Abstract
Autism spectrum disorder (ASD) comprises a range of neurodevelopmental disorders characterized by deficits in social interaction and communication as well as by restricted and repetitive behaviours1. ASD has a strong genetic component with high heritability. Exome sequencing analysis has recently identified many de novo mutations in a variety of genes in individuals with ASD2,3, with CHD8, a gene encoding a chromatin remodeller, being most frequently affected4,5,6,7,8. Whether CHD8 mutations are causative for ASD and how they might establish ASD traits have remained unknown. Here we show that mice heterozygous for Chd8 mutations manifest ASD-like behavioural characteristics including increased anxiety, repetitive behaviour, and altered social behaviour. CHD8 haploinsufficiency did not result in prominent changes in the expression of a few specific genes but instead gave rise to small but global changes in gene expression in the mouse brain, reminiscent of those in the brains of patients with ASD. Gene set enrichment analysis revealed that neurodevelopment was delayed in the mutant mouse embryos. Furthermore, reduced expression of CHD8 was associated with abnormal activation of RE-1 silencing transcription factor (REST), which suppresses the transcription of many neuronal genes. REST activation was also observed in the brains of humans with ASD, and CHD8 was found to interact physically with REST in the mouse brain. Our results are thus consistent with the notion that CHD8 haploinsufficiency is a highly penetrant risk factor for ASD, with disease pathogenesis probably resulting from a delay in neurodevelopment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Abrahams, B. S. & Geschwind, D. H. Advances in autism genetics: on the threshold of a new neurobiology. Nat. Rev. Genet. 9, 341–355 (2008)
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012)
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–299 (2012)
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–250 (2012)
Talkowski, M. E. et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell 149, 525–537 (2012)
Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012)
O’Roak, B. J. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012)
Bernier, R. et al. Disruptive CHD8 mutations define a subtype of autism early in development. Cell 158, 263–276 (2014)
Hall, J. A. & Georgel, P. T. CHD proteins: a diverse family with strong ties. Biochem. Cell Biol. 85, 463–476 (2007)
Marfella, C. G. & Imbalzano, A. N. The Chd family of chromatin remodelers. Mutat. Res. 618, 30–40 (2007)
Thompson, B. A., Tremblay, V., Lin, G. & Bochar, D. A. CHD8 is an ATP-dependent chromatin remodeling factor that regulates β-catenin target genes. Mol. Cell. Biol. 28, 3894–3904 (2008)
Nishiyama, M. et al. CHD8 suppresses p53-mediated apoptosis through histone H1 recruitment during early embryogenesis. Nat. Cell Biol. 11, 172–182 (2009)
Nishiyama, M., Skoultchi, A. I. & Nakayama, K. I. Histone H1 recruitment by CHD8 is essential for suppression of the Wnt-β-catenin signaling pathway. Mol. Cell. Biol. 32, 501–512 (2012)
Nishiyama, M. et al. Early embryonic death in mice lacking the β-catenin-binding protein Duplin. Mol. Cell. Biol. 24, 8386–8394 (2004)
Nakatani, J. et al. Abnormal behavior in a chromosome-engineered mouse model for human 15q11-13 duplication seen in autism. Cell 137, 1235–1246 (2009)
Chao, H. T. et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468, 263–269 (2010)
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011)
Han, S. et al. Autistic-like behaviour in Scn1a+/− mice and rescue by enhanced GABA-mediated neurotransmission. Nature 489, 385–390 (2012)
Peñagarikano, O. et al. Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell 147, 235–246 (2011)
Sugathan, A. et al. CHD8 regulates neurodevelopmental pathways associated with autism spectrum disorder in neural progenitors. Proc. Natl Acad. Sci. USA 111, E4468–E4477 (2014)
Cotney, J. et al. The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment. Nat. Commun. 6, 6404 (2015)
Voineagu, I. et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474, 380–384 (2011)
Parikshak, N. N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 155, 1008–1021 (2013)
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014)
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011)
Krumm, N., O’Roak, B. J., Shendure, J. & Eichler, E. E. A de novo convergence of autism genetics and molecular neuroscience. Trends Neurosci. 37, 95–105 (2014)
Ballas, N., Grunseich, C., Lu, D. D., Speh, J. C. & Mandel, G. REST and its corepressors mediate plasticity of neuronal gene chromatin throughout neurogenesis. Cell 121, 645–657 (2005)
Thiel, G., Ekici, M. & Rössler, O. G. RE-1 silencing transcription factor (REST): a regulator of neuronal development and neuronal/endocrine function. Cell Tissue Res. 359, 99–109 (2015)
Johnson, R. et al. REST regulates distinct transcriptional networks in embryonic and neural stem cells. PLoS Biol. 6, e256 (2008)
Willsey, A. J. et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 155, 997–1007 (2013)
Gu, H., Zou, Y. R. & Rajewsky, K. Independent control of immunoglobulin switch recombination at individual switch regions evidenced through Cre-loxP-mediated gene targeting. Cell 73, 1155–1164 (1993)
Sado, Y. et al. Establishment by the rat lymph node method of epitope-defined monoclonal antibodies recognizing the six different α chains of human type IV collagen. Histochem. Cell Biol. 104, 267–275 (1995)
Kamura, T. et al. VHL-box and SOCS-box domains determine binding specificity for Cul2-Rbx1 and Cul5-Rbx2 modules of ubiquitin ligases. Genes Dev. 18, 3055–3065 (2004)
Cluny, N. L., Keenan, C. M., Lutz, B., Piomelli, D. & Sharkey, K. A. The identification of peroxisome proliferator-activated receptor alpha-independent effects of oleoylethanolamide on intestinal transit in mice. Neurogastroenterol. Motil. 21, 420–429 (2009)
Takao, K. et al. Deficiency of schnurri-2, an MHC enhancer binding protein, induces mild chronic inflammation in the brain and confers molecular, neuronal, and behavioral phenotypes related to schizophrenia. Neuropsychopharmacology 38, 1409–1425 (2013)
Koshimizu, H., Takao, K., Matozaki, T., Ohnishi, H. & Miyakawa, T. Comprehensive behavioral analysis of cluster of differentiation 47 knockout mice. PLoS One 9, e89584 (2014)
Kazdoba, T. M., Leach, P. T. & Crawley, J. N. Behavioral phenotypes of genetic mouse models of autism. Genes Brain Behav. 15, 7–26 (2016)
Shoji, H., Hagihara, H., Takao, K., Hattori, S. & Miyakawa, T. T-maze forced alternation and left-right discrimination tasks for assessing working and reference memory in mice. J. Vis. Exp. 60, 3300 (2012)
Deacon, R. M. Assessing nest building in mice. Nat. Protocols 1, 1117–1119 (2006)
Odawara, J. et al. The classification of mRNA expression levels by the phosphorylation state of RNAPII CTD based on a combined genome-wide approach. BMC Genomics 12, 516 (2011)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Nieuwenhuis, S., Forstmann, B. U. & Wagenmakers, E. J. Erroneous analyses of interactions in neuroscience: a problem of significance. Nat. Neurosci. 14, 1105–1107 (2011)
Benjamini, Y., Drai, D., Elmer, G., Kafkafi, N. & Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 125, 279–284 (2001)
Hayashi, S. & McMahon, A. P. Efficient recombination in diverse tissues by a tamoxifen-inducible form of Cre: a tool for temporally regulated gene activation/inactivation in the mouse. Dev. Biol. 244, 305–318 (2002)
Subtil-Rodriguez, A. et al. The chromatin remodeller CHD8 is required for E2F-dependent transcription activation of S-phase genes. Nucleic Acids Res. 42, 2185–2196 (2014)
Acknowledgements
We thank Y. Kita, K. Tsunematsu, K. Maehara, S. Hirata, M. Kato, Y. Nakajo, T. Akasaka, M. Tanaka, Y. Yamada and K. Takeda for technical assistance; as well as K. Tamada for discussion. Computed tomography was supported by the Center for Advanced Instrumental and Educational Support, Faculty of Agriculture, Kyushu University. This study was supported in part by KAKENHI and by a Grant-in-Aid for Scientific Research on Innovative Areas (Comprehensive Brain Science Network) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Author information
Authors and Affiliations
Contributions
M.N. and A.K. assisted with animal preparation and molecular biology experiments. H.S. and T.M. conducted behavioural studies. Y.O., T.S. and M.S. performed sequencing and data analysis. Y.K. performed all other experiments and data analysis. T.T. interpreted results. K.I.N. coordinated the study and wrote the manuscript. All authors discussed the data and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks E. Eichler, C. Powell and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Generation of Chd8L-specific knockout mice.
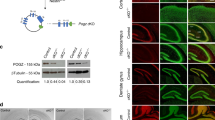
a, Structural organization of CHD8 isoforms. Circles4, triangles7 and asterisks8 indicate point mutations identified in ASD patients. In addition, copy number variants in ASD patients include a translocation mutation in the 5′ untranslated region5 and gene duplication8. b, Schematic representation of the wild-type mouse Chd8 allele, the targeting vector, the Chd8 allele with a loxP-neo cassette (Flox-neo), the floxed Chd8 allele, and the floxed Chd8 allele after removal of exons 11–13 by Cre recombinase (deletion). Crossing and noncrossing dotted lines indicate the regions of homologous recombination and deletion, respectively. The expected sizes of DNA fragments in Southern blot analysis with the indicated probe are shown. Exons and loxP sites are indicated by numbered boxes and by open triangles, respectively. Bg, BglII; DT, diphtheria toxin cassette. c, Southern blot analysis of BglII-digested genomic DNA from embryonic stem cells of the indicated Chd8 genotypes with the probe shown in b. d, Immunoblot analysis of CHD8 in tamoxifen-treated mouse embryonic fibroblasts (MEFs) derived from animals of the indicated genotypes. CAG-Cre-ERT2/Chd8LF/F mice were obtained from crosses of Chd8LF/F mice with CAG-Cre-ERT2 transgenic mice45. e, Chd8+/+, Chd8+/∆L and Chd8∆L/∆L mouse embryos at E8.5–12.5. Scale bars, 1 mm. f, Immunoblot analysis of CHD8 in the whole brain or olfactory bulb of E10.5, E14.5, E18.5 or adult (9 weeks of age) mice of the indicated genotypes. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 2 Abundance of Chd8 mRNA and CHD8 protein in various mouse tissues and brain regions.
a, b, Quantitative PCR with reverse transcription (qRT–PCR) analysis of Chd8L (a) and Chd8S (b) mRNAs in the whole brain or brain regions of E10.5, E14.5, E18.5 or adult (9 weeks of age) Chd8+/∆SL or Chd8+/∆L mice relative to those for wild-type mice. c–f, Abundance of CHD8L and CHD8S isoforms at both mRNA and protein levels in the indicated tissues of adult Chd8+/+, Chd8+/∆SL, and Chd8+/∆L mice (n = 6 for protein of Chd8+/∆SL, n = 3 for others). Data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 3 Macrocephaly and intestinal problem but otherwise normal general health of Chd8 mutant mice.
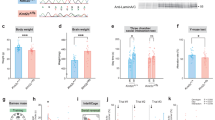
a, Brain weight of E14.5 and E18.5 mice. b, Brain volume at 9 weeks of age as determined by computed tomography (n = 8 mice per genotype). c, Body weight of E14.5, E18.5 and adult (9 weeks of age) mice. d, Intestine length and intestinal transit in 9-week-old mice (n = 30 animals of each genotype). e–h, Body temperature (e) as well as latency to falling in the wire-hang test (f), latency to the first hind-paw response in the hot-plate test (g), and latency to falling in the rotarod test (h) (n = 20 mice per genotype). Data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test (a–d) or two-way ANOVA (strain (S) = Chd8+/∆SL versus Chd8+/∆L; genotype (G) = mutant versus WT) (e–h) after correction for multiple testing by the FDR in the case of two-way ANOVA.
Extended Data Figure 4 Chd8 heterozygous mutant mice show abnormalities in startle response, PPI and social behaviour.
a, b, Acoustic startle response to stimuli of 110 or 120 dB (a) as well as inhibition of the startle response by prepulses of 74 or 78 dB (b). c–e, Protocol (c) as well as time spent around each cage during the first (d) and second (e) tasks for the sociability and social-novelty preference test. f, g, Protocol (f) and time spent around each cage (g) for the social-preference test with a cagemate as a familiar mouse. All data are for 20 mice per genotype. All quantitative data are mean ± s.e.m. **P < 0.01, ***P < 0.001, two-way ANOVA, after correction for multiple testing by the FDR.
Extended Data Figure 5 Behavioural tests in which Chd8 mutant mice show no abnormalities.
a–c, Latency to reach the target (a) and number of errors (b) in training trials as well as time spent around each hole in the probe trial (c) for the Barnes maze test. d–f, During the reversal phase of the Barnes maze test, latency to reach the target (d) and number of errors (e) in reversal training trials as well as time spent around each hole in the reversal probe trial (f). g, Total duration of self-grooming during a 10-min period. h, Nesting score in the nest-building test. i, Total immobility time in the Porsolt forced-swim test. Data are mean ± s.e.m. (n = 20 mice per genotype). P values were determined by Student’s t-test (c, f, g), two-way repeated-measures ANOVA (a, b, d, e), or two-way ANOVA (h, i).
Extended Data Figure 6 Validation of specificity for antibodies to CHD8 and classification of CHD8 binding regions in E14.5 mouse brain.
a, b, Immunoblot analysis (a) and ChIP analysis (b) of lysates of wild-type or CHD8L-null (CAG-CreERT2/Chd8LF/F cells treated with tamoxifen (4-OHT)) MEFs performed with the indicated antibodies to CHD8. Precipitated DNA in b was subjected to qPCR analysis with primers for the indicated CHD8 target genes. Quantitative data are mean ± s.e.m. from three independent experiments. *P < 0.05, **P < 0.01, Student’s t-test. For gel source data, see Supplementary Fig. 1. c, Composition of CHD8 binding peaks in E14.5 mouse brain compared with that for the whole genome. The TSS region here is defined as the region spanning 2 kb upstream to 1 kb downstream of the actual TSS. Upstream, 2–5 kb upstream of the TSS; downstream, transcription end site (TES) to 5 kb downstream; intergenic, all other regions. d, Average positions of CHD8 binding peaks in E14.5 mouse brain for genes classified according to percentile of expression level determined by RNA-seq analysis. e, Venn diagram for overlap of CHD8 target genes between E14.5 and adult mouse brain.
Extended Data Figure 7 Classification of CHD8 binding regions.
a, Heat-map clustering of CHD8 binding peaks with histone modifications in the region spanning 2 kb upstream to 2 kb downstream of the peak in adult mouse brain. b, Composition of CHD8 binding peaks. c, Box plot for the expression of CHD8 target genes in the brain of adult Chd8+/∆L mice relative to that in Chd8+/+ mice with the peaks categorized as in b. The top and bottom edges of the box indicate the twenty-fifth and seventy-fifth percentiles, the central bar indicates the median, and the whiskers indicate non-outlier extremes. d, CHD8 target genes as a proportion of all genes, ASD-related genes (SFARI gene database, updated March 2015), or ASD-related genes in which expression is downregulated in the brain of adult Chd8+/∆L mice.***P < 0.001, Wilcoxon rank-sum test (c) or hypergeometric test (d).
Extended Data Figure 9 Effects of CHD8 haploinsufficiency on gene expression.
a, Volcano plot expanded between log2(fold change) values of −1 and 1 for differentially expressed genes in the brain of adult Chd8+/∆L mice compared with Chd8+/+ mice. Green line indicates P = 0.05. b–f, Volcano plots for differentially expressed genes in the brain of E10.5 (b), E12.5 (c), E14.5 (d), E16.5 (e) or E18.5 (f) Chd8+/∆L mice compared with Chd8+/+ mice. Green lines indicate a twofold change and P = 0.05. g, Functional annotation of genes whose expression changed by a factor of >2 or <0.5 with P < 0.05 in the mutant mice. h, CHD8 binding amount (FPKM) for TSS regions of genes in the brain of adult Chd8+/∆L mice compared with Chd8+/+ mice. i, Box plot of CHD8 binding amount to TSS regions of genes classified according to change in expression level induced by CHD8 haploinsufficiency for the brain of adult Chd8+/∆L and Chd8+/+ mice. j, Absolute NES versus average expression for 1,000 randomly picked gene sets (black dots) and the ASD-related gene set (red dot indicated by the arrow). k, GSEA analysis for Wnt pathway- or p53 pathway-related gene sets (included in the Hallmark gene set collection) in the brain of E10.5, E12.5, E14.5, E16.5, E18.5 or adult Chd8+/∆L mice. *P < 0.05, Student’s t-test (a), Welch’s t-test (b–f) or Wilcoxon rank-sum test (i).
Extended Data Figure 10 REST activation with CHD8 haploinsufficiency is most prominent in E14.5 mouse brain.
a, GSEA analysis for the V$NRSF_01 gene set in the brain of E10.5, E12.5, E14.5, E16.5, E18.5 or adult Chd8+/∆L mice. b, c, GSEA plots of differentially expressed genes in the V$NRSF_01 gene set applied to the data of ref. 20 (b) or ref. 21 (c). KD, knockdown. d, qRT–PCR analysis of Rest mRNA abundance in the whole brain of E10.5, E14.5 or E18.5 Chd8+/∆SL or Chd8+/∆L mice relative to that for wild-type mice. e, Absolute NES versus average expression for 1,000 randomly picked gene sets (black dots) and the V$NRSF_01 gene set (red dot indicated by arrow). f, Lysates of HEK293T cells expressing Flag-tagged versions of CHD8L or FOXP1 (negative control) were subjected to immunoprecipitation (IP) with antibodies to Flag, and the resulting precipitates as well as the original cell lysates (input) were subjected to immunoblot analysis with antibodies to REST and to Flag. g, Box plot for the amount of REST (FPKM) bound to the RE-1 motif of REST target genes with or without bound CHD8 in the Chd8+/+ or Chd8+/∆L brain at E14.5. h, Cycloheximide (CHX) chase analysis of REST in wild-type or CHD8L-depleted (as in Extended Data Fig. 6) MEFs. Data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1, which show the original uncropped western or southern blots for Figures 1a, 4e and Extended Data Figures 1c, d, f, 2e, f, 6a, and 10f, h. The black frames denote how the gels were cropped for the final figure. It also contains a Supplementary Discussion and additional references. (PDF 4279 kb)
Supplementary Table 1
This file contains the source data for behavioural tests. (XLSX 64 kb)
Supplementary Table 2
This file contains statistical analysis of behavioural data. (XLSX 27 kb)
Supplementary Table 3
This file contains results (MACS P value, expression level, fold change, and P value) for total genes of ChIP-seq or RNA-seq analysis used in this study. (XLSX 13895 kb)
Supplementary Table 4
This file contains gene symbols of original gene sets and complete results of GSEA reported in the manuscript. (XLSX 850 kb)
Rights and permissions
About this article
Cite this article
Katayama, Y., Nishiyama, M., Shoji, H. et al. CHD8 haploinsufficiency results in autistic-like phenotypes in mice. Nature 537, 675–679 (2016). https://doi.org/10.1038/nature19357
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19357
This article is cited by
-
Transcriptomic dysregulation and autistic-like behaviors in Kmt2c haploinsufficient mice rescued by an LSD1 inhibitor
Molecular Psychiatry (2024)
-
CHD8-related disorders redefined: an expanding spectrum of dystonic phenotypes
Journal of Neurology (2024)
-
CHD8 mutations increase gliogenesis to enlarge brain size in the nonhuman primate
Cell Discovery (2023)
-
REST in the Road Map of Brain Development
Cellular and Molecular Neurobiology (2023)
-
Developmental pyrethroid exposure and age influence phenotypes in a Chd8 haploinsufficient autism mouse model
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.