Abstract
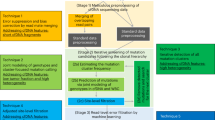
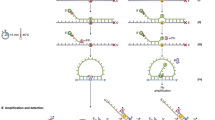
Mutations are important markers in the early detection of cancer1,2,3. Clinical specimens such as bodily fluid samples often contain a small percentage of mutated cells in a large background of normal cells. Thus, assays to detect mutations leading to cancer need to be highly sensitive and specific4,5,6,7,8. In addition, they should be possible to carry out in an automated and high-throughput manner to allow large-scale screening4,5,6,7,8. Here we describe a screening method, termed PPEM (PNA-directed PCR, primer extension, MALDI-TOF), that addresses these needs more effectively than do existing methods. DNA samples are first amplified using peptide nucleic acid (PNA)–directed PCR clamping reactions in which mutated DNA is preferentially enriched. The PCR-amplified DNA fragments are then sequenced through primer extension to generate diagnostic products. Finally, mutations are identified using matrix-assisted laser-desorption/ionization time-of-flight (MALDI-TOF) mass spectrometry. This method can detect as few as 3 copies of mutant alleles in the presence of a 10,000-fold excess of normal alleles in a robust and specific manner. In addition, the method can be adapted for simultaneous detection of multiple mutations and is amenable to high-throughput automation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Potter, J.D. Colorectal cancer: molecules and populations. J. Natl. Cancer. Inst. 91, 916–932 (1999).
Srivastava, S., Verma, M. & Henson, D.E. Biomarkers for early detection of colon cancer. Clin. Cancer. Res. 7, 1118–1126 (2001).
Hirsch, F.R., Franklin, W.A., Gazdar, A.F. & and Bunn, P.A. Jr. Early detection of lung cancer: clinical perspectives of recent advances in biology and radiology. Clin. Cancer Res. 7, 5–22 (2001).
Sidransky, D. Nucleic acid–based methods for the detection of cancer. Science 278, 1054–1058 (1997).
Ahrendt, S.A. et al. Molecular detection of tumor cells in bronchoalveolar lavage fluid from patients with early stage lung cancer. J. Natl. Cancer. Inst. 91, 332–339 (1999).
Dong, S.M. et al. Detecting colorectal cancer in stool with the use of multiple genetic targets. J. Natl. Cancer Inst. 93, 858–865 (2001).
Ahlquist, D.A. et al. Colorectal cancer screening by detection of altered human DNA in stool: feasibility of a multitarget assay panel. Gastroenterology 119, 1219–1227 (2000).
Ahrendt, S. et al. Rapid p53 sequence analysis in primary lung cancer using an oligonucleotide probe array. Proc. Natl. Acad. Sci. USA 96, 7382–7387 (1999).
Orum, H. et al. Single base pair mutation analysis by PNA directed PCR clamping. Nucleic Acids Res. 21, 5332–5336 (1993).
Behn, M. & Schuermann, M. Sensitive detection of p53 gene mutations by a "mutant enriched" PCR-SSCP technique. Nucleic Acids Res. 26, 1356–1358 (1998).
Behn, M. et al. Facilitated detection of oncogene mutations from exfoliated tissue materials by a PNA-mediated "enriched PCR" protocol. J. Pathol. 190, 69–75 (2000).
Nordhoff, E. et al. Rapid determination of short DNA sequences by the use of MALDI-MS. Nucleic Acid Res. 28, e86 (2000).
Braun, A., Little, D.P. & Koster, H. Detecting CFTR gene mutations by using primer oligo base extension and mass spectrometry. Clin. Chem. 43, 1151–1158 (1997).
Sun, X., Ding, H., Hung, K. & Guo, B.C. A new MALDI-TOF based mini-sequencing assay for genotyping of SNPs. Nucleic Acid Res. 28, e68 (2000).
Nollau, P. & Wagener, C. Methods for detection of point mutations: performance and quality assessment. IFCC Scientific Division, Committee on Molecular Biology Techniques. Clin. Chem. 43, 1114–1128 (1997).
Jacobson, D.R. & Miller, N.E. A highly sensitive assay for mutant ras genes and its application to the study of presentation and relapse genotypes in acute leukemia. Oncogene 9, 553–563 (1994).
Lehman, T.A. et al. Detection of K-ras oncogene mutation by polymerase chain reaction–based ligase chain reaction. Anal. Biochem. 239, 153–159 (1996).
Khanna, M. et al. Multiplex PCR/LDR for detection of K-ras mutations in primary colon tumors. Oncogene 17, 27–38 (1999).
Brennan, J.A. et al. Association between cigarette smoking and mutation of the p53 gene in squamous-cell carcinoma of the head and neck. N. Engl. J. Med. 332, 712–717 (1995).
Shuber, A. P., Grondin, V. J., Klinger, K. W. A simplified procedure for developing multiplex PCRs. Genome. Res. 5, 488–493 (1995).
Acknowledgements
This work was supported by NIH grants HG01815 and CA81653 (B.G.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sun, X., Hung, K., Wu, L. et al. Detection of tumor mutations in the presence of excess amounts of normal DNA. Nat Biotechnol 20, 186–189 (2002). https://doi.org/10.1038/nbt0202-186
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0202-186
This article is cited by
-
Detection of K-Ras mutations in tumour samples of patients with non-small cell lung cancer using PNA-mediated PCR clamping
British Journal of Cancer (2009)
-
Replacing PCR with COLD-PCR enriches variant DNA sequences and redefines the sensitivity of genetic testing
Nature Medicine (2008)
-
Single-tube reaction using peptide nucleic acid as both PCR clamp and sensor probe for the detection of rare mutations
Nature Protocols (2006)
-
Prevalence and clinical impact of Met Y1253D-activating point mutation in radiotherapy-treated squamous cell cancer of the oropharynx
Oncogene (2003)
-
Emerging molecular markers of cancer
Nature Reviews Cancer (2002)