Abstract
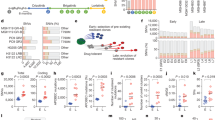
Acquired drug resistance prevents cancer therapies from achieving stable and complete responses1. Emerging evidence implicates a key role for non-mutational drug resistance mechanisms underlying the survival of residual cancer ‘persister’ cells2,3,4. The persister cell pool constitutes a reservoir from which drug-resistant tumours may emerge. Targeting persister cells therefore presents a therapeutic opportunity to impede tumour relapse5. We previously found that cancer cells in a high mesenchymal therapy-resistant cell state are dependent on the lipid hydroperoxidase GPX4 for survival6. Here we show that a similar therapy-resistant cell state underlies the behaviour of persister cells derived from a wide range of cancers and drug treatments. Consequently, we demonstrate that persister cells acquire a dependency on GPX4. Loss of GPX4 function results in selective persister cell ferroptotic death in vitro and prevents tumour relapse in mice. These findings suggest that targeting of GPX4 may represent a therapeutic strategy to prevent acquired drug resistance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Groenendijk, F. H. & Bernards, R. Drug resistance to targeted therapies: déjà vu all over again. Mol. Oncol. 8, 1067–1083 (2014)
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010)
Raha, D. et al. The cancer stem cell marker aldehyde dehydrogenase is required to maintain a drug-tolerant tumor cell subpopulation. Cancer Res. 74, 3579–3590 (2014)
Hata, A. N. et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat. Med. 22, 262–269 (2016)
Oxnard, G. R. The cellular origins of drug resistance in cancer. Nat. Med. 22, 232–234 (2016)
Viswanathan, V. S. et al. Dependency of a therapy-resistant state of cancer cells on a lipid peroxidase pathway. Nature 547, 453–457 (2017)
Fan, W. et al. MET-independent lung cancer cells evading EGFR kinase inhibitors are therapeutically susceptible to BH3 mimetic agents. Cancer Res. 71, 4494–4505 (2011)
Yagoda, N. et al. RAS–RAF–MEK-dependent oxidative cell death involving voltage-dependent anion channels. Nature 447, 865–869 (2007)
Yang, W. S. & Stockwell, B. R. Ferroptosis: death by lipid peroxidation. Trends Cell Biol. 26, 165–176 (2016)
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012)
Skouta, R. et al. Ferrostatins inhibit oxidative lipid damage and cell death in diverse disease models. J. Am. Chem. Soc. 136, 4551–4556 (2014)
Jiang, L. et al. Ferroptosis as a p53-mediated activity during tumour suppression. Nature 520, 57–62 (2015)
Friedmann Angeli, J. P. et al. Inactivation of the ferroptosis regulator Gpx4 triggers acute renal failure in mice. Nat. Cell Biol. 16, 1180–1191 (2014)
Kriska, T., Pilat, A., Schmitt, J. C. & Girotti, A. W. Sterol carrier protein-2 (SCP-2) involvement in cholesterol hydroperoxide cytotoxicity as revealed by SCP-2 inhibitor effects. J. Lipid Res. 51, 3174–3184 (2010)
Shimada, K., Hayano, M., Pagano, N. C. & Stockwell, B. R. Cell-line selectivity improves the predictive power of pharmacogenomic analyses and helps identify NADPH as biomarker for ferroptosis sensitivity. Cell Chem. Biol. 23, 225–235 (2016)
Yang, W. S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014)
Guéraud, F. et al. Chemistry and biochemistry of lipid peroxidation products. Free Radic. Res. 44, 1098–1124 (2010)
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017)
Roh, J. L., Kim, E. H., Jang, H. J., Park, J. Y. & Shin, D. Induction of ferroptotic cell death for overcoming cisplatin resistance of head and neck cancer. Cancer Lett. 381, 96–103 (2016)
Yoo, S. E. et al. Gpx4 ablation in adult mice results in a lethal phenotype accompanied by neuronal loss in brain. Free Radic. Biol. Med. 52, 1820–1827 (2012)
Salt, M. B., Bandyopadhyay, S. & McCormick, F. Epithelial-to-mesenchymal transition rewires the molecular path to PI3K-dependent proliferation. Cancer Discov. 4, 186–199 (2014)
Gelain, D. P. et al. A systematic review of human antioxidant genes. Front. Biosci. 14, 4457–4463 (2009)
Wilson, J. N., Liu, W., Brown, A. S. & Landgraf, R. Binding-induced, turn-on fluorescence of the EGFR/ERBB kinase inhibitor, lapatinib. Org. Biomol. Chem. 13, 5006–5011 (2015)
Acknowledgements
We acknowledge technical support from the UCSF ES Cell Targeting Core and UCSF Preclinical Therapeutics Core. This work was supported by grants from the National Cancer Institute (NCI) of the National Institutes of Health (NIH) (Cancer Target Discovery and Development Network grant U01CA168370 to M.T.M. and F.M., U01CA217882 to M.T.M., U01CA176152 to S.L.S., U01CA168397 to M.E.B., and R01CA212767 to M.T.M.), Susan G. Komen for the Cure Postdoctoral Fellowship KG1101214 to M.J.H., and the Howard Hughes Medical Institute (S.L.S.).
Author information
Authors and Affiliations
Contributions
M.T.M. and F.M. directed the project; M.J.H., V.S.V., S.L.S., F.M. and M.T.M. wrote the manuscript; M.J.H. performed chemical screens, RNA-seq analysis, and all cell culture experiments except V.S.V. and M.J.R. performed Kuramochi cell GPX4 inhibitor assays, D.B. assisted with ROS, glutathione and NADPH assays, and J.G. performed western blots; M.J.H. directed CRISPR editing of A375 cells and in vivo experiments that were performed at UCSF core facilities; V.S.V. and J.K.E. contributed reagents; A.M., H.D.D. and M.E.B. provided advice and project support.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks N. Chandel and P. Vandenabeele and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Additional data demonstrating persister cell ferroptosis sensitivity.
a, A375 melanoma persister cells are reversibly drug-resistant. Scale bars, 400 μm. b, Global antioxidant gene expression is downregulated in BT474 persister cells. P value calculated using a two-tailed Wilcoxon signed-rank test. c, d, MCF10A parental cells (c) and BT474 persister cells derived from carboplatin and paclitaxel (d) were treated with RSL3 or ML210 for 3 days. e, f, BT474 persister cells (e) or parental cells (f) were treated with erastin for 5 days. g, Western blot demonstrating GPX4 knockout in two distinct A375 clones (clone 1, KO1; clone 2, KO2). For gel source data, see Supplementary Fig. 1. h, BT474 parental cells co-treated with 2 μM lapatinib and RSL3 or ML210 for 3 days. Data are means and s.d. from 3 (c) or 2 (d–h) biologically independent samples. c, NS, not significant (P > 0.05); two-tailed t-test. All data are representative of two separate experiments.
Extended Data Figure 2 Additional data demonstrating that GPX4 inhibition causes ferroptosis in persister cells.
a, b, A375 persister cells were treated with RSL3 (a) or ML210 (b) and ferroptosis rescue compounds for 3 days. c, PC9 persister cells were treated with RSL3 or ML210 and ferroptosis rescue compounds for 3 days. d, Relative concentration of total labile iron in BT474 parental and persister cells. Data are means and s.d. from 2 biologically independent samples. All data are representative of two separate experiments.
Extended Data Figure 3 Additional data demonstrating that persister cells have a disabled antioxidant program and depend on GPX4 in vivo.
a, Reduced glutathione (GSH) and reduced plus oxidized glutathione (GSH + GSSG) levels in BT474 cells. b–d, Fractional viability of BT474 persister cells treated with RSL3 and antioxidant compounds (b), endogenous ROS-generating compound DMNQ (c), or the SOD1 inhibitor LCS-1 (d) for 3 days. e, ROS levels (DCF staining) in BT474 cells treated with ML210 for 1 h. f, BT474 persister cells were co-treated with ALDH inhibitor disulfiram and ferrostatin-1 for 3 days. g, Tumour volume measurements for the full time course of the experiment presented in Fig. 4d. Ferrostatin-1 was withdrawn on day 10. See Source Data for individual data points. h, Untreated A375 GPX4 wild-type or GPX4 knockout clone 1 tumour formation without ferrostatin-1 dosing. See Source Data for individual data points. i, ROS levels (DCF staining) in cells regrown without lapatinib for 15 days from BT474 persister cells and then treated with 1 μM RSL3 for 1 h. j, Persister cell GPX4 dependence model. Data are means and s.d. from three (a, e, i) or two (b–d, f) biologically independent samples or four (g) or five (h) animals. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; NS, not significant (P > 0.05); two-tailed t-tests. All data are representative of two separate experiments.
Supplementary information
Supplementary Information
This file contains the raw western blot files for Extended Data Figure 1g. (PDF 179 kb)
Supplementary Table 1
This Supplementary Table contains RNAseq expression data (FPKM) and differential expression analysis for human RefSeq genes in BT474 parental and persister cells. (XLSX 1205 kb)
Supplementary Table 2
This Supplementary Table contains Ingenuity Pathway Analysis (IPA) “Functions” results derived from analysis of BT474 parental and persister cell RNAseq data. (XLSX 135 kb)
Supplementary Table 3
This Supplementary Table contains Ingenuity Pathway Analysis (IPA) "Canonical Pathways" results derived from analysis of BT474 parental and persister cell RNAseq data. (XLSX 58 kb)
Supplementary Table 4
This Supplementary Table contains Ingenuity Pathway Analysis (IPA) "Upstream Regulators" results derived from analysis of BT474 parental and persister cell RNAseq data. (XLSX 157 kb)
Rights and permissions
About this article
Cite this article
Hangauer, M., Viswanathan, V., Ryan, M. et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 551, 247–250 (2017). https://doi.org/10.1038/nature24297
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24297
This article is cited by
-
Targeting ferroptosis for leukemia therapy: exploring novel strategies from its mechanisms and role in leukemia based on nanotechnology
European Journal of Medical Research (2024)
-
Synergism of non-thermal plasma and low concentration RSL3 triggers ferroptosis via promoting xCT lysosomal degradation through ROS/AMPK/mTOR axis in lung cancer cells
Cell Communication and Signaling (2024)
-
Low-dose hypomethylating agents cooperate with ferroptosis inducers to enhance ferroptosis by regulating the DNA methylation-mediated MAGEA6-AMPK-SLC7A11-GPX4 signaling pathway in acute myeloid leukemia
Experimental Hematology & Oncology (2024)
-
Harnessing ferroptosis for enhanced sarcoma treatment: mechanisms, progress and prospects
Experimental Hematology & Oncology (2024)
-
TrkA promotes MDM2-mediated AGPS ubiquitination and degradation to trigger prostate cancer progression
Journal of Experimental & Clinical Cancer Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.