Abstract
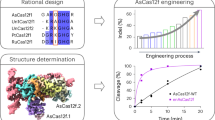
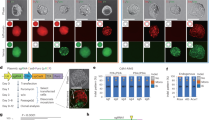
Components of the prokaryotic clustered, regularly interspaced, short palindromic repeats (CRISPR) loci have recently been repurposed for use in mammalian cells1,2,3,4,5,6. The CRISPR-associated (Cas)9 can be programmed with a single guide RNA (sgRNA) to generate site-specific DNA breaks, but there are few known rules governing on-target efficacy of this system7,8. We created a pool of sgRNAs, tiling across all possible target sites of a panel of six endogenous mouse and three endogenous human genes and quantitatively assessed their ability to produce null alleles of their target gene by antibody staining and flow cytometry. We discovered sequence features that improved activity, including a further optimization of the protospacer-adjacent motif (PAM) of Streptococcus pyogenes Cas9. The results from 1,841 sgRNAs were used to construct a predictive model of sgRNA activity to improve sgRNA design for gene editing and genetic screens. We provide an online tool for the design of highly active sgRNAs for any gene of interest.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Garneau, J.E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 39, 9275–9282 (2011).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Wang, T., Wei, J.J., Sabatini, D.M. & Lander, E.S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014).
Gagnon, J.A. et al. Efficient mutagenesis by Cas9 protein-mediated oligonucleotide insertion and large-scale assessment of single-guide RNAs. PLoS ONE 9, e98186 (2014).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Koike-Yusa, H., Li, Y., Tan, E.-P., Del Castillo Velasco-Herrera, M. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014).
Zhou, Y. et al. High-throughput screening of a CRISPR-Cas9 library for functional genomics in human cells. Nature 509, 487–491 (2014).
Luo, B. et al. Highly parallel identification of essential genes in cancer cells. Proc. Natl. Acad. Sci. USA 105, 20380–20385 (2008).
Cheung, H.W. et al. Systematic investigation of genetic vulnerabilities across cancer cell lines reveals lineage-specific dependencies in ovarian cancer. Proc. Natl. Acad. Sci. USA 108, 12372–12377 (2011).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Cho, S.W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Wu, X. et al. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat. Biotechnol. 32, 670–676 (2014).
Kuscu, C., Arslan, S., Singh, R., Thorpe, J. & Adli, M. Genome-wide analysis reveals characteristics of off-target sites bound by the Cas9 endonuclease. Nat. Biotechnol. 32, 677–683 (2014).
Reynolds, A. et al. Rational siRNA design for RNA interference. Nat. Biotechnol. 22, 326–330 (2004).
Fellmann, C. et al. Functional identification of optimized RNAi triggers using a massively parallel sensor assay. Mol. Cell 41, 733–746 (2011).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L.A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank M. Waring (Ragon Institute, Cambridge, MA) for expert assistance with flow cytometry; S. Rosenbluh (Broad Institute) for sharing the pLX_311-Cas9 vector; T. Mason (Broad Institute) for Illumina sequencing advice and execution; D. Alan, A. Brown, M. Tomko, M. Greene and T. Green (Broad Institute) for assistance in building the sgRNA design website; J. Listgarten and N. Fuso (Microsoft Research) for a critical analysis of the sgRNA activity prediction model.
Author information
Authors and Affiliations
Contributions
E.H., D.B.G., Z.T. and J.G.D. designed experiments; E.H., M.S., D.B.G. and Z.T. ran experiments; E.H. and J.G.D. analyzed experimental results; M.H. and J.G.D. analyzed sequencing data and developed analysis tools; I.S. developed the sgRNA scoring model; E.H., D.E.R. and J.G.D. wrote the manuscript with help from other authors; B.L.E., R.J.X. and D.E.R. supervised the research. J.G.D. is a Merkin Institute Fellow. Z.T. is supported by US National Institutes of Health 5T32CA009172-39.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 4–6 and 9 (PDF 4229 kb)
Supplementary Table 1
List of 6500 synthesized oligonucleotides. A small number of oligonucleotides appear more than once. (XLSX 123 kb)
Supplementary Table 2
Full dataset of Illumina sequencing reads from the mouse pool. Each sample was tagged with several different sample barcodes and thus appears in multiple columns of data. See Methods for additional details on data analysis. (XLSX 2375 kb)
Supplementary Table 3
Full dataset of Illumina sequencing reads from the human pool. (XLSX 547 kb)
Supplementary Table 7
List of 1,841 CDS-targeting sgRNAs. On-target activity is expressed as within-gene percent-rank in the marker-negative population relative to the unsorted population. The predicted sgRNA score from the final model is also given. See Methods for additional details on data analysis. (XLSX 204 kb)
Supplementary Table 8
For each single and dinucleotide feature, number of sgRNA examples, frequency of scoring in the top quintile percent-rank for that target gene, and statistical significance for over- or under-representation of sgRNAs with that feature in the most-active sgRNA quintile. (XLSX 102 kb)
Supplementary Table 10
1,278 sgRNAs targeting 414 essential genes in A375 cells. sgRNA activity is expressed as log2 fold change in abundance during two weeks of growth. See Methods for additional details on data analysis. (XLSX 252 kb)
Rights and permissions
About this article
Cite this article
Doench, J., Hartenian, E., Graham, D. et al. Rational design of highly active sgRNAs for CRISPR-Cas9–mediated gene inactivation. Nat Biotechnol 32, 1262–1267 (2014). https://doi.org/10.1038/nbt.3026
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3026
This article is cited by
-
Improved prediction of bacterial CRISPRi guide efficiency from depletion screens through mixed-effect machine learning and data integration
Genome Biology (2024)
-
Cas9-directed long-read sequencing to resolve optical genome mapping findings in leukemia diagnostics
Scientific Reports (2024)
-
A novel aminotransferase gene and its regulator acquired in Saccharomyces by a horizontal gene transfer event
BMC Biology (2023)
-
Precise DNA cleavage using CRISPR-SpRYgests
Nature Biotechnology (2023)
-
High-throughput retrieval of target sequences from complex clone libraries using CRISPRi
Nature Biotechnology (2023)