Abstract
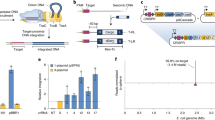
Broad host-range mini-Tn7 vectors facilitate integration of single-copy genes into bacterial chromosomes at a neutral, naturally evolved site. Here we present a protocol for employing the mini-Tn7 system in bacteria with single attTn7 sites, using the example Pseudomonas aeruginosa. The procedure involves, first, cloning of the genes of interest into an appropriate mini-Tn7 vector; second, co-transfer of the recombinant mini-Tn7 vector and a helper plasmid encoding the Tn7 site-specific transposition pathway into P. aeruginosa by either transformation or conjugation, followed by selection of insertion-containing strains; third, PCR verification of mini-Tn7 insertions; and last, optional Flp-mediated excision of the antibiotic-resistance selection marker present on the chromosomally integrated mini-Tn7 element. From start to verification of the insertion events, the procedure takes as little as 4 d and is very efficient, yielding several thousand transformants per microgram of input DNA or conjugation mixture. In contrast to existing chromosome integration systems, which are mostly based on species-specific phage or more-or-less randomly integrating transposons, the mini-Tn7 system is characterized by its ready adaptability to various bacterial hosts, its site specificity and its efficiency. Vectors have been developed for gene complementation, construction of gene fusions, regulated gene expression and reporter gene tagging.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Blatny, J.M., Brautaset, T., Winther-Larsen, H.C., Karunakaran, P. & Valla, S. Improved broad-host-range RK2 vectors for high and low regulated gene expression levels in Gram-negative bacteria. Plasmid 38, 35–51 (1997).
Trieu-Cuot, P., Carlier, C., Poyart-Salmeron, C. & Courvalin, P. An integrative vector exploiting the transposition properties of Tn1545 for insertional mutagenesis and cloning of genes from Gram-positive bacteria. Gene 106, 21–27 (1991).
Stover, C.K. et al. New use of BCG for recombinant vaccines. Nature 351, 456–460 (1991).
Hoang, T.T., Kutchma, A.J., Becher, A. & Schweizer, H.P. Integration proficient plasmids for Pseudomonas aeruginosa: site-specific integration and use for engineering of reporter and expression strains. Plasmid 43, 59–72 (2000).
Kieser, T., Bibb, M.J., Buttner, M.J., Chater, K.F. & Hopwood, D.A. Practical Streptomyces Genetics (The John Innes Foundation, Colney, UK, 2000).
Charpentier, E. et al. Novel cassette-based shuttle vector system for Gram-positive bacteria. Appl. Environ. Microbiol. 70, 6076–6085 (2004).
de Lorenzo, V., Herrero, M., Jakubzik, U. & Timmis, K.N. Mini-Tn5 transposon derivatives for insertion mutagenesis, promoter probing, and chromosomal insertion of cloned DNA in Gram-negative bacteria. J. Bacteriol. 172, 6568–6572 (1990).
Craig, N.L. Transposon Tn7. in Mobile DNA (eds. Berg, D.E. & Howe, M.M.) 211–225 (American Society for Microbiology, Washington DC, 1989).
Craig, N.L. Transposon Tn7. Curr. Top. Microbiol. Immunol. 204, 27–48 (1996).
Choi, K.-H. et al. A Tn7-based broad-range bacterial cloning and expression system. Nat. Methods 2, 443–448 (2005).
Peters, J.E. & Craig, N.L. Tn7: smarter than we thought. Nat. Rev. Mol. Cell Biol. 2, 806–814 (2001).
Bao, Y., Lies, D.P., Fu, H. & Roberts, G.P. An improved Tn7-based system for the single-copy insertion of cloned genes into the chromosomes of Gram-negative bacteria. Gene 109, 167–168 (1991).
Hojberg, O., Schnider, U., Winteler, H.V., Sorensen, J. & Haas, D. Oxygen-sensing reporter strain of Pseudomonas fluorescens for monitoring the distribution of low-oxygen habitats in soil. Appl. Environ. Microbiol. 65, 4085–4093 (1999).
Koch, B., Jensen, L.E. & Nybroe, O. A panel of Tn7-based vectors for insertion of the gfp marker gene or for delivery of cloned DNA into Gram-negative bacteria. J. Mirobiol. Methods 45, 187–195 (2001).
Klausen, M. et al. Biofilm formation by Pseudomonas aeruginosa wild type, flagella and type IV pili mutants. Mol. Microbiol. 48, 1511–1524 (2003).
Lambertsen, L., Sternberg, C. & Molin, S. Mini-Tn7 transposons for site-specific tagging of bacteria with fluorescent proteins. Environ. Microbiol. 6, 726–732 (2004).
Stellwagen, A.E. & Craig, N.L. Avoiding self: two Tn7-encoded proteins mediate target immunity in Tn7 transposition. EMBO J. 16, 6823–6834 (1997).
Choi, K.-H., DeShazer, D. & Schweizer, H.P. mini-Tn7 insertion in bacteria with multiple glmS-linked attTn7 sites: example Burkholderia mallei ATCC 23344. Nat. Protocols 10.1038/nprot2006.25 (2006).
Choi, K.-H. & Schweizer, H.P. mini-Tn7 insertion in bacteria with secondary, non-glmS-linked attTn7 sites: example Proteus mirabilis HI4320. Nat. Protocols 1, 170–178 (2006).
Becher, A. & Schweizer, H.P. Integration-proficient Pseudomonas aeruginosa vectors for isolation of single copy chromosomal lacZ and lux gene fusions. BioTechniques 29, 948–954 (2000).
Schweizer, H.P., Hoang, T.T., Propst, K.L., Ornelas, H.R. & Karkhoff-Schweizer, R.R. Vector design and development of host strains for Pseudomonas. in Genetic Engineering (ed. Setlow, J.K.) 69–81 (Kluwer-Academic/Plenum, New York, 2001).
Wyckoff, T.J. & Wozniak, D.J. Transcriptional analysis of genes involved in Pseudomonas aeruginosa biofilms. Methods Enzymol. 336, 144–151 (2001).
Sauer, K., Camper, A.K., Ehrlich, G.D., Costerton, J.W. & Davies, D.G. Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J. Bacteriol. 184, 1140–1154 (2002).
Choi, K.-H., Kumar, A. & Schweizer, H.P. A 10 min method for preparation of highly electrocompetent Pseudomonas aeruginosa cells: application for DNA fragment transfer between chromosomes and plasmid transformation. J. Microbiol. Methods 64, 391–397 (2006).
Hoang, T.T., Karkhoff-Schweizer, R.R., Kutchma, A.J. & Schweizer, H.P. A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212, 77–86 (1998).
Sambrook, J. & Russell, D.W. Molecular Cloning (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 2001).
Acknowledgements
We thank all present and former members of the Schweizer laboratory, especially R.R. Karkhoff-Schweizer, C. Lopez, J. Gaynor, S. Joshi, K. White and many undergraduate students, for their important contributions to establishing the Tn7 system in various bacteria, especially P. aeruginosa. This work was supported by a Public Health Service grant (AI058141) from the US National Institute of Allergy and Infectious Diseases (NIAID).
Author information
Authors and Affiliations
Contributions
K.H.C. performed all experiments and assisted in writing the manuscript; H.P.S. supervised the research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Choi, KH., Schweizer, H. mini-Tn7 insertion in bacteria with single attTn7 sites: example Pseudomonas aeruginosa. Nat Protoc 1, 153–161 (2006). https://doi.org/10.1038/nprot.2006.24
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.24
This article is cited by
-
Chromosomal barcodes for simultaneous tracking of near-isogenic bacterial strains in plant microbiota
Nature Microbiology (2024)
-
3-Ketoacyl-ACP synthase III FabH1 is essential for branched-chain DSF family signals in Xanthomonas oryzae pv. oryzae
Phytopathology Research (2023)
-
Two-step conversion of polyethylene into recombinant proteins using a microbial platform
Microbial Cell Factories (2023)
-
Disrupting iron homeostasis can potentiate colistin activity and overcome colistin resistance mechanisms in Gram-Negative Bacteria
Communications Biology (2023)
-
The evolution of short- and long-range weapons for bacterial competition
Nature Ecology & Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.