Abstract
Bone marrow fibrosis is a critical component of primary myelofibrosis (PMF). However, the origin of the myofibroblasts that drive fibrosis is unknown. Using genetic fate mapping we found that bone marrow leptin receptor (Lepr)-expressing mesenchymal stromal lineage cells expanded extensively and were the fibrogenic cells in PMF. These stromal cells downregulated the expression of key haematopoietic-stem-cell-supporting factors and upregulated genes associated with fibrosis and osteogenesis, indicating fibrogenic conversion. Administration of imatinib or conditional deletion of platelet-derived growth factor receptor a (Pdgfra) from Lepr+ stromal cells suppressed their expansion and ameliorated bone marrow fibrosis. Conversely, activation of the PDGFRA pathway in bone marrow Lepr+ cells led to expansion of these cells and extramedullary haematopoiesis, features of PMF. Our data identify Lepr+ stromal lineage cells as the origin of myofibroblasts in PMF and suggest that targeting PDGFRA signalling could be an effective way to treat bone marrow fibrosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Zhou, B. O., Yue, R., Murphy, M. M., Peyer, J. G. & Morrison, S. J. Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell 15, 154–168 (2014).
Schepers, K., Campbell, T. B. & Passegue, E. Normal and leukemic stem cell niches: insights and therapeutic opportunities. Cell Stem Cell 16, 254–267 (2015).
Abdel-Wahab, O. I. & Levine, R. L. Primary myelofibrosis: update on definition, pathogenesis, and treatment. Annu. Rev. Med. 60, 233–245 (2009).
Tefferi, A. Myeloproliferative neoplasms: a decade of discoveries and treatment advances. Am. J. Hematol. 91, 50–58 (2016).
Araki, M. et al. Activation of the thrombopoietin receptor by mutant calreticulin in CALR-mutant myeloproliferative neoplasms. Blood 127, 1307–1316 (2016).
Chachoua, I. et al. Thrombopoietin receptor activation by myeloproliferative neoplasm associated calreticulin mutants. Blood 127, 1325–1335 (2016).
Marty, C. et al. Calreticulin mutants in mice induce an MPL-dependent thrombocytosis with frequent progression to myelofibrosis. Blood 127, 1317–1324 (2016).
Klampfl, T. et al. Somatic mutations of calreticulin in myeloproliferative neoplasms. New Engl. J. Med. 369, 2379–2390 (2013).
Nangalia, J. et al. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. New Engl. J. Med. 369, 2391–2405 (2013).
Rampal, R. et al. Integrated genomic analysis illustrates the central role of JAK-STAT pathway activation in myeloproliferative neoplasm pathogenesis. Blood 123, e123–e133 (2014).
Jacobson, R. J., Salo, A. & Fialkow, P. J. Agnogenic myeloid metaplasia: a clonal proliferation of hematopoietic stem cells with secondary myelofibrosis. Blood 51, 189–194 (1978).
Lundberg, P. et al. Myeloproliferative neoplasms can be initiated from a single hematopoietic stem cell expressing JAK2-V617F. J. Exp. Med. 211, 2213–2230 (2014).
Papadantonakis, N., Matsuura, S. & Ravid, K. Megakaryocyte pathology and bone marrow fibrosis: the lysyl oxidase connection. Blood 120, 1774–1781 (2012).
Yan, X. Q. et al. Chronic exposure to retroviral vector encoded MGDF (mpl-ligand) induces lineage-specific growth and differentiation of megakaryocytes in mice. Blood 86, 4025–4033 (1995).
Villeval, J. L. et al. High thrombopoietin production by hematopoietic cells induces a fatal myeloproliferative syndrome in mice. Blood 90, 4369–4383 (1997).
Shivdasani, R. A., Fujiwara, Y., McDevitt, M. A. & Orkin, S. H. A lineage-selective knockout establishes the critical role of transcription factor GATA-1 in megakaryocyte growth and platelet development. EMBO J. 16, 3965–3973 (1997).
Vannucchi, A. M. et al. Development of myelofibrosis in mice genetically impaired for GATA-1 expression (GATA-1(low) mice). Blood 100, 1123–1132 (2002).
Jeremy Wen, Q. et al. Targeting megakaryocytic-induced fibrosis in myeloproliferative neoplasms by AURKA inhibition. Nat. Med. 21, 1473–1480 (2015).
Bonner, J. C. Regulation of PDGF and its receptors in fibrotic diseases. Cytokine Growth Factor Rev. 15, 255–273 (2004).
Olson, L. E. & Soriano, P. Increased PDGFRα activation disrupts connective tissue development and drives systemic fibrosis. Dev. Cell 16, 303–313 (2009).
Iwayama, T. et al. PDGFRα signaling drives adipose tissue fibrosis by targeting progenitor cell plasticity. Genes Dev. 29, 1106–1119 (2015).
Gersuk, G. M., Carmel, R. & Pattengale, P. K. Platelet-derived growth factor concentrations in platelet-poor plasma and urine from patients with myeloproliferative disorders. Blood 74, 2330–2334 (1989).
Tefferi, A. Pathogenesis of myelofibrosis with myeloid metaplasia. J. Clin. Oncol. 23, 8520–8530 (2005).
Morikawa, S. et al. Prospective identification, isolation, and systemic transplantation of multipotent mesenchymal stem cells in murine bone marrow. J. Exp. Med. 206, 2483–2496 (2009).
Mendez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010).
Komada, Y. et al. Origins and properties of dental, thymic, and bone marrow mesenchymal cells and their stem cells. PLoS ONE 7, e46436 (2012).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Pinho, S. et al. PDGFRα and CD51 mark human nestin+ sphere-forming mesenchymal stem cells capable of hematopoietic progenitor cell expansion. J. Exp. Med. 210, 1351–1367 (2013).
Schepers, K. et al. Myeloproliferative neoplasia remodels the endosteal bone marrow niche into a self-reinforcing leukemic niche. Cell Stem Cell 13, 285–299 (2013).
Arranz, L. et al. Neuropathy of haematopoietic stem cell niche is essential for myeloproliferative neoplasms. Nature 512, 78–81 (2014).
Reilly, J. T. et al. Guideline for the diagnosis and management of myelofibrosis. Br. J. Haematol. 158, 453–471 (2012).
Yata, Y. et al. DNase I-hypersensitive sites enhance α1(I) collagen gene expression in hepatic stellate cells. Hepatology 37, 267–276 (2003).
Lin, S. L., Kisseleva, T., Brenner, D. A. & Duffield, J. S. Pericytes and perivascular fibroblasts are the primary source of collagen-producing cells in obstructive fibrosis of the kidney. Am. J. Pathol. 173, 1617–1627 (2008).
Mederacke, I. et al. Fate tracing reveals hepatic stellate cells as dominant contributors to liver fibrosis independent of its aetiology. Nat. Commun. 4, 2823 (2013).
Oguro, H., Ding, L. & Morrison, S. J. SLAM family markers resolve functionally distinct subpopulations of hematopoietic stem cells and multipotent progenitors. Cell Stem Cell 13, 102–116 (2013).
Greenbaum, A. et al. CXCL12 in early mesenchymal progenitors is required for haematopoietic stem-cell maintenance. Nature 495, 227–230 (2013).
De Minicis, S. et al. Gene expression profiles during hepatic stellate cell activation in culture and in vivo. Gastroenterology 132, 1937–1946 (2007).
Lataillade, J. J. et al. Does primary myelofibrosis involve a defective stem cell niche? From concept to evidence. Blood 112, 3026–3035 (2008).
Yue, R., Zhou, B. O., Shimada, I. S., Zhao, Z. & Morrison, S. J. Leptin receptor promotes adipogenesis and reduces osteogenesis by regulating mesenchymal stromal cells in adult bone marrow. Cell Stem Cell 18, 782–796 (2016).
Lydon, N. B. & Druker, B. J. Lessons learned from the development of imatinib. Leuk. Res. 28, S29–S38 (2004).
Worthley, D. L. et al. Gremlin 1 identifies a skeletal stem cell with bone, cartilage, and reticular stromal potential. Cell 160, 269–284 (2015).
Kramann, R. et al. Perivascular Gli1+ progenitors are key contributors to injury-induced organ fibrosis. Cell Stem Cell 16, 51–66 (2015).
Inra, C. N. et al. A perisinusoidal niche for extramedullary haematopoiesis in the spleen. Nature 527, 466–471 (2015).
Tefferi, A. et al. Phase 2 trial of imatinib mesylate in myelofibrosis with myeloid metaplasia. Blood 99, 3854–3856 (2002).
Hasselbalch, H. C. et al. Imatinib mesylate in idiopathic and postpolycythemic myelofibrosis. Am. J. Hematol. 74, 238–242 (2003).
DeFalco, J. et al. Virus-assisted mapping of neural inputs to a feeding center in the hypothalamus. Science 291, 2608–2613 (2001).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Tallquist, M. D. & Soriano, P. Cell autonomous requirement for PDGFRα in populations of cranial and cardiac neural crest cells. Development 130, 507–518 (2003).
Sharov, A. A., Dudekula, D. B. & Ko, M. S. A web-based tool for principal component and significance analysis of microarray data. Bioinformatics 21, 2548–2549 (2005).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Acknowledgements
This work was supported by the MPN Research Foundation. L.D. and J.L. were supported by the Rita Allen Foundation and the National Heart, Lung and Blood Institute (1R01HL132074). Flow cytometry was partly supported by the NIH (S10RR027050 and S10OD020056). We thank R. Schwabe at Columbia and D. Brenner at UC San Diego for providing Col-gfp mice. We thank L. Olson at Oklahoma Medical Research Foundation and P. Soriano at Icahn School of Medicine at Mount Sinai for providing PdgfraD842V mice. We thank S. Weyn-Vanhentenryck, C. Zhang and R. Schwabe at Columbia for help in analysing gene expression data. We thank S. Ho and A. Figueroa for help with flow cytometry.
Author information
Authors and Affiliations
Contributions
G.W., M.D., L.M.-M., Y.L. and L.D. carried out all experiments with the help of Q.L. and J.L. M.D., L.M.-M., G.W. and L.D. designed the experiments, interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 2 HSCs progressively mobilize to the spleen as PMF advances. (Related to Fig. 2).
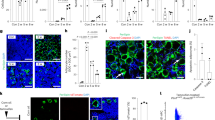
(a) Bone marrow multipotent progenitors (n = 17 mice for control, n = 5 mice for intermediate TOE, n = 10 mice for advanced TOE) and LSK hematopoietic progenitors (n = 12 mice for control, n = 4 mice for intermediate TOE, n = 6 mice for advanced TOE) increased in intermediate then decreased in advanced stage of PMF. (b) Representative flow cytometry plots for quantifying BrdU incorporation in HSCs after a 5-day BrdU pulse. (c) Spleen HSC frequency gradually increased from intermediate to advanced PMF mice (n = 22 mice for control, n = 15 mice for intermediate TOE, n = 10 mice for advanced TOE). (d) Spleen cellularity gradually increased from intermediate to advanced PMF mice (n = 15 mice for control, n = 6 mice for intermediate TOE, n = 7 mice for advanced TOE). (e,f) Spleen HSC number (n = 13 mice for control, n = 6 mice for intermediate TOE, n = 7 mice for advanced TOE), MPP number (n = 9 mice for control, n = 4 mice for intermediate TOE, n = 7 mice for advanced TOE) and HPC (LSK) number (n = 5 mice for control, n = 3 mice for intermediate TOE, n = 3 mice for advanced TOE) gradually increased from intermediate to advanced PMF mice. (g) An increased total HSC number (bone marrow plus spleen) in intermediate PMF mice was followed by a reduction in advanced PMF mice (n = 12 mice for control, n = 6 mice for intermediate TOE, n = 7 mice for advanced TOE). Data represent mean ± s.e.m. Two-tailed student’s t-tests were used to assess statistical significance: ∗P < 0.05, ∗∗∗P < 0.001.
Supplementary Figure 3 Bone marrow mesenchymal stromal lineage cells expand and assume a fibrotic cell fate. (Related to Fig. 3).
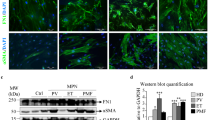
(a) Bone marrow CD45/Ter119-CD140a+ mesenchymal stromal cell frequency increased as PMF developed from intermediate to advanced stages (n = 17 mice for control, n = 12 mice for intermediate TOE, n = 8 mice for advanced TOE). (b) Bone marrow mesenchymal stromal lineage cells expanded 6.5 fold as quantified by counting TdTomato + cells on bone marrow sections from vector control and TOE mice (n = 4 images for control and TOE each, data represent mean ± s.e.m. Two-tailed student’s t-tests were used to assess statistical significance: ∗∗∗P < 0.001). (c,d) Lepr-cre-expressing mesenchymal stromal cells expanded extensively and displayed elongated fibroblast-like stromal cell morphology. Arrow heads point to elongated fibroblast-like cells. (e) In Lepr-cre; loxptdTomato; Col-gfp control or TOE mice, all tdTomato+ cells are GFP+. (f) In Lepr-cre; loxptdTomato; Col-gfp control or TOE mice, nearly all GFP+ cells are tdTomato+. Images are representative of at least 3 biological replicates.
Supplementary Figure 4 Bone marrow mesenchymal stromal cells down-regulate CXCL12 and SCF, and display a fibrotic cell morphology. (Related to Fig. 4).
(a,b) Representative images showing normal Cxcl12-DsRed expression in Cxcl12DsRed/+ mice transplanted with control virus-infected bone marrow cells. (c,d) Images showing Cxcl12-DsRed expression in TOE mice. Arrow heads point to elongated fibroblast-like stromal cells. (e) qPCR analysis showing downregulation of Scf and Cxcl12 in bone marrow mesenchymal stromal cells from intermediate stage to advanced stage PMF mice (n = 4 mice for control, n = 4 mice for intermediate PMF and n = 4 mice for advanced PMF). Data represent mean ± s.d. Two-tailed student’s t-tests were used to assess statistical significance. (f) Representative flow cytometry plots showing CD45/Ter119−Scf-GFP+ stromal cells. Images are representative of at least 3 biological replicates.
Supplementary Figure 5 Gene expression profiling analysis of mesenchymal stromal cells from PMF mice. (Related to Fig. 5).
(a) Representative flow cytometric plots showing the gates to sort CD45/Ter119/CD31−Cxcl12-DsRed+ stromal cells from PMF and control bone marrow. A total of three freshly double-sorted aliquots of cells (∼ 5,000) from PMF (from 5 mice) and control (from 3 mice) Cxcl12DsRed/+ mice were used for gene expression analysis. (b) Normalized expression levels of mesenchymal cell markers and HSC niche factors by CD45/Ter119/CD31−Cxcl12-DsRed+ stromal cells from PMF and control bone marrow. Values represent mean ± s.d. from three biological replicates. (c) List of fibrosis genes used to performed GSEA in Fig. 5c. (d) List of osteogenic genes used to performed GSEA in Fig. 5d.
Supplementary Figure 6 Lepr-cre; Pdgfrafl/− mice have normal HSC function and Lepr-cre; Pdgfrafl/fl TOE mice fail to develop bone marrow fibrosis (Related to Fig. 6).
(a) Flow cytometry plots showing efficient deletion of PDGFRa. (b) A competitive reconstitution assay for Lepr-cre; Pdgfrafl/− and control mice. 5 × 105 donor bone marrow cells from Lepr-cre; Pdgfrafl/− adult mice or control Lepr-cre; Pdgfrafl+ mice were competitively transplanted with 5 × 105 recipient bone marrow cells into irradiated recipient mice. The percentages of donor-derived Mac-1+ myeloid, CD3+ T, and B220+ B cells in the blood were analyzed for 16 weeks after transplantation (n = 5 recipient mice for each genotype). Data represent mean ± s.d. Two-tailed student’s t-tests were used to assess statistical significance. (c) Lepr-cre; Pdgfrafl/fl TOE mice displayed enlarged spleens and imatinib-treated TOE mice showed normal sized spleens. (d) Representative reticulin staining on spleen sections from Lepr-cre; Pdgfrafl/fl TOE mice revealed excessive deposition of reticulin fibers, similar to control TOE mice. (e) Confocal images showing similar levels of megakaryocyte hyperplasia in the bone marrow from Lepr-cre; Pdgfrafl/fl and control TOE mice. CD41 is a marker for megakaryocytes (red). Nuclei were stained with DAPI (blue). (f) Bone marrow sections from Lepr-cre; Pdgfrafl/fl and control TOE mice were subjected to reticulin staining. While control TOE mice robustly developed bone marrow fibrosis, none of the Lepr-cre; Pdgfrafl/fl TOE mice had bone marrow fibrosis. (g) Representative flow cytometry plots showing effective suppression of bone marrow stromal cell expansion in Lepr-cre; Pdgfrafl/fl TOE and imatinib-treated TOE mice. (h) Spleen sections from control and TOE + imatinib were subjected to reticulin staining. Images are representative of at least 3 biological replicates.
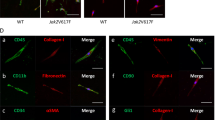
Supplementary Figure 7 Lepr-cre; PdgfraD842V/+ mice have bone marrow stromal fibrotic conversion and HSC mobilization (Related to Fig. 7).
(a) Flow cytometry analysis revealed normal bone marrow stromal cell frequency from Lepr-cre; PdgfraD842V/+ mice (n = 3 mice for control, n = 4 mice for D842V KI). (b) Representative reticulin staining on bone marrow sections from Lepr-cre; PdgfraD842V/+ and control mice. (c) Normal bone marrow cellularity (n = 4 mice for control, n = 5 mice for D842V KI) and HPC frequency of Lepr-cre; PdgfraD842V/+ mice (n = 5 mice for control, n = 6 mice for D842V KI). (d) HSC frequency in livers from Lepr-cre; PdgfraD842V/+ mice (n = 5 mice for control, n = 6 mice for D842V KI). Data represent mean ± s.d. Two-tailed student’s t-tests were used to assess statistical significance. Images are representative of at least 3 biological replicates.
Supplementary information
Supplementary information
Supplementary information (PDF 9451 kb)
Rights and permissions
About this article
Cite this article
Decker, M., Martinez-Morentin, L., Wang, G. et al. Leptin-receptor-expressing bone marrow stromal cells are myofibroblasts in primary myelofibrosis. Nat Cell Biol 19, 677–688 (2017). https://doi.org/10.1038/ncb3530
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3530
This article is cited by
-
Extramedullary hematopoiesis in cancer
Experimental & Molecular Medicine (2024)
-
Cellular niches for hematopoietic stem cells in bone marrow under normal and malignant conditions
Inflammation and Regeneration (2023)
-
Mdm2/p53 levels in bone marrow mesenchymal stromal cells are essential for maintaining the hematopoietic niche in response to DNA damage
Cell Death & Disease (2023)
-
The role of vesicle trafficking genes in osteoblast differentiation and function
Scientific Reports (2023)
-
Contributions of bone marrow monocytes/macrophages in myeloproliferative neoplasms with JAK2V617F mutation
Annals of Hematology (2023)