariable loops except H3 for the 689 antibody structures in the PIGS dataset with the corresponding models obtained using a leave-one-out procedure.
Supplementary Figure 3 RMSD values for loop H3 alone after cross-validation of the PIGS dataset.
Distribution of the Cα RMSD calculated on loop H3 alone for the 689 antibody structures in the PIGS dataset with the corresponding models obtained using a leave-one-out procedure.
Supplementary Figure 4 Superimposition of the models and the experimentally solved structure of 13PL antibody obtained following two different modeling strategies.
The cartoon representations of the hypervariable loops of the heavy (H) and light (L) chains obtained using the Same antibody and the Same canonical structure options are respectively blue- and red-colored. The solved structure is shown in grey. The figure was produced with PyMOL (The PyMOL Molecular Graphics System, Version 1.5.0.4 Schrödinger, LLC. http://www.pymol.org/).
Supplementary Figure 5 Results of the modeling of the structure of antibody ED-10.
a) The top templates obtained for monoclonal antibody ED-10 before the manual alignment; b) Automatic alignment generated for the heavy chain sequence; c) Manual alignment for the heavy chain; d) the top templates obtained for monoclonal antibody ED-10 after the manual alignment.
Supplementary information
Supplementary Figure 1
RMSD value versus sequence similarity after cross-validation of the PIGS dataset. (PDF 413 kb)
Supplementary Figure 2
RMSD values for all the hypervariable loops except H3 after cross-validation of the PIGS dataset. (PDF 272 kb)
Supplementary Figure 3
RMSD values for loop H3 alone after cross-validation of the PIGS dataset. (PDF 271 kb)
Supplementary Figure 4
Superimposition of the models and the experimentally solved structure of 13PL antibody obtained following two different modeling strategies. (PDF 288 kb)
Supplementary Figure 5
Results of the modeling of the structure of antibody ED-10. (PDF 2355 kb)
Rights and permissions
About this article
Cite this article
Marcatili, P., Olimpieri, P., Chailyan, A. et al. Antibody modeling using the Prediction of ImmunoGlobulin Structure (PIGS) web server. Nat Protoc 9, 2771–2783 (2014). https://doi.org/10.1038/nprot.2014.189
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.189
This article is cited by
-
32A9, a novel human antibody for designing an immunotoxin and CAR-T cells against glypican-3 in hepatocellular carcinoma
Journal of Translational Medicine (2020)
-
Automated shape-based clustering of 3D immunoglobulin protein structures in chronic lymphocytic leukemia
BMC Bioinformatics (2018)
-
Structure-function analysis and therapeutic efficacy of antibodies to fungal melanin for melanoma radioimmunotherapy
Scientific Reports (2018)
-
Rapid and accurate in silico solubility screening of a monoclonal antibody library
Scientific Reports (2017)
-
Superposition-free comparison and clustering of antibody binding sites: implications for the prediction of the nature of their antigen
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.