Abstract
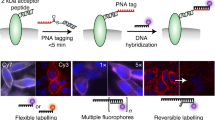
In this paper, we provide a general protocol for labeling proteins with the membrane-permeant fluorogenic biarsenical dye fluorescein arsenical hairpin binder–ethanedithiol (FlAsH-EDT2). Generation of the tetracysteine-tagged protein construct by itself is not described, as this is a protein-specific process. This method allows site-selective labeling of proteins in living cells and has been applied to a wide variety of proteins and biological problems. We provide here a generally applicable labeling procedure and discuss the problems that can occur as well as general considerations that must be taken into account when designing and implementing the procedure. The method can even be applied to proteins with expression below 1 pmol mg−1 of protein, such as G protein–coupled receptors, and it can be used to study the intracellular localization of proteins as well as functional interactions in fluorescence resonance energy transfer experiments. The labeling procedure using FlAsH-EDT2 as described takes 2–3 h, depending on the number of samples to be processed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Sletten, E.M. & Bertozzi, C.R. Bioorthogonal chemistry: fishing for selectivity in a sea of functionality. Angew. Chem. Int. Ed. Engl. 48, 6974–6998 (2009).
Shaner, N.C., Steinbach, P.A. & Tsien, R.Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Giepmans, B.N., Adams, S.R., Ellisman, M.H. & Tsien, R.Y. The fluorescent toolbox for assessing protein location and function. Science 312, 217–224 (2006).
Gautier, A. et al. An engineered protein tag for multiprotein labeling in living cells. Chem. Biol. 15, 128–136 (2008).
Gronemeyer, T., Godin, G. & Johnsson, K. Adding value to fusion proteins through covalent labelling. Curr. Opin. Biotechnol. 16, 453–458 (2005).
Keppler, A., Pick, H., Arrivoli, C., Vogel, H. & Johnsson, K. Labeling of fusion proteins with synthetic fluorophores in live cells. Proc. Natl. Acad. Sci. USA 101, 9955–9959 (2004).
Maurel, D. et al. Cell-surface protein-protein interaction analysis with time-resolved FRET and snap-tag technologies: application to GPCR oligomerization. Nat. Methods 5, 561–567 (2008).
Miller, L.W., Cai, Y., Sheetz, M.P. & Cornish, V.W. In vivo protein labeling with trimethoprim conjugates: a flexible chemical tag. Nat. Methods 2, 255–257 (2005).
Los, G.V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
George, N., Pick, H., Vogel, H., Johnsson, N. & Johnsson, K. Specific labeling of cell surface proteins with chemically diverse compounds. J. Am. Chem. Soc. 126, 8896–8897 (2004).
Soh, N. Selective chemical labeling of proteins with small fluorescent molecules based on metal-chelation methodology. Sensors 8, 1004–1102 (2008).
Hauser, C.T. & Tsien, R.Y. A hexahistidine-Zn2+-dye label reveals STIM1 surface exposure. Proc. Natl. Acad. Sci. USA 104, 3693–3697 (2007).
Kapanidis, A.N., Ebright, Y.W. & Ebright, R.H. Site-specific incorporation of fluorescent probes into protein: hexahistidine-tag-mediated fluorescent labeling with Ni2+:nitrilotriacetic acid (n)-fluorochrome conjugates. J. Am. Chem. Soc. 123, 12123–12125 (2001).
Guignet, E.G., Hovius, R. & Vogel, H. Reversible site-selective labeling of membrane proteins in live cells. Nat. Biotechnol. 22, 440–444 (2004).
Litowski, J.R. & Hodges, R.S. Designing heterodimeric two-stranded alpha-helical coiled-coils. Effects of hydrophobicity and alpha-helical propensity on protein folding, stability, and specificity. J. Biol. Chem. 277, 37272–37279 (2002).
Yano, Y. et al. Coiled-coil tag-probe system for quick labeling of membrane receptors in living cell. ACS Chem. Biol. 3, 341–345 (2008).
Griffin, B.A., Adams, S.R. & Tsien, R.Y. Specific covalent labeling of recombinant protein molecules inside live cells. Science 281, 269–272 (1998).
Gaietta, G. et al. Multicolor and electron microscopic imaging of connexin trafficking. Science 296, 503–507 (2002).
Stroffekova, K., Proenza, C. & Beam, K.G. The protein-labeling reagent FLASH-EDT2 binds not only to CCXXCC motifs but also non-specifically to endogenous cysteine-rich proteins. Pflügers Arch. 442, 859–866 (2001).
Adams, S.R. et al. New biarsenical ligands and tetracysteine motifs for protein labeling in vitro and in vivo: synthesis and biological applications. J. Am. Chem. Soc. 124, 6063–6076 (2002).
Martin, B.R., Giepmans, B.N., Adams, S.R. & Tsien, R.Y. Mammalian cell-based optimization of the biarsenical-binding tetracysteine motif for improved fluorescence and affinity. Nat. Biotechnol. 23, 1308–1314 (2005).
Wang, T., Yan, P., Squier, T.C. & Mayer, M.U. Prospecting the proteome: identification of naturally occurring binding motifs for biarsenical probes. Chembiochem 8, 1937–1940 (2007).
Spagnuolo, C.C., Vermeij, R.J. & Jares-Erijman, E.A. Improved photostable FRET-competent biarsenical-tetracysteine probes based on fluorinated fluoresceins. J. Am. Chem. Soc. 128, 12040–12041 (2006).
Adams, S.R. & Tsien, R.Y. Preparation of the membrane-permeant biarsenicals FlAsH-EDT2 and ReAsH-EDT2 for fluorescent labeling of tetracysteine-tagged proteins. Nat. Protoc. 3, 1527–1534 (2008).
Madani, F. et al. Hairpin structure of a biarsenical-tetracysteine motif determined by NMR spectroscopy. J. Am. Chem. Soc. 131, 4613–4615 (2009).
Martin, B.R., Deerinck, T.J., Ellisman, M.H., Taylor, S.S. & Tsien, R.Y. Isoform-specific PKA dynamics revealed by dye-triggered aggregation and DAKAP1alpha-mediated localization in living cells. Chem. Biol. 14, 1031–1042 (2007).
Chaumont, S. & Khakh, B.S. Patch-clamp coordinated spectroscopy shows P2X2 receptor permeability dynamics require cytosolic domain rearrangements but not Panx-1 channels. Proc. Natl. Acad. Sci. USA 105, 12063–12068 (2008).
Hoffmann, C. et al. A FlAsH-based FRET approach to determine G protein-coupled receptor activation in living cells. Nat. Methods 2, 171–176 (2005).
Nikolaev, V.O., Hoffmann, C., Bünemann, M., Lohse, M.J. & Vilardaga, J.P. Molecular basis of partial agonism at the neurotransmitter a2A-adrenergic receptor and Gi-protein heterotrimer. J. Biol. Chem. 281, 24506–24511 (2006).
Böhme, I., Morl, K., Bamming, D., Meyer, C. & Beck-Sickinger, A.G. Tracking of human Y receptors in living cells—a fluorescence approach. Peptides 28, 226–234 (2007).
Ju, W. et al. Activity-dependent regulation of dendritic synthesis and trafficking of AMPA receptors. Nat. Neurosci. 7, 244–253 (2004).
Enninga, J., Mounier, J., Sansonetti, P. & Tran Van Nhieu, G. Secretion of type III effectors into host cells in real time. Nat. Methods 2, 959–965 (2005).
Ignatova, Z. & Gierasch, L.M. Monitoring protein stability and aggregation in vivo by real-time fluorescent labeling. Proc. Natl. Acad. Sci. USA 101, 523–528 (2004).
Rudner, L. et al. Dynamic fluorescent imaging of human immunodeficiency virus type 1 gag in live cells by biarsenical labeling. J. Virol. 79, 4055–4065 (2005).
Das, S.C., Panda, D., Nayak, D. & Pattnaik, A.K. Biarsenical labeling of vesicular stomatitis virus encoding tetracysteine-tagged m protein allows dynamic imaging of m protein and virus uncoating in infected cells. J. Virol. 83, 2611–2622 (2009).
Venken, K.J. et al. Recombineering-mediated tagging of Drosophila genomic constructs for in vivo localization and acute protein inactivation. Nucleic Acids Res. 36, e114 (2008).
Tour, O., Meijer, R.M., Zacharias, D.A., Adams, S.R. & Tsien, R.Y. Genetically targeted chromophore-assisted light inactivation. Nat. Biotechnol. 21, 1505–1508 (2003).
Marek, K.W. & Davis, G.W. Transgenically encoded protein photoinactivation (FlAsH-FALI): acute inactivation of synaptotagmin I. Neuron 36, 805–813 (2002).
Roberti, M.J., Bertoncini, C.W., Klement, R., Jares-Erijman, E.A. & Jovin, T.M. Fluorescence imaging of amyloid formation in living cells by a functional, tetracysteine-tagged alpha-synuclein. Nat. Methods 4, 345–351 (2007).
Chen, B., Mayer, M.U., Markillie, L.M., Stenoien, D.L. & Squier, T.C. Dynamic motion of helix A in the amino-terminal domain of calmodulin is stabilized upon calcium activation. Biochemistry 44, 905–914 (2005).
Jost, C.A., Reither, G., Hoffmann, C. & Schultz, C. Contribution of fluorophores to protein kinase C FRET probe performance. Chembiochem 9, 1379–1384 (2008).
Zürn, A. et al. Fluorescence resonance energy transfer analysis of a2A-adrenergic receptor activation reveals distinct agonist-specific conformational changes. Mol. Pharmacol. 75, 534–541 (2009).
Luedtke, N.W., Dexter, R.J., Fried, D.B. & Schepartz, A. Surveying polypeptide and protein domain conformation and association with FlAsH and ReAsH. Nat. Chem. Biol. 3, 779–784 (2007).
Chen, B., Cao, H., Yan, P., Mayer, M.U. & Squier, T.C. Identification of an orthogonal peptide binding motif for biarsenical multiuse affinity probes. Bioconjug. Chem. 18, 1259–1265 (2007).
Zürn, A. et al. Site-specific, orthogonal labeling of proteins in intact cells with two small biarsenical fluorophores. Bioconjug. Chem. 21, 853–859 (2010).
Van Engelenburg, S.B., Nahreini, T. & Palmer, A.E. FACS-based selection of tandem tetracysteine peptides with improved ReAsH brightness in live cells. Chembiochem 11, 489–493 (2010).
Andresen, M., Schmitz-Salue, R. & Jakobs, S. Short tetracysteine tags to beta-tubulin demonstrate the significance of small labels for live cell imaging. Mol. Biol. Cell 15, 5616–5622 (2004).
Langhorst, M.F., Genisyuerek, S. & Stuermer, C.A. Accumulation of FlAsH/Lumio Green in active mitochondria can be reversed by b-mercaptoethanol for specific staining of tetracysteine-tagged proteins. Histochem Cell Biol. 125, 743–747 (2006).
Tour, O. et al. Calcium Green FlAsH as a genetically targeted small-molecule calcium indicator. Nat. Chem. Biol. 3, 423–431 (2007).
Gaietta, G.M. et al. Golgi twins in late mitosis revealed by genetically encoded tags for live cell imaging and correlated electron microscopy. Proc. Natl. Acad. Sci. USA 103, 17777–17782 (2006).
Griffin, B.A., Adams, S.R., Jones, J. & Tsien, R.Y. Fluorescent labeling of recombinant proteins in living cells with FlAsH. Methods Enzymol. 327, 565–578 (2000).
Campbell, R.E. Fluorescent-protein-based biosensors: modulation of energy transfer as a design principle. Anal. Chem. 81, 5972–5979 (2009).
Maier-Peuschel, M. et al. A fluorescence resonance energy transfer-based M2 muscarinic receptor sensor reveals rapid kinetics of allosteric modulation. J. Biol. Chem. 285, 8793–8800 (2010).
Robia, S.L., Flohr, N.C. & Thomas, D.D. Phospholamban pentamer quaternary conformation determined by in-gel fluorescence anisotropy. Biochemistry 44, 4302–4311 (2005).
Acknowledgements
Research in the authors' laboratories is supported by the US National Institutes of Health, the Howard Hughes Medical Institute, the Deutsche Forschungsgemeinschaft (German Research Foundation) and the European Research Council.
Author information
Authors and Affiliations
Contributions
C.H., G.G., A.Z., S.R.A. and S.T. conducted experiments; C.H. and G.G. drafted the paper; and M.H.E., R.Y.T. and M.J.L. finalized the paper.
Corresponding author
Ethics declarations
Competing interests
The University of California, San Diego owns a patent on FlAsH, ReAsH, and tetracysteine sequences. R.Y.T. shares inventors' royalties from these patents. The University of Washington owns a patent on fluorescent GPCR sensors. C.H. and M.J.L. share inventors' royalties from these patents.
Rights and permissions
About this article
Cite this article
Hoffmann, C., Gaietta, G., Zürn, A. et al. Fluorescent labeling of tetracysteine-tagged proteins in intact cells. Nat Protoc 5, 1666–1677 (2010). https://doi.org/10.1038/nprot.2010.129
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2010.129
This article is cited by
-
GPCR kinase knockout cells reveal the impact of individual GRKs on arrestin binding and GPCR regulation
Nature Communications (2022)
-
β-arrestin1 and 2 exhibit distinct phosphorylation-dependent conformations when coupling to the same GPCR in living cells
Nature Communications (2022)
-
Cartilage oligomeric matrix protein is an endogenous β-arrestin-2-selective allosteric modulator of AT1 receptor counteracting vascular injury
Cell Research (2021)
-
Single-molecule localization to study cytoskeletal structures, membrane complexes, and mechanosensors
Biophysical Reviews (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.