Abstract
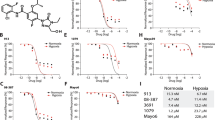
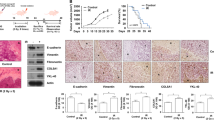
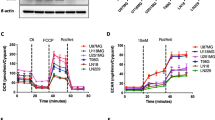
Cross-talk among oncogenic signaling and metabolic pathways may create opportunities for new therapeutic strategies in cancer. Here we show that although acute inhibition of EGFR-driven glucose metabolism induces only minimal cell death, it lowers the apoptotic threshold in a subset of patient-derived glioblastoma (GBM) cells. Mechanistic studies revealed that after attenuated glucose consumption, Bcl-xL blocks cytoplasmic p53 from triggering intrinsic apoptosis. Consequently, targeting of EGFR-driven glucose metabolism in combination with pharmacological stabilization of p53 with the brain-penetrant small molecule idasanutlin resulted in synthetic lethality in orthotopic glioblastoma xenograft models. Notably, neither the degree of EGFR-signaling inhibition nor genetic analysis of EGFR was sufficient to predict sensitivity to this therapeutic combination. However, detection of rapid inhibitory effects on [18F]fluorodeoxyglucose uptake, assessed through noninvasive positron emission tomography, was an effective predictive biomarker of response in vivo. Together, these studies identify a crucial link among oncogene signaling, glucose metabolism, and cytoplasmic p53, which may potentially be exploited for combination therapy in GBM and possibly other malignancies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Brennan, C.W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Vivanco, I. et al. Differential sensitivity of glioma- versus lung cancer-specific EGFR mutations to EGFR kinase inhibitors. Cancer Discov. 2, 458–471 (2012).
Cloughesy, T.F., Cavenee, W.K. & Mischel, P.S. Glioblastoma: from molecular pathology to targeted treatment. Annu. Rev. Pathol. 9, 1–25 (2014).
Lee, E.Q. et al. Phase I/II study of sorafenib in combination with temsirolimus for recurrent glioblastoma or gliosarcoma: North American Brain Tumor Consortium study 05-02. Neuro-oncol. 14, 1511–1518 (2012).
Wen, P.Y. et al. Phase I/II study of erlotinib and temsirolimus for patients with recurrent malignant gliomas: North American Brain Tumor Consortium trial 04-02. Neuro-oncol. 16, 567–578 (2014).
Lee, M.J. et al. Sequential application of anticancer drugs enhances cell death by rewiring apoptotic signaling networks. Cell 149, 780–794 (2012).
Vander Heiden, M.G. et al. Growth factors can influence cell growth and survival through effects on glucose metabolism. Mol. Cell. Biol. 21, 5899–5912 (2001).
Altman, B.J. & Rathmell, J.C. Metabolic stress in autophagy and cell death pathways. Cold Spring Harb. Perspect. Biol. 4, a008763 (2012).
Verhaak, R.G.W. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Babic, I. et al. EGFR mutation-induced alternative splicing of Max contributes to growth of glycolytic tumors in brain cancer. Cell Metab. 17, 1000–1008 (2013).
Lee, J. et al. Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 9, 391–403 (2006).
Nathanson, D.A. et al. Targeted therapy resistance mediated by dynamic regulation of extrachromosomal mutant EGFR DNA. Science 343, 72–76 (2014).
Masui, K. et al. mTOR complex 2 controls glycolytic metabolism in glioblastoma through FoxO acetylation and upregulation of c-Myc. Cell Metab. 18, 726–739 (2013).
Haq, R. et al. Oncogenic BRAF regulates oxidative metabolism via PGC1α and MITF. Cancer Cell 23, 302–315 (2013).
Zhao, Y. et al. Glucose metabolism attenuates p53 and Puma-dependent cell death upon growth factor deprivation. J. Biol. Chem. 283, 36344–36353 (2008).
Deng, J. et al. BH3 profiling identifies three distinct classes of apoptotic blocks to predict response to ABT-737 and conventional chemotherapeutic agents. Cancer Cell 12, 171–185 (2007).
Montero, J. et al. Drug-induced death signaling strategy rapidly predicts cancer response to chemotherapy. Cell 160, 977–989 (2015).
Kruse, J.P. & Gu, W. Modes of p53 regulation. Cell 137, 609–622 (2009).
Maddocks, O.D. & Vousden, K.H. Metabolic regulation by p53. J. Mol. Med. (Berl.) 89, 237–245 (2011).
Jiang, D. et al. Analysis of p53 transactivation domain mutants reveals Acad11 as a metabolic target important for p53 pro-survival function. Cell Rep. 10, 1096–1109 (2015).
Chipuk, J.E. et al. Direct activation of Bax by p53 mediates mitochondrial membrane permeabilization and apoptosis. Science 303, 1010–1014 (2004).
Mihara, M. et al. p53 has a direct apoptogenic role at the mitochondria. Mol. Cell 11, 577–590 (2003).
Liu, J.C. et al. High mitochondrial priming sensitizes hESCs to DNA-damage-induced apoptosis. Cell Stem Cell 13, 483–491 (2013).
Strom, E. et al. Small-molecule inhibitor of p53 binding to mitochondria protects mice from gamma radiation. Nat. Chem. Biol. 2, 474–479 (2006).
Tasdemir, E. et al. Regulation of autophagy by cytoplasmic p53. Nat. Cell Biol. 10, 676–687 (2008).
Green, D.R. & Kroemer, G. Cytoplasmic functions of the tumour suppressor p53. Nature 458, 1127–1130 (2009).
Chipuk, J.E., Bouchier-Hayes, L., Kuwana, T., Newmeyer, D.D. & Green, D.R. PUMA couples the nuclear and cytoplasmic proapoptotic function of p53. Science 309, 1732–1735 (2005).
Lessene, G. et al. Structure-guided design of a selective BCL-X(L) inhibitor. Nat. Chem. Biol. 9, 390–397 (2013).
The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Zhang, Y., Xiong, Y. & Yarbrough, W.G. ARF promotes MDM2 degradation and stabilizes p53: ARF-INK4a locus deletion impairs both the Rb and p53 tumor suppression pathways. Cell 92, 725–734 (1998).
Pomerantz, J. et al. The Ink4a tumor suppressor gene product, p19Arf, interacts with MDM2 and neutralizes MDM2′s inhibition of p53. Cell 92, 713–723 (1998).
Tovar, C. et al. Small-molecule MDM2 antagonists reveal aberrant p53 signaling in cancer: implications for therapy. Proc. Natl. Acad. Sci. USA 103, 1888–1893 (2006).
Lehár, J. et al. Synergistic drug combinations tend to improve therapeutically relevant selectivity. Nat. Biotechnol. 27, 659–666 (2009).
Vaseva, A.V., Marchenko, N.D. & Moll, U.M. The transcription-independent mitochondrial p53 program is a major contributor to nutlin-induced apoptosis in tumor cells. Cell Cycle 8, 1711–1719 (2009).
Chipuk, J.E., Maurer, U., Green, D.R. & Schuler, M. Pharmacologic activation of p53 elicits Bax-dependent apoptosis in the absence of transcription. Cancer Cell 4, 371–381 (2003).
DeBerardinis, R.J., Lum, J.J., Hatzivassiliou, G. & Thompson, C.B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 7, 11–20 (2008).
Ding, Q. et al. Discovery of RG7388, a potent and selective p53-MDM2 inhibitor in clinical development. J. Med. Chem. 56, 5979–5983 (2013).
Tannous, B.A. Gaussia luciferase reporter assay for monitoring biological processes in culture and in vivo. Nat. Protoc. 4, 582–591 (2009).
Qu, L. et al. Endoplasmic reticulum stress induces p53 cytoplasmic localization and prevents p53-dependent apoptosis by a pathway involving glycogen synthase kinase-3β. Genes Dev. 18, 261–277 (2004).
Han, M.-K. et al. SIRT1 regulates apoptosis and Nanog expression in mouse embryonic stem cells by controlling p53 subcellular localization. Cell Stem Cell 2, 241–251 (2008).
Yang, W.H. et al. Modification of p53 with O-linked N-acetylglucosamine regulates p53 activity and stability. Nat. Cell Biol. 8, 1074–1083 (2006).
Leu, J.I.J., Dumont, P., Hafey, M., Murphy, M.E. & George, D.L. Mitochondrial p53 activates Bak and causes disruption of a Bak–Mcl1 complex. Nat. Cell Biol. 6, 443–450 (2004).
Follis, A.V. et al. PUMA binding induces partial unfolding within BCL-xL to disrupt p53 binding and promote apoptosis. Nat. Chem. Biol. 9, 163–168 (2013).
Reardon, D.A., Wen, P.Y. & Mellinghoff, I.K. Targeted molecular therapies against epidermal growth factor receptor: past experiences and challenges. Neuro. Oncol. 16 (Suppl. 8), viii7–vii13 (2014).
Wei, W. et al. Single-cell phosphoproteomics resolves adaptive signaling dynamics and informs targeted combination therapy in glioblastoma. Cancer Cell 29, 563–573 (2016).
Clark, P.M., Ebiana, V.A., Gosa, L., Cloughesy, T.F. & Nathanson, D.A. Harnessing preclinical molecular imaging to inform advances in personalized cancer medicine. J. Nucl. Med. 58, 689–696 (2017).
Spence, A.M. et al. 18F-FDG PET of gliomas at delayed intervals: improved distinction between tumor and normal gray matter. J. Nucl. Med. 45, 1653–1659 (2004).
Nathanson, D. et al. Co-targeting of convergent nucleotide biosynthetic pathways for leukemia eradication. J. Exp. Med. 211, 473–486 (2014).
Takanaga, H. & Frommer, W.B. Facilitative plasma membrane transporters function during ER transit. FASEB J. 24, 2849–2858 (2010).
Cheng, E.H. et al. BCL-2, BCL-X(L) sequester BH3 domain-only molecules preventing BAX- and BAK-mediated mitochondrial apoptosis. Mol. Cell 8, 705–711 (2001).
Dai, H., Marbach, P., Lemaire, M., Hayes, M. & Elmquist, W.F. Distribution of STI-571 to the brain is limited by P-glycoprotein-mediated efflux. J. Pharmacol. Exp. Ther. 304, 1085–1092 (2003).
Magi, A. et al. EXCAVATOR: detecting copy number variants from whole-exome sequencing data. Genome Biol. 14, R120 (2013).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Acknowledgements
We thank H. Herschman, M. Teitell, and T. Graeber for their critical review of the manuscript. This work was funded by National Institutes of Health (NIH)/National Cancer Institute (NCI) R01 CA 213133 (D.A.N.), NIH/NCI UCLA SPORE in Brain Cancer P50 CA 211015 (D.A.N., T.F.C., S.J.B., H.I.K., and W.H.Y.), Nanosystems Biology Cancer Center U54 CA 199090 (D.A.N., T.F.C., and J.T.L.), a Jonsson Comprehensive Cancer Center Foundation/Seed Grant (D.A.N.), the Uncle Kory Foundation (D.A.N. and T.F.C.), the B Hasso Family Foundation (D.A.N. and T.F.C.), the Spiegelman Family Foundation in Memory of Barry Spiegelman (D.A.N. and T.F.C.), the Art of the Brain (T.F.C.), the Ziering Family Foundation (T.F.C. and P.S.M.), the National Brain Tumor Society (T.F.C. and P.S.M.), and the Ben And Catherine Ivy Foundation (T.F.C. and P.S.M.). A.L. and V.W.D. were supported by the Ludwig Institute for Cancer Research and NIH grant R01 CA 205967. H.I.K. was supported by the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation and NIH grant NS 052563. W.X.M. was supported by National Science Foundation (NSF) Graduate Research Fellowship Program (GRFP) DGE-114087. We thank L. Abrey, J. Garcia, G. Nichols, S. Middleton, and L. Chen (all at Roche Pharmaceuticals) for their medical and scientific advice regarding idasanutlin; G. Coppola (UCLA) and Y. Qin (UCLA) for assistance in analyzing exome-sequencing data; K. Faull (UCLA) for help with mass spectrometry; L. Baufeld (UCLA) for technical assistance; J. Heath (Caltech) and M. Phelps (UCLA) for helpful discussions; and W. Fromer (Stanford), S. Korsmeyer (Dana-Farber Cancer Institute), R. Agami (The Netherlands Cancer Institute), and G. Lahav (Harvard) for reagents.
Author information
Authors and Affiliations
Contributions
W.X.M., T.F.C., and D.A.N. conceived the study. W.X.M., L.G., V.W.D., L.T., J.E.T., B.H., W.B.G., N.A.B., M.D.H., J.T.L., W.H.Y., P.N.R., A.L., and D.A.N. designed and/or conducted the experiments and analyzed data. B.H., H.I.K., S.J.B., P.S.M., P.M.C. and A.L. provided reagents, cell lines, and critical input. W.X.M. and D.A.N. wrote the original manuscript. All authors read and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.H. is an employee of Roche. T.F.C. has received consulting fees from Roche.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 (PDF 8016 kb)
Rights and permissions
About this article
Cite this article
Mai, W., Gosa, L., Daniels, V. et al. Cytoplasmic p53 couples oncogene-driven glucose metabolism to apoptosis and is a therapeutic target in glioblastoma. Nat Med 23, 1342–1351 (2017). https://doi.org/10.1038/nm.4418
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4418
This article is cited by
-
A cytosolic mutp53(E285K) variant confers chemoresistance of malignant melanoma
Cell Death & Disease (2023)
-
Pacritinib inhibits glucose consumption in squamous cell lung cancer cells by targeting FLT3
Scientific Reports (2023)
-
Early Reduction of Glucose Consumption Is a Biomarker of Kinase Inhibitor Efficacy Which Can Be Reversed with GLUT1 Overexpression in Lung Cancer Cells
Molecular Imaging and Biology (2023)
-
Leveraging extrachromosomal DNA to fine-tune trials of targeted therapy for glioblastoma: opportunities and challenges
Nature Reviews Clinical Oncology (2022)
-
Repression of p53 function by SIRT5-mediated desuccinylation at Lysine 120 in response to DNA damage
Cell Death & Differentiation (2022)