Abstract
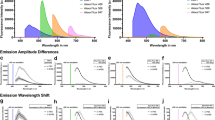
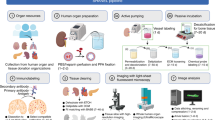
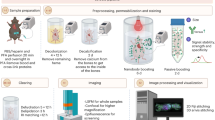
The examination of tissue histology by light microscopy is a fundamental tool for investigating the structure and function of organs under normal and disease states. Many current techniques for tissue sectioning, imaging and analysis are time-consuming, and they present major limitations for 3D tissue reconstruction. The introduction of methods to achieve the optical clearing and subsequent light-sheet laser scanning of entire transparent organs without sectioning represents a major advance in the field. We recently developed a highly reproducible and versatile clearing procedure called 3D imaging of solvent-cleared organs, or 3DISCO, which is applicable to diverse tissues including brain, spinal cord, immune organs and tumors. Here we describe a detailed protocol for performing 3DISCO and present its application to various microscopy techniques, including example results from various mouse tissues. The tissue clearing takes as little as 3 h, and imaging can be completed in ∼45 min. 3DISCO is a powerful technique that offers 3D histological views of tissues in a fraction of the time and labor required to complete standard histology studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Steward, O., Zheng, B. & Tessier-Lavigne, M. False resurrections: distinguishing regenerated from spared axons in the injured central nervous system. J. Comp. Neurol. 459, 1–8 (2003).
Tuchin, V. Tissue Optics: Light Scattering Methods and Instruments for Medical Diagnosis (SPIE Press, 2007).
Boas, D. A fundamental limitation of linearized algorithms for diffuse optical tomography. Opt. Express 1, 404–413 (1997).
Genina, E.A., Bashkatov, A.N. & Tuchin, V.V. Tissue optical immersion clearing. Expert Rev. Med. Devices 7, 825–842 (2010).
Spalteholz, W. Über das Durchsichtigmachen von menschlichen und tierischen Präparaten (S. Hierzel, 1914).
Keller, P.J. & Dodt, H.U. Light sheet microscopy of living or cleared specimens. Curr. Opin. Neurobiol. 22, 138–143 (2012).
Zhao, Q.L. et al. [In vivo reflectance spectroscopy study of different clearing agents on human skin optical clearing]. Guang Pu Xue Yu Guang Pu Fen Xi 31, 302–307 (2011).
Bucher, D., Scholz, M., Stetter, M., Obermayer, K. & Pfluger, H.J. Correction methods for three-dimensional reconstructions from confocal images: I. Tissue shrinking and axial scaling. J. Neurosci. Methods 100, 135–143 (2000).
Gerger, A. et al. Diagnostic applicability of in vivo confocal laser scanning microscopy in melanocytic skin tumors. J. Invest. Dermatol. 124, 493–498 (2005).
Dodt, H.U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Erturk, A. et al. Three-dimensional imaging of the unsectioned adult spinal cord to assess axon regeneration and glial responses after injury. Nat. Med. 18, 166–171 (2012).
Becker, K., Jährling, N., Kramer, E.R., Schnorrer, F. & Dodt, H.U. Ultramicroscopy: 3D reconstruction of large microscopical specimens. J. Biophotonics 1, 36–42 (2008).
Jährling, N. Applications of Ultramicroscopy to Neurobiology and Methodological Improvements. Ph.D. thesis, Universität Oldenburg, 2011. (Springer, 2012, in press).
Becker, K., Jährling, N., Saghafi, S., Weiler, R. & Dodt, H.U. Chemical clearing and dehydration of GFP expressing mouse brains. PLoS ONE 7, e33916 (2012).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Jährling, N., Becker, K., Schonbauer, C., Schnorrer, F. & Dodt, H.U. Three-dimensional reconstruction and segmentation of intact Drosophila by ultramicroscopy. Front. Syst. Neurosci. 4, 1 (2010).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E.H. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Fuchs, E., Jaffe, J., Long, R. & Azam, F. Thin laser light sheet microscope for microbial oceanography. Opt. Express 10, 145–154 (2002).
Saghafi, S. et al. Image enhancement in ultramicroscopy by improved laser light sheets. J. Biophotonics 3, 686–695 (2010).
Yokomizo, T. et al. Whole-mount three-dimensional imaging of internally localized immunostained cells within mouse embryos. Nat. Protoc. 7, 421–431 (2012).
Nolte, C. et al. GFAP promoter-controlled EGFP-expressing transgenic mice: a tool to visualize astrocytes and astrogliosis in living brain tissue. Glia 33, 72–86 (2001).
Jung, S. et al. Analysis of fractalkine receptor CX(3)CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Borrell, V., Yoshimura, Y. & Callaway, E.M. Targeted gene delivery to telencephalic inhibitory neurons by directional in utero electroporation. J. Neurosci. Methods 143, 151–158 (2005).
Miyamichi, K. et al. Cortical representations of olfactory input by trans-synaptic tracing. Nature 472, 191–196 (2011).
Conte, W.L., Kamishina, H. & Reep, R.L. Multiple neuroanatomical tract-tracing using fluorescent Alexa Fluor conjugates of cholera toxin subunit B in rats. Nat. Protoc. 4, 1157–1166 (2009).
Keller, P.J. & Stelzer, E.H. Quantitative in vivo imaging of entire embryos with digital scanned laser light sheet fluorescence microscopy. Curr. Opin. Neurobiol. 18, 624–632 (2008).
Jährling, N., Becker, K. & Dodt, H.U. 3D-reconstruction of blood vessels by ultramicroscopy. Organogenesis 5, 227–230 (2009).
Gray, D.C. et al. pHUSH: a single vector system for conditional gene expression. BMC Biotechnol. 7, 61 (2007).
Acknowledgements
We thank M. Chang and F. Yeh for critically reading the manuscript, and C. Chalouni, H. Ngu and L. Komuves for assistance with large-scale imaging microscopy and macros. We thank J. Sanes (Washington University in St. Louis) for the Thy-1 GFP (GFP-M) line, T. Jacks (Massachusetts Institute of Technology) for KrasLSLG12D mice, H. Ploegh (Massachusetts Institute of Technology) for MHCII-GFP mice, M. Nussenzweig (Rockefeller University) for CD11c YFP-Venus mice, and D.R. Littman (New York University) for CX3CR1-GFP mice. We thank C. Murriel (Genentech) for providing the lung tissues and C. Sakanaka (Genentech) for providing mammary gland tissues. The work is supported by Genentech and the Hertie foundation. N. Jährling was supported by the Theodor Körner Fonds.
Author information
Authors and Affiliations
Contributions
A.E. initiated the current project, designed and performed all the experiments, analyzed the data, and made all the figures and videos. A.E. and M.S. wrote the paper. C.D.H. prepared the immune organs and J.G.E. provided transgenic mice. K.B. and N.J. developed and performed various clearing protocols. A.E., K.B., N.J., C.P.M., F.H., F.B., M.S. and H.-U.D. contributed to the development of the clearing protocol at various stages. H.-U.D. led the development of the ultramicroscopy and clearing technology. All authors edited the paper.
Corresponding author
Ethics declarations
Competing interests
A.E., C.D.H., J.G.E. and M.S. are employees of Genentech, a member of the Roche Group.
Supplementary information
Supplementary video 1
A cleared spinal cord tissue of a GFP-M mouse imaged with 2-photon microscopy. In this high resolution scans, the individual axons can be readily traced. (MOV 15907 kb)
Supplementary video 2
The simulation of a scan through entire cleared brain from a GFP-M mouse in two different orientations: horizontal and sagittal. The bottom two windows show the horizontal scan in higher magnifications (left showing the hippocampus and right showing the cerebellum). (MOV 9533 kb)
Supplementary video 3
3D reconstruction and animation of the hippocampus from a GFP-M mouse. (MOV 9680 kb)
Supplementary video 4
3D reconstruction and animation of the spinal cord vasculature from a wild type mouse traced with lectin-FITC labeling. (MOV 18079 kb)
Rights and permissions
About this article
Cite this article
Ertürk, A., Becker, K., Jährling, N. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat Protoc 7, 1983–1995 (2012). https://doi.org/10.1038/nprot.2012.119
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2012.119
This article is cited by
-
Reflective multi-immersion microscope objectives inspired by the Schmidt telescope
Nature Biotechnology (2024)
-
Virtual reality-empowered deep-learning analysis of brain cells
Nature Methods (2024)
-
Whole-body cellular mapping in mouse using standard IgG antibodies
Nature Biotechnology (2024)
-
Current and future applications of light-sheet imaging for identifying molecular and developmental processes in autism spectrum disorders
Molecular Psychiatry (2024)
-
TSA-PACT: a method for tissue clearing and immunofluorescence staining on zebrafish brain with improved sensitivity, specificity and stability
Cell & Bioscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.