Abstract
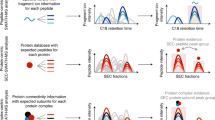
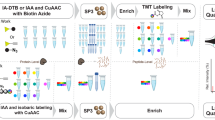
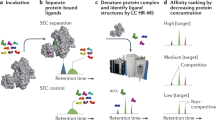
Ligand-receptor interactions represent essential biological triggers that regulate many diverse and important cellular processes. We have developed a discovery-based proteomic biochemical protocol that couples affinity purification with multidimensional liquid chromatographic tandem mass spectrometry (LCLC-MS/MS) and bioinformatic analysis. Compared with previous approaches, our analysis increases sensitivity, shortens analysis duration and boosts comprehensiveness. In this protocol, receptor extracellular domains are fused with the Fc region of IgG to generate fusion proteins that are purified from transfected HEK293T cells. These 'ecto-Fcs' are coupled to protein A beads and serve as baits for binding assays with prey proteins extracted from rodent brain. After capture, the affinity-purified proteins are digested into peptides and comprehensively analyzed by LCLC-MS/MS with ion-trap mass spectrometers. In 4 working days, this protocol can generate shortlists of candidate ligand-receptor protein-protein interactions. Our 'ecto-Fc MS' approach outperforms antibody-based approaches and provides a reproducible and robust framework for identifying extracellular ligand-receptor interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chen, Y., Fournier, A., Couvineau, A., Laburthe, M. & Amiranoff, B. Purification of a galanin receptor from pig brain. Proc. Natl. Acad. Sci. USA 90, 3845–3849 (1993).
Ichtchenko, K. et al. Neuroligin 1: a splice site-specific ligand for -neurexins. Cell 81, 435–443 (1995).
Sugita, S. et al. A stoichiometric complex of neurexins and dystroglycan in brain. J. Cell Biol. 154, 435–445 (2001).
Flanagan, J.G. & Vanderhaeghen, P. The ephrins and Eph receptors in neural development. Annu. Rev. Neurosci. 21, 309–345 (1998).
Savas, J.N., Stein, B.D., Wu, C.C. & Yates, J.R. III . Mass spectrometry accelerates membrane protein analysis. Trends Biochem. Sci. 36, 388–396 (2011).
Eng, J.K., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spect. 5, 976–989 (1994).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Frei, A.P. et al. Direct identification of ligand-receptor interactions on living cells and tissues. Nat. Biotechnol. 30, 997–1001 (2012).
Ko, J., Fuccillo, M.V., Malenka, R.C. & Sudhof, T.C. LRRTM2 functions as a neurexin ligand in promoting excitatory synapse formation. Neuron 64, 791–798 (2009).
de Wit, J. et al. Unbiased discovery of glypican as a receptor for LRRTM4 in regulating excitatory synapse development. Neuron 79, 696–711 (2013).
de Wit, J. et al. LRRTM2 interacts with Neurexin1 and regulates excitatory synapse formation. Neuron 64, 799–806 (2009).
O'Sullivan, M.L. et al. FLRT proteins are endogenous latrophilin ligands and regulate excitatory synapse development. Neuron 73, 903–910 (2012).
Neeper, M. et al. Cloning and expression of a cell surface receptor for advanced glycosylation end products of proteins. J. Biol. Chem. 267, 14998–15004 (1992).
Thompson, A. et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal. Chem. 75, 1895–1904 (2003).
Aruffo, A., Stamenkovic, I., Melnick, M., Underhill, C.B. & Seed, B. CD44 is the principal cell surface receptor for hyaluronate. Cell 61, 1303–1313 (1990).
Ushkaryov, Y.A. et al. Conserved domain structure of β-neurexins. Unusual cleaved signal sequences in receptor-like neuronal cell-surface proteins. J Biol. Chem. 269, 11987–11992 (1994).
Hacker, D.L. et al. Polyethyleneimine-based transient gene expression processes for suspension-adapted HEK-293E and CHO-DG44 cells. Protein Expr. Purif. 92, 67–76 (2013).
Link, A.J. et al. Direct analysis of protein complexes using mass spectrometry. Nat. Biotechnol. 17, 676–682 (1999).
Washburn, M.P., Wolters, D. & Yates, J.R. III. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Butko, M.T. et al. In vivo quantitative proteomics of somatosensory cortical synapses shows which protein levels are modulated by sensory deprivation. Proc. Natl. Acad. Sci. USA 110, E726–E735 (2013).
Niessen, S., McLeod, I. & Yates, J.R. III . HPLC separation of digested proteins and preparation for matrix-assisted laser desorption/ionization analysis. Cold Spring Harb. Protoc. 2006 (2006).
Xu, T., Wong, C.C., Kashina, A. & Yates, J.R. III. Identification of N-terminally arginylated proteins and peptides by mass spectrometry. Nat. Protoc. 4, 325–332 (2009).
Yates, J.R. III . et al. Toward objective evaluation of proteomic algorithms. Nat. Methods 9, 455–456 (2012).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Geer, L.Y. et al. Open mass spectrometry search algorithm. J. Proteome Res. 3, 958–964 (2004).
Craig, R. & Beavis, R.C. TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20, 1466–1467 (2004).
Carvalho, P.C., Fischer, J.S., Chen, E.I., Yates, J.R. III . & Barbosa, V.C. PatternLab for proteomics: a tool for differential shotgun proteomics. BMC Bioinformatics 9, 316 (2008).
Cox, J. et al. A practical guide to the MaxQuant computational platform for SILAC-based quantitative proteomics. Nat. Protoc. 4, 698–705 (2009).
Clough, T., Thaminy, S., Ragg, S., Aebersold, R. & Vitek, O. Statistical protein quantification and significance analysis in label-free LC-MS experiments with complex designs. BMC Bioinformatics 13 (suppl. 16): S6 (2012).
Liu, H., Sadygov, R.G. & Yates, J.R. III . A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal. Chem. 76, 4193–4201 (2004).
Paoletti, A.C. et al. Quantitative proteomic analysis of distinct mammalian Mediator complexes using normalized spectral abundance factors. Proc. Natl. Acad. Sci. USA 103, 18928–18933 (2006).
Maiolica, A., Borsotti, D. & Rappsilber, J. Self-made frits for nanoscale columns in proteomics. Proteomics 5, 3847–3850 (2005).
Liao, L., McClatchy, D.B. & Yates, J.R. Shotgun proteomics in neuroscience. Neuron 63, 12–26 (2009).
McDonald, W.H. et al. MS1, MS2, and SQT-three unified, compact, and easily parsed file formats for the storage of shotgun proteomic spectra and identifications. Rapid Commun. Mass Spectrom. 18, 2162–2168 (2004).
Tabb, D.L., McDonald, W.H. & Yates, J.R. III. DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J. Proteome Res. 1, 21–26 (2002).
Siddiqui, T.J. et al. An LRRTM4-HSPG complex mediates excitatory synapse development on dentate gyrus granule cells. Neuron 79, 680–695 (2013).
Acknowledgements
We thank M. O'Sullivan, B. Fonslow, B. Stein and J. Moresco, and members of the Yates and Ghosh laboratories for their feedback on the project, and S. Von Daake for suggestions on the manuscript. We thank T. Südhof (Stanford University) for his generosity in providing the original plasmids of β-NRXN1. This plasmid has been modified to accommodate many different genes and suit a variety of applications, but the original high-expression backbone remains unchanged. J.N.S. is supported by a US National Institutes of Health (NIH) Pathway to Independence Award K99DC013805-01. The Yates laboratory is supported by R01 MH068770, P41 GM103533, R01MH100175 and HHSN268201000035C grants from NIH. We also acknowledge support from NIH grants R01NS067216 and R01MH068578 to A.G. and NIH RO1-MH092906 to D.C., as well as the Robert Wood Johnson Foundation (grant no. 67038) for their support of the Child Health Institute of New Jersey. J.D.W. is funded by a NARSAD Young Investigator Award from the Brain and Behavior Research Foundation, a European Research Council (ERC) Starting Grant (no. 311083) and a Research Foundation Flanders (FWO) Odysseus Grant.
Author information
Authors and Affiliations
Contributions
J.N.S., J.D.W., A.G. and J.R.Y. contributed to the conception and design of the project; J.N.S., J.D.W., D.C. and R.Z. performed the experiments; and J.N.S., J.D.W. and D.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Savas, J., De Wit, J., Comoletti, D. et al. Ecto-Fc MS identifies ligand-receptor interactions through extracellular domain Fc fusion protein baits and shotgun proteomic analysis. Nat Protoc 9, 2061–2074 (2014). https://doi.org/10.1038/nprot.2014.140
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.140
This article is cited by
-
Synapse type-specific proteomic dissection identifies IgSF8 as a hippocampal CA3 microcircuit organizer
Nature Communications (2020)
-
Specification of synaptic connectivity by cell surface interactions
Nature Reviews Neuroscience (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.