Abstract
Metabolism has been shown to integrate with epigenetics and transcription to modulate cell fate and function1,2,3. Beyond meeting the bioenergetic and biosynthetic demands of T-cell differentiation4,5,6,7,8, whether metabolism might control T-cell fate by an epigenetic mechanism is unclear. Here, through the discovery and mechanistic characterization of a small molecule, (aminooxy)acetic acid, that reprograms the differentiation of T helper 17 (TH17) cells towards induced regulatory T (iTreg) cells, we show that increased transamination, mainly catalysed by GOT1, leads to increased levels of 2-hydroxyglutarate in differentiating TH17 cells. The accumulation of 2-hydroxyglutarate resulted in hypermethylation of the Foxp3 gene locus and inhibited Foxp3 transcription, which is essential for fate determination towards TH17 cells. Inhibition of the conversion of glutamate to α-ketoglutaric acid prevented the production of 2-hydroxyglutarate, reduced methylation of the Foxp3 gene locus, and increased Foxp3 expression. This consequently blocked the differentiation of TH17 cells by antagonizing the function of transcription factor RORγt and promoted polarization into iTreg cells. Selective inhibition of GOT1 with (aminooxy)acetic acid ameliorated experimental autoimmune encephalomyelitis in a therapeutic mouse model by regulating the balance between TH17 and iTreg cells. Targeting a glutamate-dependent metabolic pathway thus represents a new strategy for developing therapeutic agents against TH17-mediated autoimmune diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Wellen, K. E. & Thompson, C. B. A two-way street: reciprocal regulation of metabolism and signalling. Nat. Rev. Mol. Cell Biol. 13, 270–276 (2012)
Lu, C. & Thompson, C. B. Metabolic regulation of epigenetics. Cell Metab. 16, 9–17 (2012)
Kaelin, W. G. Jr & McKnight, S. L. Influence of metabolism on epigenetics and disease. Cell 153, 56–69 (2013)
Pearce, E. L., Poffenberger, M. C., Chang, C. H. & Jones, R. G. Fueling immunity: insights into metabolism and lymphocyte function. Science 342, 1242454 (2013)
Pearce, E. L. & Pearce, E. J. Metabolic pathways in immune cell activation and quiescence. Immunity 38, 633–643 (2013)
Wang, R. & Green, D. R. Metabolic checkpoints in activated T cells. Nat. Immunol. 13, 907–915 (2012)
Shi, L. Z. et al. HIF1α-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011)
Berod, L. et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat. Med. 20, 1327–1333 (2014)
Yang, X. O. et al. Molecular antagonism and plasticity of regulatory and inflammatory T cell programs. Immunity 29, 44–56 (2008)
DeBerardinis, R. J. et al. Beyond aerobic glycolysis: transformed cells can engage in glutamine metabolism that exceeds the requirement for protein and nucleotide synthesis. Proc. Natl Acad. Sci. USA 104, 19345–19350 (2007)
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013)
Wise, D. R. et al. Hypoxia promotes isocitrate dehydrogenase-dependent carboxylation of α-ketoglutarate to citrate to support cell growth and viability. Proc. Natl Acad. Sci. USA 108, 19611–19616 (2011)
Terunuma, A. et al. MYC-driven accumulation of 2-hydroxyglutarate is associated with breast cancer prognosis. J. Clin. Invest. 124, 398–412 (2014)
Zhou, L. et al. TGF-β-induced Foxp3 inhibits TH17 cell differentiation by antagonizing RORγt function. Nature 453, 236–240 (2008)
Yang, R. et al. Hydrogen sulfide promotes Tet1- and Tet2-mediated Foxp3 demethylation to drive regulatory T cell differentiation and maintain immune homeostasis. Immunity 43, 251–263 (2015)
Yue, X. et al. Control of Foxp3 stability through modulation of TET activity. J. Exp. Med. 213, 377–397 (2016)
Kim, H. P. & Leonard, W. J. CREB/ATF-dependent T cell receptor-induced FoxP3 gene expression: a role for DNA methylation. J. Exp. Med. 204, 1543–1551 (2007)
Xu, T. et al. Ursolic acid suppresses interleukin-17 (IL-17) production by selectively antagonizing the function of RORγt protein. J. Biol. Chem. 286, 22707–22710 (2011)
Bettelli, E. et al. Myelin oligodendrocyte glycoprotein-specific T cell receptor transgenic mice develop spontaneous autoimmune optic neuritis. J. Exp. Med. 197, 1073–1081 (2003)
Munger, J. et al. Systems-level metabolic flux profiling identifies fatty acid synthesis as a target for antiviral therapy. Nat. Biotechnol. 26, 1179–1186 (2008)
Kernytsky, A. et al. IDH2 mutation-induced histone and DNA hypermethylation is progressively reversed by small-molecule inhibition. Blood 125, 296–303 (2015)
Buescher, J. M., Moco, S., Sauer, U. & Zamboni, N. Ultrahigh performance liquid chromatography–tandem mass spectrometry method for fast and robust quantification of anionic and aromatic metabolites. Anal. Chem. 82, 4403–4412 (2010)
Ichiyama, K. et al. The methylcytosine dioxygenase Tet2 promotes DNA demethylation and activation of cytokine gene expression in T cells. Immunity 42, 613–626 (2015)
Toker, A. et al. Active demethylation of the Foxp3 locus leads to the generation of stable regulatory T cells within the thymus. J. Immunol. 190, 3180–3188 (2013)
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007)
Zhang, Y. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Acknowledgements
S.D. is supported by funding from the Gladstone Institutes and Tsinghua University. X.W. is supported by a grant from NFSC (31570884). A Race to Erase MS Young Investigator Award, and American Heart Association Scientist Development Grant were awarded to J.K.R. and NIH/NINDS R35 NS097976 was awarded to K.A. We thank G. Howard for help with editing the manuscript, and H. W. Lim for discussion.
Author information
Authors and Affiliations
Contributions
T.X. and S.D. conceived the project; T.X., K.M.S., X.W., C.D., E.M.D. and S.D. designed the experiments. T.X., K. Li, T.M., H.W., S.Z., N.C. and Y.Z. performed the in vitro experiments; T.X. prepared the samples for metabolic analysis; K.M.S. conducted all metabolomics and flux analysis; D.Z. helped to analyse 2-HG levels. X.W. and L.N. performed active EAE experiments; T.X. and J.K.R. performed transfer EAE experiment (T.X. prepared cells and injected cells into mice, and J.K.R. scored the mice, and T.X. and J.K.R. analysed T cells infiltrated into the central nervous system). M.X. prepared chemicals. K. Liu analysed the data from (h)MeDIP–seq. K.A. designed experiments and analysed data. T.X., K.M.S., X.W., C.D., E.M.D. and S.D. wrote the manuscript; K.A., C.D., E.M.D. and S.D. edited the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks L. O’Neill and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 AOA reprograms TH17 cell differentiation towards iTreg cells.
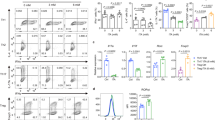
a, Schematic of the screening procedure. b, Effects of AOA on TH17 cell differentiation. c, The effect of AOA on mTOR, AMPK and c-Myc. Differentiating TH17 cells (with or without 0.75 mM AOA) were collected for analysis of phosphorylated AMPKα1 (p-AMPKα1), the mTOR downstream signalling proteins S6K and 4E-BP1, and c-Myc and β-actin. Dimethylsulfoxide (DMSO) was used as a control. d, The effect of AOA on the proliferation of TH17 cells. CD4 naive T cells were labelled with CellTrace Violet, and then cultured under TH17 or iTreg cell conditions (which were the same as TH17 cell conditions except without IL-1β, IL-6 and IL-23). At the end of experiment, the cells were analysed by intracellular staining. e, AOA promoted iTreg cell differentiation. f, AOA did not affect the survival of TH17 cell culture. The cells were differentiated under TH17 cell conditions with or without AOA. At the end of the differentiation, 7-AAD was added and the cells were immediately collected for analysis.
Extended Data Figure 2 15N-labelling analysis showed that GOT1 mediates most transamination reactions and represents the main target for AOA in TH17 cells.
a, Schematic of [15N]α-glutamine metabolism in the transamination reaction. b, The ratios of 15N-labelled amino acids to their respective intracellular amino acid pool (related to Fig. 1f). Differentiating TH17 cells or iTreg cells (68 h) were fed with 2 mM [15N]α-glutamine for 4 h. The cells were collected for intracellular metabolites analysis. The ratios of 15N-labelled aspartate, glycine, alanine and serine to their total respective amino acid pools were calculated. c, AOA, as a pan-transaminase inhibitor, inhibited de novo synthesis for several amino acids (via transamination) in both cell types in addition to aspartate. The ratio of [15N]amino acid to [15N]glutamate was calculated, and this ratio was further normalized to that in TH17 cells to demonstrate that AOA inhibited de novo synthesis for several amino acids. Our rescue results clearly showed (in Fig. 2f, g) that dimethyl-α-KG can largely rescued the effect of AOA on both TH17 and iTreg cell differentiation. Thus, from a metabolic perspective, the effects of AOA can be largely attributed to its inhibitory effect on α-KG formation (carbon metabolism), rather than its inhibitory effect on amino acid synthesis (nitrogen metabolism). Therefore, we focus on carbon metabolism of glutamate in this study. d, GOT1 is the main target for AOA in TH17 cells. Differentiating TH17 cells were infected with retrovirus containing shRNA against Got1 or control shRNA. The GFP+ cells were then purified on day 3, and further cultured under TH17 cell conditions with/or without AOA. At the end of differentiation (day 6), the cells were collected for analysis of FOXP3 and IL-17 by intracellular staining. It is clear that AOA can further inhibit TH17 cell differentiation; however, the effect is subtle and supports the conclusion that GOT1 is the main target for AOA under TH17 conditions. Data in b and c are mean ± s.d. of three technical replicates from a representative experiment. In d, representative flow cytometry data from two experiments are presented.
Extended Data Figure 3 Metabolic profiling of TH17 cells and iTreg cells.
a, Intracellular metabolites profiling of differentiating TH17 cells and iTreg cells performed by LC–MS. Asterisks denote metabolites with differential abundance between TH17 and iTreg cells, and this effect is inhibited by AOA. b, The 2-HG concentration is much higher in differentiating TH17 cells than in iTreg cells along the differentiation time line. c, AOA significantly decreased the abundance of 2-HG, whereas it did not affect the levels of l-lactic acid, glutathione, oxidized glutathione, l-aspartate, and slightly decreased α-KG and slightly increased glutamate relative to a. The relative levels of 2-HG, l-lactic acid, l-glutamate, glutathione, oxidized glutathione, l-aspartate and α-KG from a were re-plotted in c. Data in b and c are mean ± s.d. of three replicates from a representative experiment of three independent experiments.
Extended Data Figure 4 Exogenously added cell-permeable metabolites (citrate, succinate, fumarate and malate) did not rescue the effects of AOA on TH17 and iTreg cell differentiation.
a, Cell-permeable dimethyl esters of succinate, fumarate, malate, citrate, NAC or GSH did not rescue the inhibitory effects of AOA on TH17 cell differentiation. Cell-permeable metabolites (0.5 mM succinate, 50 μM fumarate, 0.5 mM malate, 0.5 mM citrate, 1 mM NAC, and 1 mM GSH) were individually added to differentiating TH17 cells in the presence of AOA. At the end of the differentiation (day 6), the cells were re-stimulated and analysed by intracellular staining of FOXP3 and IL-17. b, Cell-permeable dimethyl esters of succinate, fumarate, malate or citrate, did not rescue the effects of AOA on iTreg cell differentiation. Cell permeable metabolites (0.5 mM succinate, 50 μM fumarate, 0.5 mM malate, 0.5 mM citrate) were individually added to differentiating iTreg cells in the presence of AOA. At the end of the differentiation (day 5), the cells were directly analysed for FOXP3–GFP. Representative flow cytometry data (left panels) are from three independent experiments. Bar graphs are mean ± s.d. of three independent experiments. **P < 0.01; ***P < 0.001 by Student’s t-test.
Extended Data Figure 5 The effect of cell-permeable metabolites (α-KG, 2-HG, citrate, succinate, fumarate and malate) on the differentiation of TH17 or iTreg cells.
a, Indicated dimethyl metabolites were added into TH17 cell culture, and the cells were analysed on day 4 by intracellular staining of FOXP3 and IL-17. b, The indicated metabolites were added to iTreg cell culture, and cells were analysed on day 5 by intracellular staining of FOXP3 and IL-17. c, d, 2-HG did not affect cell proliferation and survival. c, CD4 naive T cells were labelled with CellTrace Violet according to the manufacturer’s protocol. The cells were then differentiated under TH17 cell conditions. The cells were then stimulated and collected for intracellular staining at day 4. d, CD4 naive T cells from double-reporter mice were differentiated under TH17 cell conditions, and live cells were analysed at day 4; 7-AAD was added just before analysis. Representative flow cytometry data are from three independent experiments. Bar graphs are mean ± s.d. of three independent experiments. ***P < 0.001 by Student’s t-test.
Extended Data Figure 6 Differentiating TH17 cells highly express IDH1 and IDH2, and shRNA against IDH1 and IDH2 suppresses the expression of IDH1 and IDH2, decreases DNA methylation at the Foxp3 locus, and suppresses TH17 cell differentiation.
a, Differentiating TH17 cells or iTreg cells (day 3) were collected for mRNA expression analysis. All expression levels were normalized to β-actin (Actb), and the expression level of each enzyme was normalized to that in differentiating iTreg cells. b, Differentiated TH17 cells and iTreg cells have similar expression levels of IDH3. The experiment was performed as described in a. mRNA expression of IDH3 subunits (Idh3a, Idh3b and Idh3g) was normalized to β-actin, and plotted relative to the gene expression level in iTreg cells. c, shRNA knockdown of IDH1 and IDH2 efficiently suppressed the mRNA expression of Idh1, Idh2, Il17a and Il17f, and increases mRNA expression of Foxp3. Infected cells (GFP+ cells) containing shRNA against IDH1 and IDH2 were FACS sorted and re-stimulated with anti-CD3 and anti-CD28 for mRNA expression analysis. Expression was normalized to β-actin. d, shRNA knockdown of IDH1 and IDH2 decreased the methylation level at the Foxp3 locus. Differentiating TH17 cells were infected with retrovirus containing Idh1 and Idh2 shRNA or control shRNA (shControl). At the end of the differentiation (day 6), GFP+ cells were collected for DNA methylation analysis of the Foxp3 promoter and its intronic CpG island by bisulfate sequencing. Filled circles denote methylated cytosine; open circles denote demethylated cytosine. Male mice were used owing to X-chromosome inactivation. e, Knockdown of IDH1 and IDH2 suppressed TH17 cell differentiation, and this can be reversed by addition of cell permeable (R)-2-HG. Cell permeable (R)-2-HG was added to differentiating TH17 cells at 6 h. The cells were infected with retrovirus containing shRNA. GFP+ cells were purified at day 3 for further culture under TH17 cell conditions, and cell permeable (R)-2-HG was added to the culture until the end of the experiment. Cells were then collected for analysis of FOXP3 and IL-17. f, Knockdown of IDH1 and IDH2 has minimal effect on HIF1α expression. Differentiating TH17 cells were infected with retrovirus containing Idh1 shRNA, Idh2 shRNA and Idh1 plus Idh2 shRNA or control shRNA. Cells were then analysed for IDH1, IDH2 and HIF1α expression. Data in a–c are mean ± s.d.(n = 3) of three technical replicates from a representative experiments of three experiments. Data in e (n = 3) are mean ± s.d. of three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001 by Student’s t-test.
Extended Data Figure 7 TET1 and TET2 control FOXP3 expression during TH17 cell differentiation.
a, Tet1 and Tet2 double-knockout (DKO; Tet1−/−Tet2fl/flCD4-Cre+) promoted TH17 cell differentiation, and largely abrogated the promoting effect of (R)-2-HG on TH17 cell differentiation in wild-type T cells. CD4 naive T cells from Tet1/2 DKO or control mice were differentiated under TH17 cell conditions with or without cell permeable (R)-2-HG. At day 4, cells were collected for analysis of FOXP3 and IL-17 by intracellular staining. Results are consistent with a previous study that showed that Tet1/2 DKO regulatory T cells can be more easily and efficiently converted into TH17 cells15. b, Tet1/2 DKO partially diminished the inhibitory effect of AOA on TH17 cell differentiation. CD4 naive T cells from Tet1/2 DKO or control mice were differentiated under TH17 cell conditions with or without AOA. Cells were collected for analysis of FOXP3 and IL-17 at day 6. c, A reduced concentration of TGFβ largely abolished the effect of Tet1/2 DKO on TH17 cell differentiation, and Tet1/2 DKO decreased IL-17 expression when the TGFβ concentration is low enough, indicating that TET1/2 proteins have a dual function in the fate determination of TH17 cell differentiation. CD4 naive T cells derived from Tet1/2 DKO or control mice were differentiated under TH17 cell conditions with varied concentrations of TGFβ. Cells were collected for intracellular analysis at day 4. A previous study showed that TET2 positively regulate TH17 differentiation by binding to the Il17a gene locus23. However, our results show that Tet1/2 DKO T cells enhance TH17 cell differentiation, as determined by increased IL-17 and reduced FOXP3 expression at day 4 in TH17 cell culture. Because Tet enzyme activity is very sensitive to various exogenous stimuli, as described previously16, the discrepancy between our results and those of Ichiyama et al.23 could be caused by different culture conditions, in which our TH17 polarizing condition yielded a high amount of FOXP3 even at later stage of TH17 differentiation (day 4). To test this, we reduced the amount of TGFβ in our cultures, and consistent with Ichiyama et al.23, Tet1/2 DKO caused a significant reduction in TH17 cell differentiation under low TGFβ concentrations (c). A high TGFβ concentration may induce strong and persistent activation of SMAD3 and STAT5, which then recruit TET enzymes to the Foxp3 gene locus and promote its expression by inducing or maintaining the demethylation status15. Conversely, a low TGFβ concentration results in recruitment of TET enzymes to the Il17 gene locus, which plays a dominant role in regulating TH17 differentiation. We have thus identified dual functions of TET proteins in the fate determination of TH17 cell differentiation. In summary, the role of TET1/2 during TH17 cell differentiation is dynamic and dependent on the expression of FOXP3. In a–c, representative flow cytometry data from five (a), three (b) and two (c) independent experiments are shown. In a and b, bar graphs are mean ± s.d. of five (a) or three (b) independent experiments. In c, bar graphs are mean ± s.d. of three technical replicates from a representative experiment of two independent experiments. *P < 0.05; ***P < 0.001 by Student’s t-test.
Extended Data Figure 8 AOA and (R)-2-HG selectively affect DNA hydroxymethylation and methylation at the Foxp3 locus, but not other important lineage-specific signature gene loci examined by (h)MeDIP–seq.
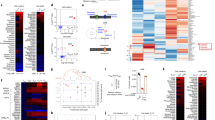
a, b, Exogenous addition of dimethyl-(R)-2-HG selectively decreases the 5hmC signal and increases the 5mC signal at the Foxp3 locus. DNA extracted from differentiating TH17 cells (day 4) in the absence or presence of 0.75 mM dimethyl-(R)-2-HG was immunoprecipitated with antibodies to 5hmC (a) or 5mC (b), followed by deep sequencing (see Methods). a, Differential peaks from hMeDIP. 5hmC peaks located in the Foxp3 conservative region or in other conservative regions such as Il4, Il5, Il13, Il10, Ifng and Rorc were highlighted. Peaks 1, 2 and 11 were located in Gata3; peak 3 in Rorc; peak 4 in Il4; peaks 5 and 16 in Il10; peak 6 in Ifng; peaks 7 and 10 in Il5; peaks 8 and 9 in Tbx21; peaks 12 in Il13; peaks 13 and 15 in Il17a; peak 14 in Il17f. (R)-2-HG decreased the 5hmC signal in 17,676 out of 330,582 peaks detected. However, (R)-2-HG did not decrease DNA hydroxymethylation at Il4, Il5, Il10, Il13, Il17a/f, Ifng, Rorc and Tbx21 loci. b, The 5mC signal was increased in 3,402 out of the 17,676 peaks with a decreased 5hmC signal. (R)-2-HG selectively decreased the 5hmC signal and increased the 5mC signal at the Foxp3 promoter and CNS2 region, but did not affect the 5hmC or 5mC signal at Ifng, Il4/5/10/13, Il17a/f, Tbx21 or Rorc. Notably, exogenous dimethyl-(R)-2-HG did not affect the 5hmC or 5mC signal at only the Foxp3 locus, and instead had a more broad effect at many loci. c, d, AOA treatment selectively affect DNA hydroxymethylation and methylation at the Foxp3 locus. 5hmC peaks located in the Foxp3 conservative region or in other conservative regions such as at Il4, Il5, Il13, Il10, Ifng and Rorc were highlighted; labels are as in a. AOA increased the 5hmC signal in 11,896 out of 330,582 peaks detected. AOA increased hydroxymethylation at the Foxp3 promoter and CNS2 region, but did not affect DNA hydroxymethylation at regions such as Ifng, Il4, Il5, Il10, Il13, Rorc and Tbx21. Notably, AOA treatment reduced the 5hmC signal at Il17a/f loci, probably owing to the antagonistic effect of FOXP3 on RORγt to recruit TET proteins to Il17a/f loci. d, The 5mC signal was increased in 1,643 out of the 11,896 peaks with an increased 5hmC signal. AOA decreased the 5mC signal at the Foxp3 promoter. Notably, changes in the 5mC signal at the Foxp3 CNS2 region were not detected, probably because the CNS2 region is largely methylated (around 70–80% of it is methylated in iTreg cells), and changes in 5mC are more subtle and harder to detect than changes in 5hmC. Notably, both AOA and 2-HG affected the 5hmC, but not 5mC, signal at Gata3. However, Gata3 expression is not regulated by DNA hydroxymethylation as shown in a previous study23, therefore 2-HG and AOA are unlikely to affect Gata3 expression. e, 17,676 peaks with increased 5hmC signal by AOA (from a, blue) overlapped with 11,896 peaks with decreased 5hmC signal by 2-HG (from c, green). f, 3,402 peaks with increased 5mC by 2-HG (from b, purple) overlapped with 1,643 peaks with decreased 5mC signal by AOA (from d, red). These epigenetic analyses showed that the effect of AOA and 2-HG on Foxp3 compared with other T-lineage-related genes is highly selective, despite their more broad effects on genome-wide DNA methylation and hydroxymethylation. Although demethylation at the Foxp3 locus promotes FOXP3 expression, the expression of many other genes is not regulated by DNA demethylation, such as Rorc and Gata323. Treatment of T cells with AOA under TH17 cell conditions stabilizes FOXP3 expression, but does not have much of an effect on the expression of other lineage-specific transcription factors, such as Gata3 and Rorc. FOXP3 can antagonize the function of RORγt to suppress expression of TH17 cell signature genes as well as recruit DNMT1 to the gene loci of proinflammatory cytokines or signature genes to promote methylation at these loci, suppressing their expression24. In addition, FOXP3 functions as both a transcriptional activator to directly activate its target genes required for iTreg cell differentiation/function, and a transcriptional repressor to directly suppress the genes associated with effector T cell function, resulting in iTreg cell fate25.
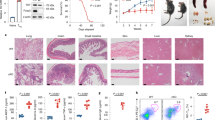
Extended Data Figure 9 shRNA knockdown of GOT1 ameliorated mouse EAE in a TH17-polarized adoptive transfer EAE model.
a, Disease score. b, The T cells infiltrated into the central nervous system were analysed (gated on Thy1.1+ cells), and representative flow cytometry data are shown. c, Statistics for the T cells infiltrated into the central nervous system (CNS). Data in a and c are mean ± s.d. *P < 0.05; **P < 0.01; ***P < 0.001 by Student’s t-test. d, Quantification of the numbers of cells infiltrated into the central nervous system regions. e, Schematic of our mechanistic model. The cell number in Fig. 4d–g was retrieved from flow cytometry data. Although we did not count the number of individual populations, we noticed that the total number of cells infiltrating into the central nervous system was reduced by Got1 shRNA; fewer than 50,000 cells were recovered from the Got1 shRNA group, whereas more than 50,000 cells were recovered from control mice. Thus, for mice receiving T cells infected with shRNA control virus, the cell number was calculated from a total of 50,000 live cells acquired by flow cytometry (this group of mice had a much higher number of total infiltrated cells, and we stopped acquiring cells after the total number reached the 50,000 threshold). Mice receiving T cells infected with Got1 shRNA virus had fewer cells infiltrated into the central nervous system, and collected cells did not reach the 50,000 threshold during flow cytometry; therefore, the cell numbers represent all of the cells infiltrated into the central nervous system. Despite the low accuracy, Got1 shRNA decreased the total absolute number of cells infiltrated into the central nervous system. For two other EAE experiments, cell numbers were counted immediately after isolating cells from the central nervous system.
Supplementary information
Supplementary Information
This file contains the uncropped blots from Extended Data Figures 1 and 5 and general information for characterization of (R)-dimethyl 2-hydroxyglutarate
Supplementary Data 1
Supplementary Data 1 contains the list of primer for qRT-PCR.
Supplementary Data 2
Supplementary Data 1 contains the list of genes.
Rights and permissions
About this article
Cite this article
Xu, T., Stewart, K., Wang, X. et al. Metabolic control of TH17 and induced Treg cell balance by an epigenetic mechanism. Nature 548, 228–233 (2017). https://doi.org/10.1038/nature23475
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23475
This article is cited by
-
Dual inhibition of glycolysis and glutaminolysis for synergistic therapy of rheumatoid arthritis
Arthritis Research & Therapy (2023)
-
Reply to: GOT1 constrains TH17 cell differentiation, while promoting iTreg cell differentiation
Nature (2023)
-
GOT1 constrains TH17 cell differentiation, while promoting iTreg cell differentiation
Nature (2023)
-
Organoids transplantation attenuates intestinal ischemia/reperfusion injury in mice through L-Malic acid-mediated M2 macrophage polarization
Nature Communications (2023)
-
Itaconate ameliorates autoimmunity by modulating T cell imbalance via metabolic and epigenetic reprogramming
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.