Abstract
Cleavage of cohesins and cyclin-dependent kinase (CDK) inhibition are thought to be sufficient for triggering chromosome segregation. Here we identify an essential requirement for anaphase chromosome movement. We show that, at anaphase onset, the phosphatase Cdc14 and the polo-like kinase Cdc5 are redundantly required to drive spindle elongation. This role of Cdc14 is mediated by the FEAR network, a group of proteins that activates Cdc14 at anaphase onset, and we suggest that Cdc5 facilitates both Cdc14 activation and CDK inhibition. We further identify the kinesin-5 motor protein Cin8 as a key target of Cdc14. Indeed, Cin8 mutants lacking critical CDK phosphorylation sites suppress the requirement for Cdc14 and Cdc5 in anaphase spindle elongation. Our results indicate that cohesin dissolution and CDK inhibition per se are not sufficient to drive sister chromatid segregation but that the motor protein Cin8 must be activated to elongate the spindle.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Nasmyth, K. Segregating sister genomes: the molecular biology of chromosome separation. Science 297, 559–565 (2002).
Uhlmann, F., Lottspeich, F. & Nasmyth, K. Sister-chromatid separation at anaphase onset is promoted by cleavage of the cohesin subunit Scc1. Nature 400, 37–42 (1999).
Uhlmann, F., Wernic, D., Poupart, M. A., Koonin, E. V. & Nasmyth, K. Cleavage of cohesin by the CD clan protease separin triggers anaphase in yeast. Cell 103, 375–386 (2000).
Thornton, B. R. et al. An architectural map of the anaphase-promoting complex. Genes Dev. 20, 449–460 (2006).
Cohen-Fix, O. & Koshland, D. The anaphase inhibitor of Saccharomyces cerevisiae Pds1p is a target of the DNA damage checkpoint pathway. Proc. Natl Acad. Sci. USA 94, 14361–14366 (1997).
Searle, J. S., Schollaert, K. L., Wilkins, B. J. & Sanchez, Y. The DNA damage checkpoint and PKA pathways converge on APC substrates and Cdc20 to regulate mitotic progression. Nat. Cell Biol. 6, 138–145 (2004).
Musacchio, A. & Salmon, E. D. The spindle-assembly checkpoint in space and time. Nat. Rev. Mol. Cell Biol. 8, 379–393 (2007).
Sullivan, M. & Morgan, D. O. Finishing mitosis, one step at a time. Nat. Rev. Mol. Cell Biol. 8, 894–903 (2007).
Stegmeier, F. & Amon, A. Closing mitosis: the functions of the Cdc14 phosphatase and its regulation. Annu. Rev. Genet. 38, 203–232 (2004).
Bouchoux, C. & Uhlmann, F. A quantitative model for ordered cdk substrate dephosphorylation during mitotic exit. Cell 147, 803–814 (2011).
Visintin, C. et al. APC/C-Cdh1-mediated degradation of the Polo kinase Cdc5 promotes the return of Cdc14 into the nucleolus. Genes Dev. 22, 79–90 (2008).
Visintin, R., Stegmeier, F. & Amon, A. The role of the polo kinase Cdc5 in controlling Cdc14 localization. Mol. Biol. Cell 14, 4486–4498 (2003).
Llamazares, S. et al. Polo encodes a protein kinase homolog required for mitosis in Drosophila. Genes Dev. 5, 2153–2165 (1991).
D’Amours, D. & Amon, A. At the interface between signaling and executing anaphase–Cdc14 and the FEAR network. Genes Dev. 18, 2581–2595 (2004).
Rock, J. M. & Amon, A. The FEAR network. Curr. Biol. 19, R1063–R1068 (2009).
Manzoni, R. et al. Oscillations in Cdc14 release and sequestration reveal a circuit underlying mitotic exit. J. Cell Biol. 190, 209–222 (2010).
Alexandru, G., Uhlmann, F., Mechtler, K., Poupart, M. A. & Nasmyth, K. Phosphorylation of the cohesin subunit Scc1 by Polo/Cdc5 kinase regulates sister chromatid separation in yeast. Cell 105, 459–472 (2001).
Park, C. J. et al. Requirement for the budding yeast polo kinase Cdc5 in proper microtubule growth and dynamics. Eukaryot. Cell 7, 444–453 (2008).
Lee, K. S., Park, J. E., Asano, S. & Park, C. J. Yeast polo-like kinases: functionally conserved multitask mitotic regulators. Oncogene 24, 217–229 (2005).
Liang, F., Jin, F., Liu, H. & Wang, Y. The molecular function of the yeast polo-like kinase Cdc5 in Cdc14 release during early anaphase. Mol. Biol. Cell 20, 3671–3679 (2009).
Snead, J. L. et al. A coupled chemical-genetic and bioinformatic approach to Polo-like kinase pathway exploration. Chem. Biol. 14, 1261–1272 (2007).
Hotz, M., Lengefeld, J. & Barral, Y. The MEN mediates the effects of the spindle assembly checkpoint on Kar9-dependent spindle pole body inheritance in budding yeast. Cell Cycle 11, 3109–3116 (2012).
Tinker-Kulberg, R. L. & Morgan, D. O. Pds1 and Esp1 control both anaphase and mitotic exit in normal cells and after DNA damage. Genes Dev. 13, 1936–1949 (1999).
Wang, H., Liu, D., Wang, Y., Qin, J. & Elledge, S. J. Pds1 phosphorylation in response to DNA damage is essential for its DNA damage checkpoint function. Genes Dev. 15, 1361–1372 (2001).
Agarwal, R., Tang, Z., Yu, H. & Cohen-Fix, O. Two distinct pathways for inhibiting pds1 ubiquitination in response to DNA damage. J. Biol. Chem. 278, 45027–45033 (2003).
Sullivan, M., Hornig, N. C., Porstmann, T. & Uhlmann, F. Studies on substrate recognition by the budding yeast separase. J. Biol. Chem. 279, 1191–1196 (2004).
Dougherty, W. G., Cary, S. M. & Parks, T. D. Molecular genetic analysis of a plant virus polyprotein cleavage site: a model. Virology 171, 356–364 (1989).
Parks, T. D., Howard, E. D., Wolpert, T. J., Arp, D. J. & Dougherty, W. G. Expression and purification of a recombinant tobacco etch virus NIa proteinase: biochemical analyses of the full-length and a naturally occurring truncated proteinase form. Virology 210, 194–201 (1995).
Toone, W. M., Aerne, B. L., Morgan, B. A. & Johnston, L. H. Getting started: regulating the initiation of DNA replication in yeast. Annu. Rev. Microbiol. 51, 125–149 (1997).
Piatti, S., Lengauer, C. & Nasmyth, K. Cdc6 is an unstable protein whose de novo synthesis in G1 is important for the onset of S phase and for preventing a ‘reductional’ anaphase in the budding yeast Saccharomyces cerevisiae. EMBO J. 14, 3788–3799 (1995).
Goh, P. Y. & Kilmartin, J. V. NDC10: a gene involved in chromosome segregation in Saccharomyces cerevisiae. J. Cell Biol. 121, 503–512 (1993).
Fraschini, R., Beretta, A., Lucchini, G. & Piatti, S. Role of the kinetochore protein Ndc10 in mitotic checkpoint activation in Saccharomyces cerevisiae. Mol. Genet. Genomics 266, 115–125 (2001).
Winey, M. & Bloom, K. Mitotic spindle form and function. Genetics 190, 1197–1224 (2012).
Mallavarapu, A., Sawin, K. & Mitchison, T. A switch in microtubule dynamics at the onset of anaphase B in the mitotic spindle of Schizosaccharomyces pombe. Curr. Biol. 9, 1423–1426 (1999).
Higuchi, T. & Uhlmann, F. Stabilization of microtubule dynamics at anaphase onset promotes chromosome segregation. Nature 433, 171–176 (2005).
Liang, F., Richmond, D. & Wang, Y. Coordination of chromatid separation and spindle elongation by antagonistic activities of mitotic and S-phase CDKs. PLoS Genet. 9, e1003319 (2013).
Rahal, R. & Amon, A. Mitotic CDKs control the metaphase-anaphase transition and trigger spindle elongation. Genes Dev. 22, 1534–1548 (2008).
Charles, J. F. et al. The Polo-related kinase Cdc5 activates and is destroyed by the mitotic cyclin destruction machinery in S. cerevisiae. Curr. Biol. 8, 497–507 (1998).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Woodbury, E. L. & Morgan, D. O. Cdk and APC activities limit the spindle-stabilizing function of Fin1 to anaphase. Nat. Cell Biol. 9, 106–112 (2007).
Woodbury, E. L. & Morgan, D. O. The role of self-association in Fin1 function on the mitotic spindle. J. Biol. Chem. 282, 32138–32143 (2007).
Pereira, G. & Schiebel, E. Separase regulates INCENP-Aurora B anaphase spindle function through Cdc14. Science 302, 2120–2124 (2003).
Khmelinskii, A., Lawrence, C., Roostalu, J. & Schiebel, E. Cdc14-regulated midzone assembly controls anaphase B. J. Cell Biol. 177, 981–993 (2007).
Khmelinskii, A. & Schiebel, E. Assembling the spindle midzone in the right place at the right time. Cell Cycle 7, 283–286 (2008).
Khmelinskii, A., Roostalu, J., Roque, H., Antony, C. & Schiebel, E. Phosphorylation-dependent protein interactions at the spindle midzone mediate cell cycle regulation of spindle elongation. Dev. Cell 17, 244–256 (2009).
Rozelle, D. K., Hansen, S. D. & Kaplan, K. B. Chromosome passenger complexes control anaphase duration and spindle elongation via a kinesin-5 brake. J. Cell Biol. 193, 285–294 (2011).
Avunie-Masala, R. et al. Phospho-regulation of kinesin-5 during anaphase spindle elongation. J. Cell Sci. 124, 873–878 (2011).
Hoyt, M. A., He, L., Loo, K. K. & Saunders, W. S. Two Saccharomyces cerevisiae kinesin-related gene products required for mitotic spindle assembly. J. Cell Biol. 118, 109–120 (1992).
Taxis, C., Stier, G., Spadaccini, R. & Knop, M. Efficient protein depletion by genetically controlled deprotection of a dormant N-degron. Mol. Syst. Biol. 5, 267–273 (2009).
Brennan, I. M., Peters, U., Kapoor, T. M. & Straight, A. F. Polo-like kinase controls vertebrate spindle elongation and cytokinesis. PLoS ONE 2, e409 (2007).
Santamaria, A. et al. The Plk1-dependent phosphoproteome of the early mitotic spindle. Mol. Cell. Proteomics 10, M110 004457 (2011).
Donaldson, M. M., Tavares, A. A., Ohkura, H., Deak, P. & Glover, D. M. Metaphase arrest with centromere separation in polo mutants of Drosophila. J. Cell Biol. 153, 663–676 (2001).
Yeong, F. M., Lim, H. H., Padmashree, C. G. & Surana, U. Exit from mitosis in budding yeast: biphasic inactivation of the Cdc28-Clb2 mitotic kinase and the role of Cdc20. Mol. Cell 5, 501–511 (2000).
Stern, B. M. & Murray, A. W. Lack of tension at kinetochores activates the spindle checkpoint in budding yeast. Curr. Biol. 11, 1462–1467 (2001).
Amon, A. Synchronization procedures. Methods Enzymol. 351, 457–467 (2002); J. Cell Biol. 201, 843–862 (2013)
Lianga, N. et al. A Wee1 checkpoint inhibits anaphase onset. J. Cell Biol. 201, 843–862 (2013).
Amon, A., Irniger, S. & Nasmyth, K. Closing the cell cycle circle in yeast: G2 cyclin proteolysis initiated at mitosis persists until the activation of G1 cyclins in the next cycle. Cell 77, 1037–1050 (1994).
Visintin, R., Hwang, E. S. & Amon, A. Cfi1 prevents premature exit from mitosis by anchoring Cdc14 phosphatase in the nucleolus. Nature 398, 818–823 (1999).
Acknowledgements
We thank F. Uhlmann (LRI, UK), L. Gheber (Ben Gurion University, Israel), U. Surana (IMCB, Singapore), A. Amon (MIT, USA), S. Gasser (FMI, Switzerland), A. Rudner (Ottawa Institute of Systems Biology, Canada), E. Schiebel (ZMBH, Germany) and A. Marston (Wellcome Trust Centre for Cell Biology, UK) for strains and reagents; L. Fornasari for helping with statistical analyses; S. Piatti for critical discussion and A. Amon, P. De Wulf, M. Foiani, A. Rudner and members of our laboratory for critical reading of the manuscript. Work in the R.V. laboratory was supported in part by an International Early Career Scientist grant from the Howard Hughes Medical Institute, by a grant from the Associazione Italiana Ricerca sul Cancro (AIRC-IG-12878) to R.V. and by an ‘Armanda e Enrico Mirto’ FIRC Fellowship to F.T.
Author information
Authors and Affiliations
Contributions
M.R. and C.V. performed all the experiments. F.T. prepared the strains and made the initial observation for the experiment shown in Fig. 5c and Supplementary Fig. 4c. R.V. conceived the experiments with the person(s) performing them. M.R. and R.V. wrote the paper. All authors have approved of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 cdc14 cdc5 loss-of-function mutants arrest with undivided nuclei and short bipolar spindles, related to Fig. 1.
(a) Growth of serially diluted wild type, cdc14-3, cdc5-1, cdc5-2, cdc14-3 cdc5-1, cdc14-3 cdc5-2, cdc5-as1, cdc14-1 cdc5-as1, cdc14-3 cdc5-as1 and cdc14-1 cdc5-1 strains are shown. Serial dilutions (1:5) of yeast cell suspensions starting from OD600 = 1 were spotted onto YEPD plate and incubated at 23 °C, 25 °C, 28 °C and 30 °C for 48 h. (b) Wild type, cdc5-as1, cdc14-1 and cdc14-1 cdc5-as1 cells were arrested in G1 byα-factor in YEPD at 23 °C. When more than 90% of cells was unbudded, cells were released in fresh YEPD media supplemented with the CMK inhibitor23 and incubated at 37 °C to inactivate the cdc5-as1 and cdc14-1 alleles, respectively. At the indicated time-points, cells were collected to determine the DNA content by FACS analysis. (c) cdc14-1 cdc5-as1 (open circles), cdc14-3 cdc5-as1 (closed circles), cdc14-1 cdc5-1 (closed diamonds), cdc14-3 cdc5-1 (closed triangles) and cdc14-3 cdc5-2 (closed squares) cells were treated as in (b). Samples were taken at the indicated times to determine the percentage of cells with metaphase spindles. N = 100 cells were scored for each data point. The cell drawn in each graph is representative of the terminal phenotype of the analysed strain. Representative experiments are shown (see Methods).
Supplementary Figure 2 The cdc14 cdc5 arrest is checkpoint independent, related to Fig. 2.
(a) cdc14-1 cdc5-as1 (open circles) and cdc14-1 cdc5-as1 mec1Δ sml1Δ (closed circles), cdc14-1 cdc5-as1 rad53K227A (closed triangles) and cdc14-1 cdc5-as1 rad9Δ (closed squares) cells were arrested in G1 byα-factor in YEPD at 23 °C. When arrest was complete, cells were released in fresh YEPD medium containing the CMK inhibitor23 and incubated at 37 °C to inactivate the cdc5-as1 and cdc14-1 alleles, respectively. Samples were taken at the indicated times to determine the percentage of cells with metaphase spindles. (b) cdc14-1 cdc5-as1 (open circles) and cdc14-1 cdc5-as1 mad2Δ (closed circles), cdc14-1 cdc5-as1 mad1Δ (closed triangles) and cdc14-1 cdc5-as1 mad1Δ mad2Δ (closed squares) cells were treated as described in (a). Samples were taken at the indicated times to determine the percentage of cells with metaphase spindles. One hundred cells were scored for each data point. The cell drawn in each graph is representative of the terminal phenotype of the analysed strain. Representative experiments are shown (see Methods).
Supplementary Figure 3 Characterization of the spindle elongation defect exhibited by cdc14 cdc5 cells, related to Fig. 4.
(a) pMET-CDC20, cdc23-1 and cdc14-1 cdc5-as1 cells were arrested in G1 byα-factor in YEPD at 23 °C. When more then 90% of cells were in G1, cells were released in fresh YEPD media supplemented with the CMK inhibitor, methionine and/or incubated at 37 °C to inactivate the cdc5-as1, pMET-CDC20 and/or cdc23-1 and cdc14-1 alleles. At the terminal phenotype (240 min after G1 release spindle length measurements were taken. There is not a significant difference between the frequency of spindles >4 μm in cdc14-1 cdc5-as1 cells compared with both cdc23-1 (on average 3.3% versus 7.7%; unpaired samples t = 1.3, df = 4, p = 0.2562), and pMET-CDC20 (on average 3.3% versus 1.7%; unpaired samples t = 1.4, df = 4, p = 0.2327) cells. (b,c) cdc14-1 cdc5-as1 and pGAL-CDC6 cdc6Δmad1Δ cdc14-1 cdc5-as1 cells were treated as in Fig. 4a. Spindle lengths were measured at the 120 min (b) and 240 min (c) time-points. Mean and s.e.m. (error bars) deriving from n = three independent experiments are shown, 100 cells counted for each time-point in each experiment (a,b).
Supplementary Figure 4 Lack of FEAR network activity arrest cells at anaphase entry, related to Fig. 5.
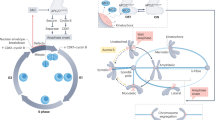
(a) Schematic representation of different models for the FEAR network. Albeit discovered a decade ago the organization of the FEAR network remains unclear. One model proposes that the Esp1–Slk19 branch act in parallel to the Spo12/Bns1 one12. A second model places all FEAR network components in the same branch36. Given the central role of Cdc5 in the release of Cdc14 (readout for FEAR network activity) it has been difficult so far to place the kinase within the network. Data in the literature are consistent with Cdc5 acting in parallel to the Spo12/Bns1 branch of the network15. Our data are consistent with a new model indicating that the network is likely composed of three branches, the Spo12/Bns1, the Esp1–Slk19 and finally the one controlled by Cdc5. (b,c) esp1-1 mcd1-1 mad1Δ and spo12Δ bns1Δ mad1Δcells (b) and cdc5-as1 mad1Δ, cdc5-as1 mad1Δesp1-1 mcd1-1, cdc5-as1 mad1Δ spo12Δbns1Δ and cdc5-as1 mad1Δ esp1-1 mcd1-1 spo12Δ bns1Δ cells (c) were arrested in G1 in YEPD at 23 °C. When more than 90% of cells reached the G1 block, cells were released in fresh YEPD medium supplemented with the CMK inhibitor and incubated at 37 °C, to inactivate esp1-1 and mcd1-1 alleles. Samples were taken at the indicated times to determine the percentage of cells with metaphase (closed circles) and anaphase (open squares) spindles (b). Clb2, Pds1 and Pgk1 protein levels were assessed by western blot analyses. Pgk1 was used as an internal loading control in immunoblots (c). n = 100 cells counted from each time-point in each experiment. Representative experiments are shown (see Methods).
Supplementary Figure 5 Deletion of CLB5 does not allow cdc14 cdc5 cells to elongate their spindles, related to Fig. 6.
(a,b) cdc5-as1 clb5Δ, cdc14-1 clb5Δ and cdc14-1 cdc5-as1 clb5Δ cells (a) and cdc5-as1 swe1Δ, cdc14-1 swe1Δ and cdc14-1 cdc5-as1 swe1Δ cells (b) were arrested in G1 byα-factor in YEPD at 23 °C. When arrest was complete, cells were released in fresh YEPD medium containing the CMK inhibitor and incubated at 37 °C to inactivate the cdc5-as1 and cdc14-1 allele, respectively. Samples were taken at the indicated times to determine the percentage of cells with metaphase (closed circles) and anaphase (open squares) spindles. n = 100 cells counted from each time-point in each experiment. The cell drawn in each graph is representative of the terminal phenotype of the analysed strain. Representative experiments are shown (see Methods).
Supplementary information
Supplementary Information
Supplementary Information (PDF 855 kb)
Rights and permissions
About this article
Cite this article
Roccuzzo, M., Visintin, C., Tili, F. et al. FEAR-mediated activation of Cdc14 is the limiting step for spindle elongation and anaphase progression. Nat Cell Biol 17, 251–261 (2015). https://doi.org/10.1038/ncb3105
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3105
This article is cited by
-
Topoisomerase II deficiency leads to a postreplicative structural shift in all Saccharomyces cerevisiae chromosomes
Scientific Reports (2021)
-
DNA double-strand breaks in telophase lead to coalescence between segregated sister chromatid loci
Nature Communications (2019)
-
Protein kinases in mitotic phosphorylation of budding yeast CENP-A
Current Genetics (2019)
-
Regulation of kinetochore configuration during mitosis
Current Genetics (2018)
-
Functions and regulation of the Polo-like kinase Cdc5 in the absence and presence of DNA damage
Current Genetics (2018)