Abstract
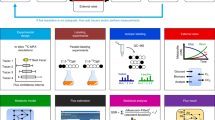
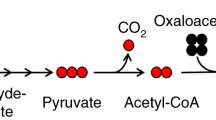
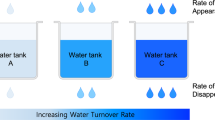
This protocol provides a method for quantitating the intracellular concentrations of endogenous metabolites in cultured cells. The cells are grown in stable isotope-labeled media to near-complete isotopic enrichment and then extracted in organic solvent containing unlabeled internal standards in known concentrations. The ratio of endogenous metabolite to internal standard in the extract is determined using mass spectrometry (MS). The product of this ratio and the unlabeled standard amount equals the amount of endogenous metabolite present in the cells. The cellular concentration of the metabolite can then be calculated on the basis of intracellular volume of the extracted cells. The protocol is exemplified using Escherichia coli and primary human fibroblasts fed uniformly with 13C-labeled carbon sources, with detection of 13C-assimilation by liquid chromatography–tandem MS. It enables absolute quantitation of several dozen metabolites over ∼1 week of work.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Rabinowitz, J.D. & Kimball, E. Acidic acetonitrile for cellular metabolome extraction from Escherichia coli. Anal. Chem. 79, 6167–6173 (2007).
Lu, W., Kwon, Y.K. & Rabinowitz, J.D. Isotope ratio-based profiling of microbial folates. J. Am. Soc. Mass Spectrom. 18, 898–909 (2007).
Yuan, J., Fowler, W.U., Kimball, E., Lu, W. & Rabinowitz, J.D. Kinetic flux profiling of nitrogen assimilation in Escherichia coli. Nat. Chem. Biol. 2, 529–530 (2006).
Lu, W., Kimball, E. & Rabinowitz, J.D. A high-performance liquid chromatography-tandem mass spectrometry method for quantitation of nitrogen-containing intracellular metabolites. J. Am. Soc. Mass Spectrom. 17, 37–50 (2006).
Wu, L. et al. Quantitative analysis of the microbial metabolome by isotope dilution mass spectrometry using uniformly 13C-labeled cell extracts as internal standards. Anal. Biochem. 336, 164–171 (2005).
Mashego, M.R. et al. MIRACLE: mass isotopomer ratio analysis of U-C-13-labeled extracts. A new method for accurate quantification of changes in concentrations of intracellular metabolites. Biotechnol. Bioeng. 85, 620–628 (2004).
Brauer, M.J. et al. Conservation of the metabolomic response to starvation across two divergent microbes. Proc. Natl. Acad. Sci. USA 103, 19302–19307 (2006).
Ikeda, T.P., Shauger, A.E. & Kustu, S. Salmonella typhimurium apparently perceives external nitrogen limitation as internal glutamine limitation. J. Mol. Biol. 259, 589–607 (1996).
Rabinowitz, J.D. Cellular metabolomics of Escherchia coli. Expert Rev. Proteomics 4, 187–198 (2007).
Villas-Boas, S. & Bruheim, P. Cold glycerol-saline: the promising quenching solution for accurate intracellular metabolite analysis of microbial cells. Anal. Biochem. 370, 87–97 (2007).
Wittmann, C., Kromer, J.O., Kiefer, P., Binz, T. & Heinzle, E. Impact of the cold shock phenomenon on quantification of intracellular metabolites in bacteria. Anal. Biochem. 327, 135–139 (2004).
Villas-Boas, S.G., Hojer-Pedersen, J., Akesson, M., Smedsgaard, J. & Nielsen, J. Global metabolite analysis of yeast: evaluation of sample preparation methods. Yeast 22, 1155–1169 (2005).
Maharjan, R.P. & Ferenci, T. Global metabolite analysis: the influence of extraction methodology on metabolome profiles of Escherichia coli. Anal. Biochem. 313, 145–154 (2003).
Kimball, E. & Rabinowitz, J.D. Identifying decomposition products in extracts of cellular metabolites. Anal. Biochem. 358, 273–280 (2006).
Meyer, R.C. et al. The metabolic signature related to high plant growth rate in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 104, 4759–4764 (2007).
Monton, M.R. & Soga, T. Metabolome analysis by capillary electrophoresis-mass spectrometry. J. Chromatogr. A 1168, 237–246 discussion 236 (2007).
Bajad, S.U. et al. Separation and quantitation of water soluble cellular metabolites by hydrophilic interaction chromatography-tandem mass spectrometry. J. Chromatogr. A 1125, 76–88 (2006).
Tolstikov, V.V., Lommen, A., Nakanishi, K., Tanaka, N. & Fiehn, O. Monolithic silica-based capillary reversed-phase liquid chromatography/electrospray mass spectrometry for plant metabolomics. Anal. Chem. 75, 6737–6740 (2003).
Maruska, A., Rocco, A., Kornysova, O. & Fanali, S. Synthesis and evaluation of polymeric continuous bed (monolithic) reversed-phase gradient stationary phases for capillary liquid chromatography and capillary electrochromatography. J. Biochem. Biophys. Methods 70, 47–55 (2007).
Horie, K. et al. Highly efficient monolithic silica capillary columns modified with poly(acrylic acid) for hydrophilic interaction chromatography. J. Chromatogr. A 1164, 198–205 (2007).
Wilson, I.D. et al. HPLC-MS-based methods for the study of metabonomics. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 817, 67–76 (2005).
Nguyen, D.T. et al. High throughput liquid chromatography with sub-2 microm particles at high pressure and high temperature. J. Chromatogr. A 1167, 76–84 (2007).
Guillarme, D., Nguyen, D.T., Rudaz, S. & Veuthey, J.L. Method transfer for fast liquid chromatography in pharmaceutical analysis: application to short columns packed with small particle. Part I: isocratic separation. Eur. J. Pharm. Biopharm. 66, 475–482 (2007).
Yu, K., Little, D., Plumb, R. & Smith, B. High-throughput quantification for a drug mixture in rat plasma—a comparison of ultra performance liquid chromatography/tandem mass spectrometry with high-performance liquid chromatography/tandem mass spectrometry. Rapid Commun. Mass Spectrom. 20, 544–552 (2006).
Cai, S.S., Hanold, K.A. & Syage, J.A. Comparison of atmospheric pressure photoionization and atmospheric pressure chemical ionization for normal-phase LC/MS chiral analysis of pharmaceuticals. Anal. Chem. 79, 2491–2498 (2007).
Hu, Q. et al. The Orbitrap: a new mass spectrometer. J. Mass Spectrom 40, 430–443 (2005).
Wilson, S.D. & Horne, D.W. Evaluation of ascorbic acid in protecting labile folic acid derivatives. Proc. Natl. Acad. Sci. USA 80, 6500–6504 (1983).
Luo, B., Groenke, K., Takors, R., Wandrey, C. & Oldiges, M. Simultaneous determination of multiple intracellular metabolites in glycolysis, pentose phosphate pathway and tricarboxylic acid cycle by liquid chromatography-mass spectrometry. J. Chromatogr. A 1147, 153–164 (2007).
van Winden, W.A. et al. Metabolic-flux analysis of Saccharomyces cerevisiae CEN.PK113-7D based on mass isotopomer measurements of 13C-labeled primary metabolites. FEMS Yeast Res. 5, 559–568 (2005).
Coulier, L. et al. Simultaneous quantitative analysis of metabolites using ion-pair liquid chromatography-electrospray ionization mass spectrometry. Anal. Chem. 78, 6573–6582 (2006).
Brown, A.F. & Dunn, G.A. Microinterferometry of the movement of dry matter in fibroblasts. J. Cell. Sci. 92 (Part 3): 379–389 (1989).
Acknowledgements
This research was supported by the Beckman Foundation, National Science Foundation–Dynamic Data-Driven Application Systems grant CNS-0540181, American Heart Association grant 0635188N, National Science Foundation CAREER award MCB-0643859, National Institutes of Health grant AI078063 and National Institutes of Health grant GM071508 for the Center of Quantitative Biology at Princeton University. We thank Wenyun Lu for contributing LC-MS expertise.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bennett, B., Yuan, J., Kimball, E. et al. Absolute quantitation of intracellular metabolite concentrations by an isotope ratio-based approach. Nat Protoc 3, 1299–1311 (2008). https://doi.org/10.1038/nprot.2008.107
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.107
This article is cited by
-
Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Nature Communications (2023)
-
Metabolite interactions in the bacterial Calvin cycle and implications for flux regulation
Communications Biology (2023)
-
A parallel glycolysis provides a selective advantage through rapid growth acceleration
Nature Chemical Biology (2023)
-
Rapid LC–MS assay for targeted metabolite quantification by serial injection into isocratic gradients
Analytical and Bioanalytical Chemistry (2023)
-
Quick tips for re-using metabolomics data
Nature Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.