Abstract
The role of the T-cell receptor (TCR) in commitment of thymocytes to regulatory CD4+Foxp3+ and conventional CD4+Foxp3− T-cell lineages remains controversial. According to the prevailing view, commitment to the former lineage, in contrast to the latter, requires that high affinity TCRs bind rare class II MHC/peptide complexes presented in ‘thymic niches’, which could explain differences between their TCR repertoires. Here we challenge this view and show that the binding of identical TCRs to the same ubiquitously expressed MHC/peptide complex often directs thymocytes to both CD4+ lineages, indicating that the TCR affinity does not play the instructive role, and that restricted presentation of peptides in ‘thymic niches’ is not necessary for selection of CD4+Foxp3+ T cells. However, depending on whether immature thymocytes bound the ligand predominantly with low or high affinity, the repertoires of regulatory and conventional CD4+ T cells were correspondingly similar or mostly different, suggesting that negative rather than positive selection sets them apart.
Similar content being viewed by others
Introduction
The primary repertoire of αβ T-cell receptors (TCR) is shaped by positive and negative selection of immature CD4+CD8+ thymocytes whose fate depends on the strength of the TCR signal, triggered by the recognition of self-peptides bound by major histocompatibility complex (MHC) molecules. The TCR affinity for selecting peptide and the number of TCR engaged (that is, avidity) are two essential factors determining the strength of this signal. Thymocytes that receive signals that are too weak or too strong are negatively selected and die by apoptosis, while those that receive signals of intermediate strength are positively selected, survive and differentiate1,2. Thymocytes positively selected by peptides bound by class II MHC molecules differentiate to two functionally different CD4+ T cells: regulatory (Treg, CD4+Foxp3+) and conventional (CD4+Foxp3−; refs 3, 4). A question that remains unresolved is whether commitment to these T-cell lineages occurs within the same or different range of TCR affinities.
Several studies suggested that commitment to CD4+Foxp3+, in contrast to CD4+Foxp3− lineage, depends on relatively high affinity interactions of TCRs with rare, tissue-specific self-peptides presented in ‘thymic niches’5,6,7,8,9,10, which could be responsible for the observed differences between their TCR repertoires11,12,13. On the other hand, recent studies showed that the TCR repertoires of these two subsets of CD4+ T cells, as well as the range of TCR affinities supporting their positive selection, are considerably overlapping14,15. This observation implied that the commitment to CD4+Foxp3+ and CD4+Foxp3− lineages is not guided by the TCR affinity for positively selecting peptide. However, since in all these studies the selecting peptides were not identified, the possibility that CD4+Foxp3+ and CD4+Foxp3− thymocytes expressing the same TCRs were selected by different self-peptides recognized with different affinities, could not be excluded. Alternatively, the differences between the TCR repertoires of conventional and regulatory T cells could be imposed by negative selection because the susceptibility to apoptosis of CD4+Foxp3+ and CD4+Foxp3− thymocytes was shown to be different16,17,18.
To address these questions we compared repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes selected on class II MHC molecules bound exclusively with one or another variant of self-peptide recognized by predominant fraction of immature thymocytes with low or high affinity, respectively. We show that in both cases the binding of identical TCRs to the same ubiquitously expressed MHC/peptide complex often directs thymocytes to both CD4+ lineages, indicating that the TCR affinity does not play a role in selecting one over the other, and that selection of regulatory CD4+ T cells can occur on peptides with unrestricted thymic expression. However, depending on the bias of the primary TCR repertoire to bind the selecting ligand with low or high affinity, the resulting repertoires of regulatory and conventional CD4+ T cells were correspondingly similar or mostly different, suggesting that negative rather than positive selection is responsible for their separation.
Results
Generation of CD4+ T cells in ‘single-peptide’ TCRmini mice
To study the impact of TCR affinity on the commitment of thymocytes to CD4+Foxp3+ and CD4+Foxp3− lineages and on their TCR repertoires, we first analyzed the generation of CD4+ T cells in ‘single-peptide’ mice, which express a limited TCR repertoire (TCRmini; ref. 19) and class II MHC molecules (Ab) that are exclusively bound with Eα(52–68) peptide (Ep) or its analogue Ep63K (Ep63K). We crossed these mice to Foxp3GFP reporter20 mice, generating TCRminiFoxp3GFP strain (referred to hereon as TCRmini). The expression levels of AbEp and AbEp63K complexes on thymic epithelial cells are very similar and resemble that of native AbEp complexes in congenic B10.5R mice21,22,23. Moreover, the timing of TCR expression on immature thymocytes in TCRmini mice resembles the timing in wild-type (Abwt) mice, and CD4+ thymocytes differentiate into CD4+Foxp3− or CD4+Foxp3+ lineages19 (Fig. 1a first column, and Fig. 2a). The TCR genes used to create TCRmini mice were derived from a T cell originally selected by AbEp complex, which recognized the AbEp63K complex as cognate antigen24. It was therefore expected that in AbEp63KTCRmini mice a larger proportion of CD4+CD8+ thymocytes will bind selecting complex with high affinity than in AbEpTCRmini mice, while in AbEpTCRmini mice a larger proportion of CD4+CD8+ thymocytes will bind selecting complex with low affinity than in AbEp63KTCRmini mice. Importantly, due to the saturation of all Ab molecules with identical peptide, the possible differences in TCR avidity between individual thymocytes expressing identical TCRs were maximally reduced making these mice uniquely suitable to study the role of TCR affinity in commitment to CD4+Foxp3+ and CD4+Foxp3− lineages. Figure 1a shows that, in contrast to the Ab-deficient TCRmini mice, both ‘single-peptide’ TCRmini mice supported the development of a substantial number of CD4+Foxp3+ and CD4+Foxp3− thymocytes, although the proportions and total numbers of thymic and peripheral CD4+ T cells generated in these mice, due to the absence of peptide diversity, were lower than in the AbwtTCRmini mice. Interestingly, the reduction in the AbEpTCRmini mice was much weaker than in the AbEp63KTCRmini mice (Fig. 1a,b), which had higher proportion of CD4+CD8+ thymocytes with elevated expression of activation markers such as Nur77 and phosphorylated AKT kinase (Fig. 1c,d), which are associated with TCR-mediated signalling25,26. This observation suggested that lower number of CD4+ T cells generated in AbEp63KTCRmini mice could be the consequence of higher proportion of thymocytes undergoing apoptosis, reflecting higher proportion of thymocytes recognizing the cognate ligand with high affinity. In agreement with this possibility, the proportion of apoptotic cells among CD4+CD8+CD69highCD5int thymocytes, which undergo selection but not among post-selected CD69highCD5high cells25, was much higher in AbEp63KTCRmini than in AbEpTCRmini mice (Fig. 2b,c). Interestingly, in spite of the lower numbers of CD4+ T cells, the proportion of CD4+Foxp3+ to CD4+Foxp3− cells in both ‘single-peptide’ mice was higher than in AbwtTCRmini mice (Fig. 1a), possibly reflecting higher resistance of CD4+Foxp3+ thymocytes to TCR-mediated deletion16,17,27. The above results showed that thymus-derived regulatory CD4+Foxp3+ T cells (tTregs), can be generated in mice expressing only single, ubiquitously expressed Ab/peptide complex, and confirmed the initial assumption that in the AbEp63KTCRmini mice more immature thymocytes recognized cognate ligand with affinities from the high end of the spectrum than in the AbEpTCRmini mice, as indicated by higher proportion of immature thymocytes undergoing apoptosis.
(a) Representative phenotypes of thymocytes (Thc) and lymph node cells (LNc) from indicated strains of mice. Numbers indicate the proportions of CD4+ and CD4+Foxp3+ cells. (b) Total numbers of CD4+ and CD4+Foxp3+ cells in the thymus (upper panel) and lymph nodes (lower panel) of indicated mice. Bars show the mean (±s.d.) of two independent experiments, in which at least four mice per group were analyzed. (c). Expression of CD69 and Nur77 on CD4+CD8+ (DP) thymocytes. (d). Expression of phosphorylated AKT in DP thymocytes from indicated mice. Bars show the mean (±s.d.), and represent data from three to five mice. Statistical significance of data was calculated using two-sample t-test (OriginLab). GFP, green fluorescent protein.
(a) Representative TCR expression pattern on thymocytes (gated as shown) from AbEpTCRmini and AbEp63KTCRmini mice (thymocytes from AbEp63KTCRmini mice are shown). (b) Staining by the LIVE/DEAD Fixable Far-Red Dead Cell Stain Kit of CD5/CD69 subpopulations of CD4+CD8+ (DP) thymocytes from AbEpTCRmini or AbEp63KTCRmini mice, gated as shown. (c) An average proportion of apoptotic thymocytes in AbEpTCRmini (grey bars) or AbEp63KTCRmini (black bars) mice (±s.d.). Five mice per group were analyzed. One-way analysis of variance was used as statistical method.
The same peptide selects CD4+Foxp3+ and Foxp3− cells with identical TCRs
Because AbEpTCRmini and AbEp63KTCRmini mice differ in the extent of intrathymic apoptosis, comparison of TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes in these mice could be informative as to the significance of negative selection in shaping their repertoires. We also thought that if TCR affinity plays critical role in the commitment, the TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes positively selected on the same, single peptide should be largely non-overlapping. Conversely, if the affinity of TCR-ligand interaction does not dictate the commitment, the repertoires should be largely overlapping. To analyze TCR repertoires we used high-throughput sequencing. Figure 1a and Supplementary Figure 1a show that as in Ab-deficient (Ab−) mice with unlimited TCR repertoire16, some CD4+ T cells were also found in Ab−TCRmini mice. TCRs sequenced from these cells accounted for 20–40% of TCRmini repertoires selected on AbEp or AbEp63K complexes (Fig. 3a,b grey slices). These sequences (their frequencies in different TCRmini strains are shown in Supplementary Fig.1b) were excluded from further analysis to eliminate TCRs that could be selected on ligands not associated with class II MHC molecules. Figure 3a,b and Supplementary Figure 2a,b also show that within 20 most abundant TCRs found on each subset of CD4+ thymocytes, several were abundant on both subsets. Most TCRs abundant on CD4+Foxp3+ thymocytes were found at low abundance on CD4+Foxp3− thymocytes and vice versa, suggesting that qualitative differences between the repertoires may be in fact smaller than that revealed by the present limited analysis. The observed differences in the frequencies of shared TCRs in CD4+Foxp3+ and CD4+Foxp3− subsets may result from different proliferation rate, thymic dwell time and stochastic events before and/or after selection. The expression of identical TCRs on CD4+Foxp3+ and CD4+Foxp3− thymocytes was previously observed in wt mice11,19 but their expression on both subsets in ‘single-peptide’ mice excluded the possibility that these TCRs were selected on different self-peptides recognized with different affinities. To compare the repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes more comprehensively, we examined the TCRs that accounted for no less than 0.01% in at least one sequenced population. Figure 3c shows the alignment of individual CDR3 regions of TCRα chain from CD4+Foxp3− thymocytes of AbEpTCRmini mice according to their frequency, overlaid by CDR3 regions from CD4+Foxp3+ thymocytes. This analysis revealed (in line with the results obtained by single cell PCR, ref. 19) that in AbEpTCRmini mice 50% of examined TCRs—each different TCR was counted as one, irrespective of its frequency—were shared by both subsets of CD4+ thymocytes (Fig. 3c), demonstrating that binding to the same MHC/peptide ligand frequently directed thymocytes with identical TCRs to different CD4+ lineages (see also Fig. 3e and Supplementary Fig. 3). However, the proportion of TCRs shared between CD4+Foxp3+ and CD4+Foxp3− repertoires was lower in AbEp63KTCRmini and in AbwtTCRmini mice (20 and 18%, respectively, Fig. 3d), indicating that in mice, in which negative selection was more extensive, the similarity of TCR repertoires was strongly reduced. This conclusion was supported by the calculated similarity indices28, which placed the repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes from AbEp63KTCRmini mice on the most extreme opposite branches of the illustrating dendrogram (Fig. 3f), while the corresponding CD4+ lineages from AbEpTCRmini mice clustered in close proximity. To determine whether reduced similarity of TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes in AbEp63KTCRmini mice, as compared with AbEpTCRmini mice, is associated with their reduced diversity, and if so, applies to both subsets or to one of them, we used TCR libraries prepared from the same number of CD4+Foxp3+ and CD4+Foxp3− thymocytes. This analysis revealed that in AbEpTCRmini mice, the diversity of TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes was comparable, but in AbEp63KTCRmini mice the diversity of TCRs on CD4+Foxp3− thymocytes was significantly reduced (Fig. 3g). This last finding suggested that the dissimilarity between repertoires of CD4+ thymocytes committed to conventional and regulatory lineages in AbEp63KTCRmini mice is primarily a consequence of enhanced negative selection of CD4+Foxp3− thymocytes.
(a,b) Pie charts depict the 20 most dominant TCRα CDR3 regions from CD4+Foxp3+ and CD4+Foxp3− thymocytes selected on AbEp or AbEp63K complexes. Each dominant CDR3 is represented by a unique colour, and the size of the slice represents its frequency (for sequences see Supplementary Fig. 2a,b). Protruding slices indicate the abundant TCRs on CD4+Foxp3− thymocytes, found on CD4+Foxp3+ thymocytes at low abundance and vice versa. Pink and grey slices depict, respectively, the portions of the repertoires that cover all less abundant CDR3 regions or CDR3 regions also found on CD4+ T cells from Ab−TCRmini mice (numbers indicate unique CDR3 sequences) (see also Supplementary Fig. 1b). (c,d) Charts show the frequencies of individual TCRs found in CD4+Foxp3− and in CD4+Foxp3+ repertoires. TCRs accounting for no less than 0.01% in at least one of these repertoires are shown. The frequencies of TCRs found exclusively in Foxp3− repertoire are depicted by the height of the red lines while the frequencies of TCRs found exclusively in Foxp3+ repertoire are depicted by the height of the green lines. The TCRs found in both repertoires are marked by blue line. The height of the blue line indicates the frequency in the Foxp3+ repertoire, while the frequency in Foxp3− repertoire is indicated by the slope of the red lines histogram (e). 100 most dominant TCRs expressed on CD4+Foxp3− thymocytes in AbEpTCRmini mice. Their frequencies on CD4+Foxp3+ thymocytes (confirmed by TCR sequencing from single sorted thymocytes19) are shown as Supplementary Fig. 3 (f) The hierarchical clusters depict similarity of TCR repertoires based on comparative analysis of the overlap using mutual information index (MII) (based on high-throughput sequencing, see Supplementary Table 1 for the number of TCR CDR3 sequences analyzed). (g) Diversity indices, that is, effective number of species (ENS) and sample coverage (CVG) of TCRα CDR3 repertoires from CD4+Foxp3+ and CD4+Foxp3− thymocytes were calculated as described in Methods. TCR sequences were retrieved from indicated subsets from three mice of each kind, isolated and sequenced separately.
Impact of negative selection on TCR repertoire
To further investigate the role of negative versus positive selection in separation of TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes we also analyzed AbEpTCRmini→AbEp63KTCRα− chain-deficient (TCRα−) and AbEp63KTCRmini→AbEpTCRα− irradiation chimeras29. In the first type of chimera Ep peptide was expressed on bone marrow-derived cells, which primarily mediate negative selection, while Ep63K peptide was expressed on thymic epithelial cells, which primarily mediate positive selection30. In the second type of chimera the expression pattern of selecting peptides was reversed. The results are shown in Fig. 4. Thymii from both types of chimeras had similar proportions of CD4+Foxp3+ and CD4+Foxp3− thymocytes (Fig. 4a), but their TCR repertoires resembled those of the donor of bone marrow cells, confirming the dominant role of negative selection in setting these repertoires apart (compare Fig. 4b–d with Fig. 3a–d, f). Furthermore, in AbEpTCRmini→Ep63KTCRα chimera, positive selection of CD4+Foxp3+ thymocytes was not significantly enhanced, even though negative selection, due to the absence of this complex on bone marrow-derived cells was reduced. Accordingly, similarity analysis showed (Fig. 4d) that repertoires from AbEpTCRmini mice clustered with repertoires from AbEp63KTCRmini mice reconstituted with AbEpTCRmini bone marrow, and were separate from the rest of the repertoires selected in the presence of AbEp63K complexes on bone marrow-derived cells. Analysis of the diversity and richness of TCRs in chimeric mice also showed that introduction of AbEp63K complex to mice expressing exclusively AbEp complex, reduced this repertoire in size and diversity, particularly in case of the CD4+Foxp3− subset (Fig. 4e), confirming the dominant impact of negative selection on allocation of particular TCRs to given subset of CD4+ thymocytes.
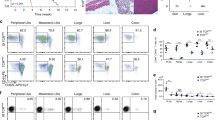
(a) Phenotype and recovery of thymocytes from the indicated bone marrow chimeras 6 weeks after bone marrow transplantation. Numbers indicate proportions of gated cells, and bar graphs show the mean values ±s.d. from two experiments (four mice per group). Statistical significance of data was calculated using two-sample t-test (OriginLab). (b) Repertoire of TCRαCDR3 regions from CD4+Foxp3− and CD4+Foxp3+ thymocytes from indicated chimeras. Colour coding of the particular TCRs is the same as in Fig. 3a,b, and Supplementary Fig. 4 shows CDR3 sequences of dominant TCRs. (c) An overlay of all CDR3 sequences obtained from thymocytes from indicated chimeras. CDR3 regions represented at least at 0.01% are shown. (d) The hierarchical diagrams depict similarity indices (MII) for TCR repertoires from CD4+Foxp3− and CD4+Foxp3+ thymocytes (based on HTS, see Supplementary Table1 for the number of TCR CDR3 sequences analyzed). Note: the clustering of CD4+Foxp3+ and CD4+Foxp3− repertoires in AbEp63KTCRmini→AbEpTCRα− chimeras was not due to their high similarity but to their low TCR diversity (see e), and the similarity between repertoires of CD4+Foxp3− subsets from AbEp63KTCRmini and AbEpKTCRmini→AbEp63KTCRα− chimeras is likely due to the negative selection by AbEp63K complex expressed on thymic medullary epithelial cells. (e) Diversity (ENS) and coverage (CVG) of TCRα CDR3 regions on CD4+Foxp3+ thymocytes selected in the indicated chimeras. CD4+ subsets were separately isolated from three chimeras of each kind and their TCRs were sequenced. GFP, green fluorescent protein.
Additional support for the dominant impact of negative selection on TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− lineages came from the analysis of CDR3 length and amino acid composition of TCRs selected on AbEp versus AbEp63K complexes. As shown in Fig. 5a there was a correlation between the abundance of a 10–11 amino acid-long CDR3 regions characteristic for AbEp63K-specific TCRs and the extent of negative selection. As compared with AbwtTCRmini mice, the frequency of 10–11 amino acid-long CDR3 regions in both CD4+ subsets was strongly reduced in AbEp63KTCRmini mice and less in AbEpTCRmini mice. Moreover, analysis of amino acid composition showed, in agreement with statistical analysis of TCR similarity, that the allocation of the same amino acid to particular position in CDR3 of the same length in both CD4+ subsets was more evident among TCRs selected on AbEp than on AbEp63K complexes (Fig. 5b).
(a) Approximately 5 × 104 of randomly selected CDR3 sequences from each repertoire were used for this comparison. (b) Amino acid distribution and entropy for CDR3 regions of 8–12 amino acid long from CD4+Foxp3− and CD4+Foxp3+ thymocytes derived from AbEp TCRmini (upper panels) or AbEp63KTCRmini (lower panels) mice. Approximately 2 × 104 of randomly selected CDR3 sequences from each repertoire were used for this comparison. This analysis was performed using IgAT software40.
TCR affinity requirements for commitment to CD4+Foxp3+ lineage
Another way to study the role of TCR in lineage commitment of CD4+ thymocytes is to analyze the frequency and distribution of TCRs that recognize selecting ligand as agonist and TCRs that do not recognize this ligand as cognate antigen. For this purpose we established a panel of AbEp63K-specific and non-specific hybridomas from CD4+Foxp3+ cells of Ep mice, and measured the mean fluorescence intensity of equilibrium tetramer binding to their TCRs, which provides a relative assessment of monomeric TCR:Ab/peptide affinity31. As shown in Fig. 6a–c and Supplementary Fig. 5a, antigen responsive (#1–6) in contrast to non-responsive (#7–11) hybridomas (both responded to CD3 stimulation) stained weakly with AbEp63K tetramer and in functional assays recognized AbEp63 complex as an agonist (Fig. 6b–d, Supplementary Fig. 5b). As shown in Fig. 6e, TCRs that recognized the selecting ligand as agonist were rare or not found on CD4+ thymocytes in AbEp63KTCRmini mice, in contrast to TCRs that did not bind this ligand with detectable affinity, which were found on both CD4+ subsets. These results also argued against the view that high affinity, self-reactive TCRs are requisite for commitment to regulatory lineage.
(a) Staining of AbEp63K-specific hybridomas from AbEpTCRmini mice with AbEp63K tetramer. CDR3 sequences of their TCRα chains are shown above each histogram. Staining of BS55 hybridoma served as positive and staining with AbCLIP tetramers as negative control. (b) The relative meanfluorescence index (MFI) of AbEp63K tetramer staining (±s.d. from three independent experiments) of AbEp63K-specific and non-specific hybridomas (also see Supplementary Fig. 5a). (c) IL-2 response and (d) CD69 upregulation in response to indicated stimuli. The mean values ±s.d. from three experiments are shown (also see Supplementary Fig.5b). (e) Frequencies of AbEp63K-specific (black/white triangles) and non-specific (grey/white triangles) TCRs on CD4+ thymocytes in AbEp63KTCRmini mice. Each symbol represents data from individual mice and horizontal lines show the mean value. One-way analysis of variance was used. (f) Body weight of AbwtTCRα- (squares) and AbEpTCRα- (circles) recipients of naive CD4+ T cells from AbEpTCRmini mice. (g) Staining by haematoxilin and eosin of tissue sections showing infiltrates of transferred AbEpTCRmini CD4+ T cells in AbwtTCRα- recipients: lung (L), large intestine (LI) salivary glands (SG) and joints (FJ). (h) Frequencies of identified autoreactive TCRs in the repertoire of healthy AbwtTCRmini mice (approximately one third of them were found). Heat maps represent abundance of Abwt-reactive TCRs on CD4+ T cells, aligned according to their clonal expansion in response to self-peptides in lymphopenic Abwt mice. Asterisks indicate that TCR self-reactivity was confirmed in vitro using T-cell hybridomas. Statistical significance was calculated using two-sample t-test (OriginLab).
To verify our conclusions derived from analysis of ‘single-peptide’ mice we examined whether in Abwt mice, TCRs selected within the higher range of affinities are preferentially expressed on CD4+Foxp3+ thymocytes. In healthy Abwt mice, direct identification of high affinity, self-reactive CD4+ clones is hardly possible due to their intrathymic deletion and self-tolerance. To overcome this difficulty, we first identified the TCRs specific for wt peptides in the repertoire of AbEp63KTCRmini mice, which due to exclusive expression of single peptide are not tolerant to wt peptides. For this purpose, purified naive CD4+ T cells from AbEp63KTCRmini mice were adoptively transferred to lymphopenic AbwtTCRα− or AbEp63KTCRα− hosts. The former, but not the latter recipients started losing weight and developed multiorgan wasting disease (Fig. 6f,g), indicating that the transferred CD4+ T cells were triggered by specific, endogenous self-antigens. Then, the TCRs of in vivo expanded self-reactive CD4+ T cells were sequenced and cross-referenced to the repertoire of CD4+ thymocytes from AbwtTCRmini mice. As shown in Fig. 6h, self-reactive TCRs were randomly distributed on CD4+Foxp3+ and CD4+Foxp3− thymocytes, indicating that TCRs which during positive selection bind self-peptides with high affinity are not preferentially expressed on CD4+Foxp3+ T cells (their CDR3 lengths and sequences are shown in Supplementary Fig. 6). This observation supported the conclusion from the analysis of ‘single-peptide’ mice, that intrathymic commitment into CD4+Foxp3+ and CD4+Foxp3− lineages is not guided by the affinity of TCR for the selecting self-peptide.
Discussion
Our study provides new observations of importance for understanding the intrathymic mechanisms that underlie the generation of conventional and regulatory CD4+ T cells, and mechanisms responsible for differences between their repertoires. First, CD4+Foxp3+ thymocytes expressing diverse TCRs were efficiently generated in ‘single-peptide’ mice lacking peptide-specific niches, indicating that interactions with a restricted set of low abundant self-peptides is not obligatory for their development. Second, both subsets shared a large fraction of identical TCRs in both ‘single-peptide’ strains of mice, demonstrating that CD4+Foxp3+ and CD4+Foxp3− cells can be generated from immature thymocytes as a result of recognition of the same ubiquitously expressed peptide by identical TCRs. This finding implies that TCR affinity for selecting ligand and its restricted expression in ‘thymic niches’ do not control commitment to regulatory T-cell lineage. Third, TCR repertoires of CD4+Foxp3+ and CD4+Foxp3− thymocytes positively selected from CD4+CD8+ thymocytes predominantly recognizing self-peptide with affinities from the higher end of the spectrum were less similar than the repertoires selected from CD4+CD8+ thymocytes predominantly recognizing self-peptide with affinities from the lower end of the spectrum. This observation, together with the results from bone marrow chimeras, indicate that it is negative, not positive selection that is responsible for observed dissimilarity between the TCR repertoires of conventional and regulatory CD4+ T cells, which differ with regard to the sensitivity to apoptosis. The biological significance of this difference could stem from the fact that effectors of autoimmunity are mostly recruited from conventional T cells and therefore more stringent purging of autoreactive CD4+Foxp3− than CD4+Foxp3+ thymocytes would be important for preventing ‘horror autotoxicus’. Conversely higher resistance to apoptosis of CD4+Foxp3+ thymocytes resulting in higher TCR diversity of their positively selected progeny may be advantageous for their regulatory role in the immune system.
The important question remains concerning the mechanism allowing thymocytes with the same TCR sequence to adopt different fates. As discussed in recent reviews32,33, several factors including stochastic variation in expression of molecules, can be involved and be responsible for commitment to CD4+Foxp3+ or to CD4+Foxp3− lineages in normal and TCR transgenic mice; for example, predisposition of individual CD4+CD8+ thymocytes, ‘stochastic overlap’ within intermediate range of affinities haphazardly deciding about their fate, encounter of ligand at different stages of development and/or on different antigen presenting cells, TCR-ligand dwell time and avidity. All these factors could also be involved in deciding about the fate of thymocytes with identical TCRs in single-peptide mice analyzed here, leaving open the question which of them may play the essential role.
Methods
Mice
The AbwtTCRmini, AbEpTCRmini, AbEp63K strains and mice with Foxp3GFP reporter were produced as previously described19,20,32. TCRα-, Ab-, Ii- and Abwt mice were purchased from Jackson Laboratory. The AbwtTCRmini mice were obtained by crossing transgenic mice expressing Vα2 and Jα26 segments in germ line configuration with transgenic mice expressing TCRβ chain of the AbEp63K-specific TCR and TCRα-mice. All mice expressing AbEp and/or AbEp63K complexes lacked expression of invariant chain and endogenous Abβ chain, ref. 34. To ensure that each T cell expresses only a single type of αβTCR, all TCRmini mice were heterozygous for TCRmini locus and lacked endogenous TCRα genes (TCRα−). To generate the AbEp63KTCRminiFoxp3GFP strain, the AbEp63K mice were back-crossed with AbEpTCRminiFoxp3GFP mice for eight generations. Ultimately, four new mouse strains were obtained: AbEpTCRminiFoxp3GFP(AbEpTCRmini), AbEp63KTCRminiFoxp3GFP(AbEp63KTCRmini) and two lymphopenic lines AbEpTCRα- and AbEp63KTCRα-. All mice (females and males) were 6 to 8 weeks old and were housed in the laboratory animal care facility at Georgia Regents University according to the protocols approved by the Institutional Animal Care and Use Committee at our University.
Phenotype analysis and cell sorting
Cell surface staining with monoclonal antibodies was done by standard procedures. The following antibodies were used: anti-CD4 (clone GK1.5, 0.4 μg ml−1), anti-CD4 (clone RM4-5, 0.4 μg ml−1), anti-CD8a (clone 53–6.7, 0.4 μg ml−1), anti-CD25 (clone PC61, 0.4 μg ml−1), anti-CD5 (clone 53–7.3, 0.4 μg ml−1), anti-CD69 (clone H1.2F3, 0.4 μg ml−1), anti-TCR Vα2 (clone B20.1, 0.33 μg ml−1), anti-TCR Vβ14 (clone 14–2, 1.25 μg ml−1), anti TCRαβ (clone H57-597, 0.5 μg ml−1). Intra-cellular staining was performed after treatment with cell fixation/permabilization kit (eBioscience), according to the manufacturer's protocol. Antibodies used were anti-mouse Nur77 (clone 14.14, 2 μg ml−1) and anti-phospho-Akt (clone D9E; antibody detects endogenous levels of Akt1 only when phosphorylated at Ser473 0.5 μg ml−1). Monoclonal antibodies were obtained from: BD Biosciences, BioLegend or eBioscience. After staining, cells were analyzed using FACSCanto cytometer, and data were analyzed using FacsDiva software (BD Biosciences) and Flowjo V10 (TreeStar Inc.). To determine the number of the apoptotic cells within thymocytes subpopulations, after cell surface staining, the LIVE/DEAD Fixable Far-Red Dead Cell Stain Kit (Life Technologies) was used according to the manufacturer's procedure.
For sorting, thymocytes or lymph node cells were stained with anti-CD4, anti-CD8 and anti-CD25 or anti-TCR Vα2. CD4+Foxp3+ and CD4+Foxp3− subsets were separated on the basis of expression of Foxp3GFP reporter. All samples were sorted using MoFlo cell sorter (Beckman Coulter) with purity>98%.
Synthesis of TCR cDNA libraries and Ion Torrent sequencing
Total RNA was isolated from the sorted subsets of thymocytes using RNeasey Mini Kit (Qiagen) according to the manufacturer’s procedure. Synthesis of the first complementary DNA (cDNA) strand was performed with a primer specific for the TCR Cα region (5′-TCGGCACATTGATTTGGGAGTC-3′) using Superscript III cDNA synthesis kit (Invitrogen). Incorporation of Ion Torrent sequencing primers (A-Kay and P1-Kay) to each TCR cDNA together with ‘bar-coding’ of DNA material was performed during the first amplification step using Accuprime Taq Polymerase (Invitrogen). The PCR reaction was carried out with a pair of primers specific to the Vα2 and Cα regions of the TCRα chain. The sequence of the 1st primer was as follows: 5′-CATCCCTGCGTG TCTCCGACTCAG XXXXXXXXXXGACTCTCAGCCTGGAGACT-3′, where the embolden font highlights sequence specific to the Vα2 segment of TCR, italic font marks a bar-code sequence, and the bold font is a sequence of the Ion Torrent forward sequencing primer A-Kay. The sequence of 2nd primer was as follows: 5′-CCTCTCTATGGGCAGTCGGTGATTGGTACACAGCA GGTTCTG-3′, where embolden font highlights nucleotides specific to constant region of TCRα and bold font marks sequence of Ion Torrent reverse sequencing primer P1-Kay. After denaturation (95 °C for 2 min) four cycles of PCR proceeded as follows: three cycles at 94 °C for 15 s/54 °C for 15 s/72 °C for 40 s and one cycle 94 °C for 15 s/58 °C for 15 s/72 °C for 40 s. The obtained cDNA was cleaned using AMPure XP kit (Beckman Coulter) and used for emulsion PCR amplification according to the described protocol35. PCR products were cleaned again with the AMPure XP kit and the molarity of each product containing various CDR3 libraries was calculated using quantitative PCR performed with A-Kay and P1-Kay primers, according to the manufacture’s protocol (Sybergreen PCR master mix, BioRad). The libraries were mixed in equal molarity, subjected to the final amplification in the same conditions as described above, and sequenced at EdgeBio Systems (Gaithersburg, MD, USA). Low quality reads and incomplete and erroneous sequences were eliminated during data processing using filters provided by Ion Torrent Suite software. All sequences were aligned to the constant Vα2 and Cα regions and examined as described36.
Bone marrow chimeras
Sublethally irradiated, AbEpTCRα- or AbEp63KTCRα- recipients received intravenous injection of T-cell-depleted bone marrow cells (2 × 106 per mouse) from AbEp63KTCRmini or AbEpTCRmini donors, respectively. After 6 weeks CD4+Foxp3GFP+ and CD4+FoxpGFP− thymocytes were sorted from each recipient and their TCRα CDR3 regions were sequenced as described above.
Adoptive transfer
For adoptive transfer, 2 × 106 of sorted CD4+CD25−Foxp3GFP− cells from AbEp63KTCRmini mice were injected intravenously to AbwtTCRα- or AbEp63KTCRα- hosts. Mouse weight was monitored daily and the experiment was terminated when the recipient’s weight reached 80% of the starting weight. The sorted CD4+TCRVα2+ cells from recipient’s lymph nodes and spleens were used to generate T-cell hybridomas. The antigen-specific hybridomas were identified and their TCRs were sequenced. For histology, organs were fixed with 10% neutral buffered formalin sections, cut at 8 mm, stained with haematoxylin and eosin and examined under the light microscope.
T-cell hybridomas
To identify Ep63K-specific TCRs, CD4+ T cells from AbEpTCRmini mice were adoptively transferred to AbEp63KTCRα- mice, and after 8 weeks activated CD4Vα2+ T cells were sorted and expanded in vitro for 3 days in the presence of antiCD3 10 μg ml−1 and interleukin-2 (IL-2). Production of the T-cell hybridomas from CD4+Foxp3− and CD4+Foxp3+ cells and an in vitro IL-2 assay were previously described14,37. Reponses of hybridomas to antigen presenting cells presenting Ep63K complexes were determined based on change in the relative expression of CD5 and CD69 using standard flow cytometry procedures, and by detecting the amount of IL-2 produced following activation14. The two-sample t-test and one-way analysis of variance test was used to calculate statistical significance.
Tetramer staining
To stain hybridomas with AbEp63K and AbCLIP tetramers, 5 × 104 cells were incubated with anti-CD4 (clone GK1.5) 0.4 μg ml−1 and anti-TCR (clone H57-597) 0.5 μg ml−1 antibodies, respectively, to crosslink TCRs, then gently washed and incubated additionally for 4 h with AbEp63K 10 μg ml−1 or AbCLIP 10 μg ml−1 (control) tetramers. The optimal concentrations of tetramers used for staining were estimated experimentally. Both tetramers were synthesized by NIH-supported Tetramer Core at Emory University (Atlanta, GA, USA). Binding of AbEp63K and AbCLIP tetramers to the TCR-expressing hybridomas was then analyzed by flow cytometry.
Statistical methods
Statistical analysis was performed using the R statistical software (R 2.15.0). For the comparative analysis of the overlap, a mutual information index (I-index) was used as defined28 together with a hierarchical (agglomerative) clustering procedure with Ward linkage method38. Stability of clustering was assessed via parametric (multinomial) bootstrap—the 95% confidence bounds for similarity matrices were constructed based on the Frobenius norm. The sample coverage (CVG)—that is, the conditional probability of discovering a new receptor—was estimated according to the Good–Turing formula39. The CVG index was calculated using a resampling procedure (multinomial bootstrap) with varying sample sizes (from fifty up to the size of the original sample) to assess the effect of under-sampling on the analysis. Diversity was quantified using the effective number of species and presented in the form of diversity profiles (for definitions see ref. 28). These profiles are plots of the order of the index versus the value of the index, thus enabling comparisons of diversity with more weight put on rare (the order of index lower than one) or abundant (the order of index higher than one) receptors. The 95% confidence bounds were constructed as described above. The significance of differences between individual samples or groups of mice was determined by two-sample t-test and one-way analysis of variance test using Origin 9.1 software, and P-values<0.05 were considered statistically significant.
Additional information
How to cite this article: Wojciech, L. et al. The same self-peptide selects conventional and regulatory CD4+ T cells with identical antigen receptors. Nat. Commun. 5:5061 doi: 10.1038/ncomms6061 (2014).
References
von Boehmer, H., Teh, H. S. & Kisielow, P. The thymus selects the useful, neglects the useless and destroys the harmful. Immunol. Today 10, 57–61 (1989).
Starr, T. K., Jameson, S. C. & Hogquist, K. A. Positive and negative selection of T cells. Annu. Rev. Immunol. 21, 139–176 (2003).
Ohkura, N. & Sakaguchi, S. Regulatory T cells: roles of T-cell receptor for their development and function. Semin. Immunopathol. 32, 95–106 (2010).
Josefowicz, S. Z., Lu, L. F. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Jordan, M. S. et al. Thymic selection of CD4+CD25+ regulatory T cells induced by an agonist self-peptide. Nat. Immunol. 2, 301–306 (2001).
Ribot, J., Romagnoli, P. & van Meerwijk, J. P. Agonist ligands expressed by thymic epithelium enhance positive selection of regulatory T lymphocytes from precursors with a normally diverse TCR repertoire. J. Immunol. 177, 1101–1107 (2006).
Leung, M. W., Shen, S. & Lafaille, J. J. TCR-dependent differentiation of thymic Foxp3+ cells is limited to small clonal sizes. J. Exp. Med. 206, 2121–2130 (2009).
Bautista, J. L. et al. Intraclonal competition limits the fate determination of regulatory T cells in the thymus. Nat. Immunol. 10, 610–617 (2009).
Aschenbrenner, K. et al. Selection of Foxp3(+) regulatory T cells specific for self antigen expressed and presented by Aire(+) medullary thymic epithelial cells. Nat. Immunol. 8, 351–358 (2007).
Lee, H. M., Bautista, J. L. & Hsieh, C. S. Thymic and peripheral differentiation of regulatory T cells. Adv. Immunol. 112, 25–71 (2011).
Hsieh, C. S. et al. Recognition of the peripheral self by naturally arising CD25+ CD4+ T-cell receptors. Immunity 21, 267–277 (2004).
Coutinho, A. et al. Thymic commitment of regulatory T cells is a pathway of TCR-dependent selection that isolates repertoires undergoing positive or negative selection. Curr. Top. Microbiol. Immunol. 293, 43–71 (2005).
Hsieh, C. S. et al. An intersection between the self-reactive regulatory and nonregulatory T-cell receptor repertoires. Nat. Immunol. 7, 401–410 (2006).
Pacholczyk, R. et al. Nonself-antigens are the cognate specificities of foxp3(+) regulatory T cells. Immunity 27, 493–504 (2007).
Wong, J. et al. Adaptation of TCR repertoires to self-peptides in regulatory and nonregulatory CD4+ T Cells. J. Immunol. 178, 7032–7041 (2007).
van Santen, H. M., Benoist, C. & Mathis, D. Number of T reg cells that differentiate does not increase upon encounter of agonist ligand on thymic epithelial cells. J. Exp. Med. 200, 1221–1230 (2004).
Taylor, S. R. et al. Regulatory T cells are resistant to apoptosis via TCR but not P2X7. J. Immunol. 178, 3474–3482 (2007).
Fassett, M. S. et al. Nuclear receptor Nr4a1 modulates both regulatory T-cell (Treg) differentiation and clonal deletion. Proc. Natl Acad. Sci. USA 109, 3891–3896 (2012).
Pacholczyk, R. et al. Origin and T-cell receptor diversity of Foxp3+CD4+CD25+ T cells. Immunity 25, 249–259 (2006).
Kuczma, M. et al. Foxp3-deficient regulatory T cells do not revert into conventional effector CD4+ T cells but constitute a unique cell subset. J. Immunol. 183, 3731–3741 (2009).
Rudensky, A. Y. et al. On the complexity of self. Nature 353, 660–662 (1991).
Ignatowicz, L., Kappler, J. & Marrack, P. The repertoire of T cells shaped by a single MHC/peptide ligand. Cell 84, 521–529 (1996).
Pacholczyk, R., Kraj, P. & Ignatowicz, L. Peptide specificity of thymic selection of CD4+CD25+ T cells. J. Immunol. 168, 613–620 (2002).
Chmielowski, B., Muranski, P. & Ignatowicz, L. In the normal repertoire of CD4+ T cells, a single class II MHC/peptide complex positively selects TCRs with various antigen specificities. J. Immunol. 162, 95–105 (1999).
Stritesky, G. L. et al. Murine thymic selection quantified using a unique method to capture deleted T cells. Proc. Natl Acad. Sci. USA 110, 4679–4684 (2013).
Sauer, S. et al. T-cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc. Natl Acad. Sci. USA 105, 7797–7802 (2008).
Moon, J. J. et al. Quantitative impact of thymic selection on Foxp3+ and Foxp3- subsets of self-peptide/MHC class II-specific CD4+ T cells. Proc. Natl Acad. Sci. USA 108, 14602–14607 (2011).
Rempala, G. A. & Seweryn, M. Methods for diversity and overlap analysis in T-cell receptor populations. J. Math. Biol. 67, 1339–1368 (2013).
Liston, A. et al. Differentiation of regulatory Foxp3+ T cells in the thymic cortex. Proc. Natl Acad. Sci. USA 105, 11903–11908 (2008).
Bensinger, S. J. et al. Major histocompatibility complex class II-positive cortical epithelium mediates the selection of CD4(+)25(+) immunoregulatory T cells. J. Exp. Med. 194, 427–438 (2001).
Lee, H. M. et al. A broad range of self-reactivity drives thymic regulatory T cell selection to limit responses to self. Immunity 37, 475–486 (2012).
Bains, I. et al. Models of self-peptide sampling by developing T cells identify candidate mechanisms of thymic selection. PLoS Comput. Biol. 9, 1–16 e1003102 (2013).
Klein, L. et al. Positive and negative selection of the cell repertoire: what thymocytes see (and don’t see). Nat. Rev. Immunol. 14, 377–391 (2014).
Pacholczyk, R., Kraj, P. & Ignatowicz, L. An incremental increase in the complexity of peptides bound to class II MHC changes the diversity of positively selected alpha/beta TCRs. J. Immunol. 166, 2357–2363 (2001).
Williams, R. et al. Amplification of complex gene libraries by emulsion PCR. Nat. Methods 3, 545–550 (2006).
Cebula, A. et al. Thymus-derived regulatory T cells contribute to tolerance to commensal microbiota. Nature 497, 258–362 (2013).
Singh, N. et al. Generation of T-cell hybridomas from naturally occurring FoxP3+ regulatory T cells. Methods Mol. Biol. 707, 39–44 (2011).
Kaufman, L. & Rousseeuw, P. J. Finding Groups in Data 1–67John Wiley & Sons, Inc. (1990).
Good, I. J. The population frequencies of species and the estimation of population parameters. Biometrika 40, 237–264 (1953).
T.Rogosch, T. et al. Immunoglobulin analysis tool: a novel tool for the analysis of human and mouse heavy and light chain transcripts. Front. Immunol. 3, 176–190 (2012).
Acknowledgements
We thank J. Pihkala and E. Szurek for technical assistance; P. Kraj and M. Kuczma for Foxp3GFP mice; B. Scirka for BS55 hybridoma; A. Cebula, M. Kuczma, E. Szurek, A. Miazek and M. Cebrat for discussions and Mrs Alicja Obodczuk for secretarial help. This work was supported by basic research grants from the National Institutes of Health (NIH; AI 5R01AI079277 to L.I., R01CA152158 to G.R. and NSF DMS1106485 to G.R.).
Author information
Authors and Affiliations
Contributions
L.W. performed most experiments and analyzed the data; A.I. performed TCRα RT-PCR and genotyped mice; M.S. and G.R. developed the statistical methodology and performed statistical analyses; S.S.P and R.A.M. designed the PACE programme for TCR analysis; P.K. and L.I. designed the study, analyzed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-6, Supplementary Table 1 and Supplementary Reference (PDF 1394 kb)
Rights and permissions
About this article
Cite this article
Wojciech, L., Ignatowicz, A., Seweryn, M. et al. The same self-peptide selects conventional and regulatory CD4+ T cells with identical antigen receptors. Nat Commun 5, 5061 (2014). https://doi.org/10.1038/ncomms6061
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms6061
This article is cited by
-
Self and microbiota-derived epitopes induce CD4+ T cell anergy and conversion into CD4+Foxp3+ regulatory cells
Mucosal Immunology (2021)
-
Dormant pathogenic CD4+ T cells are prevalent in the peripheral repertoire of healthy mice
Nature Communications (2019)
-
Non-canonicaly recruited TCRαβCD8αα IELs recognize microbial antigens
Scientific Reports (2018)
-
Increased generation of Foxp3+ regulatory T cells by manipulating antigen presentation in the thymus
Nature Communications (2016)
-
Controversies concerning thymus‐derived regulatory T cells: fundamental issues and a new perspective
Immunology & Cell Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.