Abstract
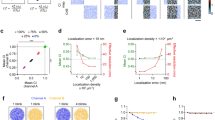
The distinctive distributions of proteins within subcellular compartments both at steady state and during signaling events have essential roles in cell function. Here we describe a method for delineating the complex arrangement of proteins within subcellular structures visualized using point-localization superresolution (PL-SR) imaging. The approach, called pair correlation photoactivated localization microscopy (PC-PALM), uses a pair-correlation algorithm to precisely identify single molecules in PL-SR imaging data sets, and it is used to decipher quantitative features of protein organization within subcellular compartments, including the existence of protein clusters and the size, density and number of proteins in these clusters. We provide a step-by-step protocol for PC-PALM, illustrating its analysis capability for four plasma membrane proteins tagged with photoactivatable GFP (PAGFP). The experimental steps for PC-PALM can be carried out in 3 d and the analysis can be done in ∼6–8 h. Researchers need to have substantial experience in single-molecule imaging and statistical analysis to conduct the experiments and carry out this analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Patterson, G., Davidson, M., Manley, S. & Lippincott-Schwartz, J. Superresolution imaging using single-molecule localization. Annu. Rev. Phys. Chem. 61, 345–367 (2010).
Toomre, D. & Bewersdorf, J. A new wave of cellular imaging. Annu. Rev. Cell Dev. Biol. 26, 285–314 (2010).
Patterson, G.H. Fluorescence microscopy below the diffraction limit. Semin. Cell Dev. Biol. 20, 886–893 (2009).
Flors, C. & Earnshaw, W.C. Super-resolution fluorescence microscopy as a tool to study the nanoscale organization of chromosomes. Curr. Opin. Chem. Biol. 15, 838–844 (2011).
Sigrist, S.J. & Sabatini, B.L. Optical super-resolution microscopy in neurobiology. Curr. Opin. Neurobiol. 22, 86–93 (2012).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S.T., Girirajan, T.P.K. & Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M.J., Bates, M. & Zhuang, X.W. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Wombacher, R. et al. Live-cell super-resolution imaging with trimethoprim conjugates. Nat. Methods 7, 717–719 (2010).
Folling, J. et al. Fluorescence nanoscopy by ground-state depletion and single-molecule return. Nat. Methods 5, 943–945 (2008).
Burnette, D.T., Sengupta, P., Dai, Y.H., Lippincott-Schwartz, J. & Kachar, B. Bleaching/blinking assisted localization microscopy for superresolution imaging using standard fluorescent molecules. Proc. Natl. Acad. Sci. USA 108, 21081–21086 (2011).
Simonson, P.D., Rothenberg, E. & Selvin, P.R. Single-molecule-based super-resolution images in the presence of multiple fluorophores. Nano Lett. 11, 5090–5096 (2011).
Thompson, R.E., Larson, D.R. & Webb, W.W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Mortensen, K.I., Churchman, L.S., Spudich, J.A. & Flyvbjerg, H. Optimized localization analysis for single-molecule tracking and super-resolution microscopy. Nat. Methods 7, 377–381 (2010).
Kanchanawong, P. et al. Nanoscale architecture of integrin-based cell adhesions. Nature 468, 580–584 (2010).
Loschberger, A. et al. Super-resolution imaging visualizes the eightfold symmetry of gp210 proteins around the nuclear pore complex and resolves the central channel with nanometer resolution. J. Cell Sci. 125, 570–575 (2012).
Kopek, B.G., Shtengel, G., Xu, C.S., Clayton, D.A. & Hess, H.F. Correlative 3D superresolution fluorescence and electron microscopy reveal the relationship of mitochondrial nucleoids to membranes. Proc. Natl. Acad. Sci. USA 109, 6136–6141 (2012).
Sengupta, P. et al. Probing protein heterogeneity in the plasma membrane using PALM and pair correlation analysis. Nat. Methods 8, 969–975 (2011).
Gould, T.J., Verkhusha, V.V. & Hess, S.T. Imaging biological structures with fluorescence photoactivation localization microscopy. Nat. Protoc. 4, 291–308 (2009).
van de Linde, S. et al. Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat. Protoc. 6, 991–1009 (2011).
Sengupta, P. & Lippincott-Schwartz, J. Quantitative analysis of photoactivated localization microscopy (PALM) datasets using pair-correlation analysis. Bioessays 34, 396–405 (2012).
Annibale, P., Vanni, S., Scarselli, M., Rothlisberger, U. & Radenovic, A. Identification of clustering artifacts in photoactivated localization microscopy. Nat. Methods 8, 527–528 (2011).
Annibale, P., Vanni, S., Scarselli, M., Rothlisberger, U. & Radenovic, A. Quantitative photo activated localization microscopy: unraveling the effects of photoblinking. PLoS ONE 6, e22678 (2011).
Kiskowski, M.A., Hancock, J.F. & Kenworthy, A.K. On the use of Ripley's K-function and its derivatives to analyze domain size. Biophys. J. 97, 1095–1103 (2009).
Owen, D.M. et al. PALM imaging and cluster analysis of protein heterogeneity at the cell surface. J. Biophotonics 3, 446–454 (2010).
Williamson, D.J. et al. Pre-existing clusters of the adaptor Lat do not participate in early T cell signaling events. Nat. Immunol. 12, 655–662 (2011).
Dickson, R.M., Cubitt, A.B., Tsien, R.Y. & Moerner, W.E. On/off blinking and switching behaviour of single molecules of green fluorescent protein. Nature 388, 355–358 (1997).
Schwille, P., Kummer, S., Heikal, A.A., Moerner, W.E. & Webb, W.W. Fluorescence correlation spectroscopy reveals fast optical excitation-driven intramolecular dynamics of yellow fluorescent proteins. Proc. Natl. Acad. Sci. USA 97, 151–156 (2000).
Ha, T. & Tinnefeld, P. Photophysics of fluorescent probes for single-molecule biophysics and super-resolution imaging. Annu. Rev. Phys. Chem. 63, 595–617 (2012).
Smith, C.S., Joseph, N., Rieger, B. & Lidke, K.A. Fast, single-molecule localization that achieves theoretically minimum uncertainty. Nat. Methods 7, 373–375 (2010).
Holden, S.J., Uphoff, S. & Kapanidis, A.N. DAOSTORM: an algorithm for high-density super-resolution microscopy. Nat. Methods 8, 279–280 (2011).
Huang, F., Schwartz, S.L., Byars, J.M. & Lidke, K.A. Simultaneous multiple-emitter fitting for single molecule super-resolution imaging. Biomed. Opt. Express 2, 1377–1393 (2011).
Quan, T. et al. High-density localization of active molecules using Structured Sparse Model and Bayesian Information Criterion. Opt. Express 19, 16963–16974 (2011).
Cox, S. et al. Bayesian localization microscopy reveals nanoscale podosome dynamics. Nat. Methods 9, 195–200 (2012).
Wang, Y., Quan, T., Zeng, S. & Huang, Z.L. PALMER: a method capable of parallel localization of multiple emitters for high-density localization microscopy. Opt. Express 20, 16039–16049 (2012).
Veatch, S.L. et al. Correlation functions quantify super-resolution images and estimate apparent clustering due to over-counting. PLoS ONE 7, e31457 (2012).
Veatch, S.L., Chiang, E.N., Sengupta, P., Holowka, D.A. & Baird, B.A. Quantitative Nanoscale analysis of IgE-Fc RI clustering and coupling to early signaling proteins. J. Phys. Chem. B 116, 6923–6935 (2012).
Tanaka, K.A. et al. Membrane molecules mobile even after chemical fixation. Nat. Methods 7, 865–866 (2010).
Testa, I., Mazza, D., Barozzi, S., Faretta, M. & Diaspro, A. Blue-light (488 nm)-irradiation-induced photoactivation of the photoactivatable green fluorescent protein. Appl. Phys. Lett. 91, 133902–133904 (2007).
Subach, F.V. et al. Photoactivatable mCherry for high-resolution two-color fluorescence microscopy. Nat. Methods 6, 153–159 (2009).
Acknowledgements
We thank G. Patterson and Y. Fu (National Institute of Biomedical Imaging and Bioengineering) for DNA constructs and purified proteins, and H. Hess and G. Stengel (Howard Hughes Medical Institute, Janelia Farm Research Campus) for analysis software and valuable discussion.
Author information
Authors and Affiliations
Contributions
P.S. and T.J.-T. designed and performed the experiments, analyzed the data and wrote the paper; J.L.-S. designed the experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Data
Compute_Paircorrelation. Matlab code for computation of autocorrelation and cross-correlation functions from binary images of single molecule spatial distribution. (PDF 205 kb)
Rights and permissions
About this article
Cite this article
Sengupta, P., Jovanovic-Talisman, T. & Lippincott-Schwartz, J. Quantifying spatial organization in point-localization superresolution images using pair correlation analysis. Nat Protoc 8, 345–354 (2013). https://doi.org/10.1038/nprot.2013.005
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2013.005
This article is cited by
-
Motion of VAPB molecules reveals ER–mitochondria contact site subdomains
Nature (2024)
-
Determination of association equilibrium constant from single molecule fluorescence localization microscopy
Photochemical & Photobiological Sciences (2022)
-
Rapid statistical discrimination of fluorescence images of T cell receptors on immobilizing surfaces with different coating conditions
Scientific Reports (2021)
-
Potential quality improvement of stochastic optical localization nanoscopy images obtained by frame by frame localization algorithms
Scientific Reports (2020)
-
moxMaple3: a Photoswitchable Fluorescent Protein for PALM and Protein Highlighting in Oxidizing Cellular Environments
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.