Abstract
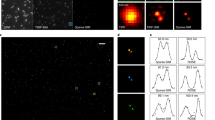
To increase the temporal resolution and maximal imaging time of super-resolution (SR) microscopy, we have developed a deconvolution algorithm for structured illumination microscopy based on Hessian matrixes (Hessian-SIM). It uses the continuity of biological structures in multiple dimensions as a priori knowledge to guide image reconstruction and attains artifact-minimized SR images with less than 10% of the photon dose used by conventional SIM while substantially outperforming current algorithms at low signal intensities. Hessian-SIM enables rapid imaging of moving vesicles or loops in the endoplasmic reticulum without motion artifacts and with a spatiotemporal resolution of 88 nm and 188 Hz. Its high sensitivity allows the use of sub-millisecond excitation pulses followed by dark recovery times to reduce photobleaching of fluorescent proteins, enabling hour-long time-lapse SR imaging of actin filaments in live cells. Finally, we observed the structural dynamics of mitochondrial cristae and structures that, to our knowledge, have not been observed previously, such as enlarged fusion pores during vesicle exocytosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hell, S.W. Far-field optical nanoscopy. Science 316, 1153–1158 (2007).
Huang, B., Bates, M. & Zhuang, X. Super-resolution fluorescence microscopy. Annu. Rev. Biochem. 78, 993–1016 (2009).
Schermelleh, L., Heintzmann, R. & Leonhardt, H. A guide to super-resolution fluorescence microscopy. J. Cell Biol. 190, 165–175 (2010).
Sengupta, P., Van Engelenburg, S. & Lippincott-Schwartz, J. Visualizing cell structure and function with point-localization superresolution imaging. Dev. Cell 23, 1092–1102 (2012).
Editorial. Artifacts of light. Nat. Methods 10, 1135 (2013).
Takakura, H. et al. Long time-lapse nanoscopy with spontaneously blinking membrane probes. Nat. Biotechnol. 35, 773–780 (2017).
Thompson, A.D. et al. Long-term live-cell STED nanoscopy of primary and cultured cells with the plasma membrane HIDE probe DiI-SiR. Angew. Chem. Int. Edn Engl. 56, 10408–10412 (2017).
Garcia-Parajo, M.F., Segers-Nolten, G.M., Veerman, J.A., Greve, J. & van Hulst, N.F. Real-time light-driven dynamics of the fluorescence emission in single green fluorescent protein molecules. Proc. Natl. Acad. Sci. USA 97, 7237–7242 (2000).
Dean, K.M. et al. Analysis of red-fluorescent proteins provides insight into dark-state conversion and photodegradation. Biophys. J. 101, 961–969 (2011).
Huang, F. et al. Video-rate nanoscopy using sCMOS camera-specific single-molecule localization algorithms. Nat. Methods 10, 653–658 (2013).
Schneider, J. et al. Ultrafast, temporally stochastic STED nanoscopy of millisecond dynamics. Nat. Methods 12, 827–830 (2015).
Carlton, P.M. et al. Fast live simultaneous multiwavelength four-dimensional optical microscopy. Proc. Natl. Acad. Sci. USA 107, 16016–16022 (2010).
Li, D. et al. Advanced imaging. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science 349, aab3500 (2015).
Gustafsson, M.G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Gustafsson, M.G. et al. Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970 (2008).
York, A.G. et al. Instant super-resolution imaging in live cells and embryos via analog image processing. Nat. Methods 10, 1122–1126 (2013).
Hayashi, S. & Okada, Y. Ultrafast superresolution fluorescence imaging with spinning disk confocal microscope optics. Mol. Biol. Cell 26, 1743–1751 (2015).
Song, L.Y. et al. Fast structured illumination microscopy using rolling shutter cameras. Meas. Sci. Technol. 27, 055401 (2016).
Nixon-Abell, J. et al. Increased spatiotemporal resolution reveals highly dynamic dense tubular matrices in the peripheral ER. Science 354, aaf3928 (2016).
Schulz, O. et al. Resolution doubling in fluorescence microscopy with confocal spinning-disk image scanning microscopy. Proc. Natl. Acad. Sci. USA 110, 21000–21005 (2013).
Schaefer, L.H., Schuster, D. & Schaffer, J. Structured illumination microscopy: artefact analysis and reduction utilizing a parameter optimization approach. J. Microsc. 216, 165–174 (2004).
Sahl, S.J. et al. Comment on “Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics”. Science 352, 527 (2016).
Chu, K. et al. Image reconstruction for structured-illumination microscopy with low signal level. Opt. Express 22, 8687–8702 (2014).
Perez, V., Chang, B.J. & Stelzer, E.H.K. Optimal 2D-SIM reconstruction by two filtering steps with Richardson-Lucy deconvolution. Sci. Rep. 6, 37149 (2016).
Müller, M., Mönkemöller, V., Hennig, S., Hübner, W. & Huser, T. Open-source image reconstruction of super-resolution structured illumination microscopy data in ImageJ. Nat. Commun. 7, 10980 (2016).
Demmerle, J. et al. Strategic and practical guidelines for successful structured illumination microscopy. Nat. Protoc. 12, 988–1010 (2017).
Keane, R.D. & Adrian, R.J. Theory of cross-correlation analysis of PIV images. Appl. Sci. Res. 49, 191–215 (1992).
Sun, T., Sun, N., Wang, J. & Tan, S. Iterative CBCT reconstruction using Hessian penalty. Phys. Med. Biol. 60, 1965–1987 (2015).
Lukinaviius, G. et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat. Methods 11, 731–733 (2014).
Nishigaki, T., Wood, C.D., Shiba, K., Baba, S.A. & Darszon, A. Stroboscopic illumination using light-emitting diodes reduces phototoxicity in fluorescence cell imaging. Biotechniques 41, 191–197 (2006).
Voets, T., Neher, E. & Moser, T. Mechanisms underlying phasic and sustained secretion in chromaffin cells from mouse adrenal slices. Neuron 23, 607–615 (1999).
Südhof, T.C. Calcium control of neurotransmitter release. Cold Spring Harb. Perspect. Biol. 4, a011353 (2012).
Zhou, Z. & Misler, S. Amperometric detection of quantal secretion from patch-clamped rat pancreatic beta-cells. J. Biol. Chem. 271, 270–277 (1996).
MacDonald, P.E., Braun, M., Galvanovskis, J. & Rorsman, P. Release of small transmitters through kiss-and-run fusion pores in rat pancreatic beta cells. Cell Metab. 4, 283–290 (2006).
Yuan, T. et al. Diacylglycerol guides the hopping of clathrin-coated pits along microtubules for exo-endocytosis coupling. Dev. Cell 35, 120–130 (2015).
Barg, S. et al. Delay between fusion pore opening and peptide release from large dense-core vesicles in neuroendocrine cells. Neuron 33, 287–299 (2002).
Ornberg, R.L. & Reese, T.S. Beginning of exocytosis captured by rapid-freezing of Limulus amebocytes. J. Cell Biol. 90, 40–54 (1981).
Jakobs, S. & Wurm, C.A. Super-resolution microscopy of mitochondria. Curr. Opin. Chem. Biol. 20, 9–15 (2014).
Shim, S.H. et al. Super-resolution fluorescence imaging of organelles in live cells with photoswitchable membrane probes. Proc. Natl. Acad. Sci. USA 109, 13978–13983 (2012).
Ball, G. et al. SIMcheck: a toolbox for successful super-resolution structured illumination microscopy. Sci. Rep. 5, 15915 (2015).
Chen, B.C. et al. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346, 1257998 (2014).
Berning, S., Willig, K.I., Steffens, H., Dibaj, P. & Hell, S.W. Nanoscopy in a living mouse brain. Science 335, 551 (2012).
Gustafsson, M.G. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl. Acad. Sci. USA 102, 13081–13086 (2005).
Shang, W. et al. Imaging Ca2+ nanosparks in heart with a new targeted biosensor. Circ. Res. 114, 412–420 (2014).
Acknowledgements
We thank B.-C. Chen and C. Shan for commenting on the optics and biological experiments and B.-C. Chen, T. Maritzen, and P. Cheng for reading the manuscript and providing suggestions. The work was supported by grants from the National Science and Technology Major Project Program (2016YFA0500400), National Natural Science Foundation of China (31327901, 31521062, 31570839, 31428004, 61375018, 61672253 and 91750203), the Major State Basic Research Program of China (2013CB531200), and Beijing Natural Science Foundation (L172003).
Author information
Authors and Affiliations
Contributions
L.C. and S.T. conceived and supervised the research; X.H. designed and built the optical system; J.F. developed the reconstruction algorithm; X.H. and L.L. performed the experiments; R.W. wrote the control software under the supervision of Y.Z. and H.M.; A.L., L.T., Y.W., H.L., Y.L., and L.W. helped with the optics, SIM reconstruction, SLM pattern generation, algorithm and biological experiments, respectively; P.X. proposed the idea of 'rolling' SIM; X.H., J.F., and L.L. analyzed the data and prepared the figures; and L.C. and S.T. wrote the paper. All of the authors participated in discussions and data interpretation.
Corresponding authors
Ethics declarations
Competing interests
L.T. works at ColdSpring Science Corporation.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–18, Supplementary Notes 1–3, Supplementary tables 1–6 (PDF 49110 kb)
Supplementary Code 1
Hessian-SIM Software package, relevant scripts and demo data. (ZIP 108449 kb)
97 Hz TIRF-SIM imaging of actin dynamics in a HUVEC for 6,800 consecutive time points.
The HUVEC labeled with Lifeact-EGFP was imaged for 6,800 consecutive time points at 97 Hz. Scale bar: 2 μm. (MOV 39034 kb)
49 Hz dual-color imaging of actin and tubulin dynamics in a HUVEC.
The HUVEC labeled with SiR-tubulin (magenta) and LifeactEGFP (green) was imaged continuously at 49 Hz. Scale bar: 2 μm. (MOV 13653 kb)
49 Hz dual-color imaging of EB3 and tubulin dynamics in an INS-1 cell.
The INS-1 cell labeled with SiR-tubulin (magenta) and EB3-EGFP (green) was imaged continuously at 49 Hz. Scale bar: 2 μm. (MOV 9481 kb)
Actin dynamics in HUVECs imaged at 10 Hz with different exposures.
Conventional SIM and Hessian-SIM compared at 10 Hz with 7ms and 0.2ms exposure time for 3,000 consecutive time points. Scale bar: 2 μm. (MOV 22129 kb)
Actin dynamics in HUVECs imaged at 1 Hz with different exposures.
Conventional SIM and Hessian-SIM compared at 1 Hz with 7ms and 0.2ms exposure time for 3,000 consecutive time points. Scale bar: 2 μm. (MOV 40057 kb)
EB3 dynamics in an INS-1 cell imaged at 188 Hz.
INS-1 cell labeled with EB3-EGFP was imaged at 188 Hz with for 3,000 consecutive time points. Scale bar: 2 μm. (MOV 18823 kb)
EB3 dynamics in an INS-1 cell imaged at 1 Hz with 0.2-ms exposure time per frame.
The INS-1 cell labeled with EB3-EGFP was imaged at 1 Hz for 3,000 consecutive time points. Scale bar: 2 μm. (MOV 47635 kb)
Resolving and tracking of rapid movements of small ER loops with 188 Hz Hessian-SIM.
The HEK293 cell was labeled with KDEL-EGFP, and imaged at 188 Hz for 6,800 consecutive time points. Scale bars: the first one is 200 nm and the later one is 2 μm. (MOV 27262 kb)
Continuous imaging of vesicle fusion labeled by VAMP2- pHluorin in an INS-1 cell for 10 min.
The INS-1 cell was labeled with VAMP2-pHluorin, and stimulated with glucose and KCl. Fusion events were detected for more than 10 min at 97 Hz frame rate (~540,000 consecutive raw images). Videos of three different durations (2-12, 280-290, and 519-529 s) were shown on the top, middle, and bottom, respectively. Scale bar: 2 μm. (MOV 5472 kb)
Vesicle fusion events with “ring” and “no ring” structures.
The INS-1 cell was labeled with VAMP2-pHluorin. Scale bar: 200 nm. (MOV 2115 kb)
Live mitochondria imaged under Hessian-SIM for 800 frames.
The mitochondria in the COS-7 cell was labeled with MitoTracker Green, and imaged under 0.5-ms exposure with an initial illumination intensity of ~18 W/cm2 light intensity (which increased by 0.05% during each SIM image). Scale bar: 2 μm. (MOV 7527 kb)
The fusion of two live mitochondria into one mitochondrion.
The COS-7 cell was labeled with MitoTracker Green. The boxed region in the left is magnified and shown on the right. Scale bars: 1 μm (left) and 0.2 μm (right). (MOV 4122 kb)
The fission of a mitochondrion into two mitochondria.
The COS-7 cell was labeled with MitoTracker Green. The boxed region in the left is magnified and shown on the right. Scale bars: 1 μm (left) and 0.5 μm (right). (MOV 859 kb)
Intra-mitochondrial cristae re-organization in which two cristae structures merged into one.
The COS-7 cell was labeled with PHB2-mScarlet and MitoTracker Green. Boxed regions in the left are magnified and shown on the right. Scale bars: 1 μm (left), 0.5 μm (center) and 0.2 μm (right). (MOV 2782 kb)
97 Hz 2D-Hessian-SIM imaging of cytosolic actin dynamics in a HUVEC for 2,000 consecutive time points.
The HUVEC labeled with Lifeact-EGFP was imaged for 2,000 consecutive time points at 97 Hz. Scale bar: 2 μm. (MOV 12998 kb)
Rights and permissions
About this article
Cite this article
Huang, X., Fan, J., Li, L. et al. Fast, long-term, super-resolution imaging with Hessian structured illumination microscopy. Nat Biotechnol 36, 451–459 (2018). https://doi.org/10.1038/nbt.4115
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4115
This article is cited by
-
Surmounting photon limits and motion artifacts for biological dynamics imaging via dual-perspective self-supervised learning
PhotoniX (2024)
-
Membrane transformations of fusion and budding
Nature Communications (2024)
-
Tracking endogenous proteins based on RNA editing-mediated genetic code expansion
Nature Chemical Biology (2024)
-
Live-cell imaging powered by computation
Nature Reviews Molecular Cell Biology (2024)
-
Bright and stable monomeric green fluorescent protein derived from StayGold
Nature Methods (2024)