Abstract
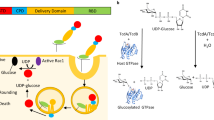
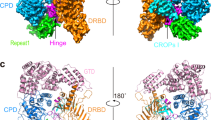
The global prevalence of severe Clostridium difficile infection highlights the profound clinical significance of clostridial glucosylating toxins1,2,3,4. Virulence is dependent on the autoactivation of a toxin cysteine protease5,6,7,8,9, which is promoted by the allosteric cofactor inositol hexakisphosphate (InsP6)10,11,12,13,14,15,16,17. Host mechanisms that protect against such exotoxins are poorly understood. It is increasingly appreciated that the pleiotropic functions attributed to nitric oxide (NO), including host immunity, are in large part mediated by S-nitrosylation of proteins18,19. Here we show that C. difficile toxins are S-nitrosylated by the infected host and that S-nitrosylation attenuates virulence by inhibiting toxin self-cleavage and cell entry. Notably, InsP6- and inositol pyrophosphate (InsP7)-induced conformational changes in the toxin enabled host S-nitrosothiols to transnitrosylate the toxin catalytic cysteine, which forms part of a structurally conserved nitrosylation motif. Moreover, treatment with exogenous InsP6 enhanced the therapeutic actions of oral S-nitrosothiols in mouse models of C. difficile infection. Allostery in bacterial proteins has thus been successfully exploited in the evolutionary development of nitrosothiol-based innate immunity and may provide an avenue to new therapeutic approaches.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chen, X. et al. A mouse model of Clostridium difficile–associated disease. Gastroenterology 135, 1984–1992 (2008).
Savidge, T.C. et al. Clostridium difficile toxin B is an inflammatory enterotoxin in human intestine. Gastroenterology 125, 413–420 (2003).
Lyras, D. et al. Toxin B is essential for virulence of Clostridium difficile. Nature 458, 1176–1179 (2009).
Kuehne, S.A. et al. The role of toxin A and toxin B in Clostridium difficile infection. Nature 467, 711–713 (2010).
Satchell, K.J. MARTX, multifunctional autoprocessing repeats-in-toxin toxins. Infect. Immun. 75, 5079–5084 (2007).
Sheahan, K.L., Cordero, C.L. & Fullner Satchell, K.J. Autoprocessing of the Vibrio cholerae RTX toxin by the cysteine protease domain. EMBO J. 26, 2552–2561 (2007).
Egerer, M. et al. Auto-catalytic cleavage of Clostridium difficile toxins A and B depends on cysteine protease activity. J. Biol. Chem. 282, 25314–25321 (2007).
Pei, J., Lupardus, P.J., Garcia, K.C. & Grishin, N.V. CPDadh: a new peptidase family homologous to the cysteine protease domain in bacterial MARTX toxins. Protein Sci. 18, 856–862 (2009).
Puri, A.W. et al. Rational design of inhibitors and activity-based probes targeting Clostridium difficile virulence factor TcdB. Chem. Biol. 17, 1201–1211 (2010).
Reineke, J. et al. Autocatalytic cleavage of Clostridium difficile toxin B. Nature 446, 415–419 (2007).
Prochazkova, K. & Fullner Satchell, K.J. Structure-function analysis of inositol hexakisphosphate-induced autoprocessing of the Vibrio cholerae multifunctional autoprocessing RTX toxin. J. Biol. Chem. 283, 23656–23664 (2008).
Lupardus, P.J., Shen, A., Bogyo, M. & Garcia, K.C. Small molecule–induced allosteric activation of the Vibrio cholera RTX cysteine protease domain. Science 322, 265–268 (2008).
Egerer, M., Giesemann, T., Herrmann, C. & Aktories, K. Auto-catalytic processing of Clostridium difficile toxin B-binding of inositol hexakisphosphate. J. Biol. Chem. 284, 3389–3395 (2009).
Prochazkova, K. et al. Structural and molecular mechanism for autoprocessing of MARTX toxin of Vibrio cholera at multiple sites. J. Biol. Chem. 284, 26557–26568 (2009).
Pruitt, R.N., Chagot, B., Cover, M., Chazin, W.J. & Lacy, D.B. Structure-function analysis of inositol hexakisphosphate-induced autoprocessing in Clostridium difficile toxin A. J. Biol. Chem. 284, 21934–21940 (2009).
Kreimeyer, I. et al. Autoproteolytic cleavage mediates cytotoxicity of Clostridium difficile toxin A. Naunyn Schmiedebergs Arch. Pharmacol. 383, 253–262 (2011).
Guttenberg, G. et al. Clostridal glucosylating toxins: inositol hexakisphosphate-dependent processing of Closterium sordellii lethal toxin and Clostridium novyi α-toxin. J. Biol. Chem. 286, 14779–14786 (2011).
Benhar, M., Forrester, M.T. & Stamler, J.S. Protein denitrosylation: enzymatic mechanisms and cellular functions. Nat. Rev. Mol. Cell Biol. 10, 721–732 (2009).
Foster, M.W., Hess, D.T. & Stamler, J.S. S-nitrosylation in health and disease-a current perspective. Trends Mol. Med. 15, 391–404 (2009).
Qiu, B., Pothoulakis, C., Castagliuolo, I., Nikulasson, Z. & LaMont, J.T. Nitric oxide inhibits rat intestinal secretion by Clostridium difficile toxin A but not Vibrio cholerae enterotoxin. Gastroenterology 111, 409–418 (1996).
Ng, J. et al. Clostridium difficile toxin–induced inflammation and intestinal injury are mediated by the inflammasome. Gastroenterology 139, 542–552 (2010).
Que, L.G. et al. Protection from experimental asthma by an endogenous bronchodilator. Science 308, 1618–1621 (2005).
Savidge, T.C. et al. Enteric glia regulate intestinal barrier function and inflammation via release of S-nitrosoglutathione. Gastroenterology 132, 1344–1358 (2007).
Hausladen, A. et al. Assessment of nitric oxide signals by triodide chemiluminescence. Proc. Natl. Acad. Sci. USA 104, 2157–2162 (2007).
Wiktorowicz, J. et al. Quantification of cysteinyl S-nitrosylation by fluorescence in unbiased proteomic studies. Biochemistry 50, 5601–5614 (2011).
Jaffrey, S.R. & Snyder, S.H. The biotin-switch method for the detection of S-nitrosylated proteins. Sci. STKE 2001, pl1 (2001).
Gow, A.J. et al. Basal and stimulated protein S-nitrosylation in multiple cell types and tissues. J. Biol. Chem. 277, 9637–9640 (2002).
Popoff, M.R. & Geny, B. Multifaceted role of Rho, Rac, Cdc42 and Ras in intercellular junctions, lessons from toxins. Biochim. Biophys. Acta 88, 797–812 (2009).
Kim, S.F., Huri, D.A. & Snyder, S.H. Inducible nitric oxide synthase binds, S-nitrosylates, and activates cyclooxygenase-2. Science 310, 1966–1970 (2005).
He, X. et al. An ultrasensitive rapid immunocytotoxicity assay for detecting Clostridium difficile toxins. J. Microbiol. Methods 78, 97–100 (2009).
Marino, S.M. & Gladyshev, V.N. Structural analysis of cysteine S-nitrosylation: a modified acid-based motif and the emerging role of trans-nitrosylation. J. Mol. Biol. 395, 844–859 (2010).
Eichinger, A. et al. Crystal structure of gingipain R: an Arg-specific bacterial cysteine proteinase with a caspase-like fold. EMBO J. 18, 5453–5462 (1999).
Chakraborty, A. et al. Inositol pyrophosphates inhibit Akt signaling, thereby regulating insulin sensitivity and weight gain. Cell 143, 897–910 (2010).
Tang, H.Y. & Speicher, D.W. Identification of alternative products and optimization of 2-nitro-5-thiocyanatobenzoic acid cyanylation and cleavage at cysteine residues. Anal. Biochem. 334, 48–61 (2004).
McMahon, T.J. et al. Functional coupling of oxygen binding and vasoactivity in S-nitrosohemoglobin. J. Biol. Chem. 275, 16738–16745 (2000).
Matsumoto, A., Comatas, K.E., Liu, L. & Stamler, J.S. Screening for nitric oxide–dependent protein-protein interactions. Science 301, 657–661 (2003).
Benhar, M., Forrester, M.T., Hess, D.T. & Stamler, J.S. Regulated protein denitrosylation by cytosolic and mitochondrial thioredoxins. Science 320, 1050–1054 (2008).
Cui, Q. & Karplus, M. Allostery and cooperativity revisited. Protein Sci. 17, 1295–1307 (2008).
Saura, M. et al. An antiviral mechanism of nitric oxide: Inhibition of a viral protease. Immunity 10, 21–28 (1999).
Eu, J.P., Sun, J., Xu, L., Stamler, J.S. & Meissner, G. The skeletal muscle calcium release channel: coupled O2 sensor and NO signaling functions. Cell 102, 499–509 (2000).
Acknowledgements
This work was supported by the Eli & Edith Broad Foundation, the John S. Dunn Gulf Coast Consortium for Chemical Genomics Robert A. Welch Collaborative Grant Program, the Howard Hughes Medical Institute and grants from the US National Institutes of Health National Institute of Allergy and Infectious Diseases (R01AI088748, N01AI30050), National Institute of Diabetes and Digestive and Kidney Diseases (R01DK084509, K01DK076549; R21-DK078032-01), National Heart, Lung, and Blood Institute (R01-HL059130, R01-HL091876, R01-HL095463, P01-HL075443-06A, NO1-HV-00245) and 1UL1RR029876-01. We thank D. Powell, S. Weinman, C.S. Schein and G. Prestwich for their critiques.
Author information
Authors and Affiliations
Contributions
T.C.S. designed the study, performed InsP6, GSNO and cytotoxicity assays, BIACORE analysis, and wrote the paper; P.U. performed the toxin S-nitrosylation and InsP6 binding studies; N.O. and W.B. performed the toxin structural modeling and molecular docking simulations; I.P. performed the SNO immunofluorescence; K.A., A.C. and V.A. performed toxin autocleavage, InsP7 phosphorylation and UDP-glucosylation assays; A.G.T. performed animal toxin studies; R.D.E. performed the mass spectrometry; J.E.W. performed the cysteine saturation labeling studies; M.L. provided clinical specimens; R.K. performed the CD spectral analysis; L.S., W.N. and H.F. developed the toxin mutants, performed InsP6 cleavage and stool cytotoxicity assays and animals studies; B.H., A.H. and J.S.S. performed or oversaw the measurements of GSNO and SNO proteins; J.S.S. assisted with the study design and writing of the paper; C.P. prepared holotoxins, performed animal toxin studies and assisted with study design and manuscript editing.
Corresponding author
Ethics declarations
Competing interests
C.P. is a paid consultant with Merck and Optimer Pharmaceuticals and a paid speaker for the Postgraduate Institute for Medicine. J.S.S. has a small financial interest in N30 Pharma, Adamas Pharma, Vindica LLC, SabrePharm and LifeHealth, early-stage companies in development of nitric oxide–related therapeutics.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Methods (PDF 1811 kb)
Rights and permissions
About this article
Cite this article
Savidge, T., Urvil, P., Oezguen, N. et al. Host S-nitrosylation inhibits clostridial small molecule–activated glucosylating toxins. Nat Med 17, 1136–1141 (2011). https://doi.org/10.1038/nm.2405
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2405
This article is cited by
-
S-nitrosothiol homeostasis maintained by ADH5 facilitates STING-dependent host defense against pathogens
Nature Communications (2024)
-
S-nitrosylation-mediated coupling of G-protein alpha-2 with CXCR5 induces Hippo/YAP-dependent diabetes-accelerated atherosclerosis
Nature Communications (2021)
-
S-nitrosylation-mediated activation of a histidine kinase represses the type 3 secretion system and promotes virulence of an enteric pathogen
Nature Communications (2020)
-
Symmetrical (SDMA) and asymmetrical dimethylarginine (ADMA) in sepsis: high plasma levels as combined risk markers for sepsis survival
Critical Care (2018)
-
Markers of nitric oxide are associated with sepsis severity: an observational study
Critical Care (2017)