Abstract
Sleep disconnects animals from the external world, at considerable risks and costs that must be offset by a vital benefit. Insight into this mysterious benefit will come from understanding sleep homeostasis: to monitor sleep need, an internal bookkeeper must track physiological changes that are linked to the core function of sleep1. In Drosophila, a crucial component of the machinery for sleep homeostasis is a cluster of neurons innervating the dorsal fan-shaped body (dFB) of the central complex2,3. Artificial activation of these cells induces sleep2, whereas reductions in excitability cause insomnia3,4. dFB neurons in sleep-deprived flies tend to be electrically active, with high input resistances and long membrane time constants, while neurons in rested flies tend to be electrically silent3. Correlative evidence thus supports the simple view that homeostatic sleep control works by switching sleep-promoting neurons between active and quiescent states3. Here we demonstrate state switching by dFB neurons, identify dopamine as a neuromodulator that operates the switch, and delineate the switching mechanism. Arousing dopamine4,5,6,7,8 caused transient hyperpolarization of dFB neurons within tens of milliseconds and lasting excitability suppression within minutes. Both effects were transduced by Dop1R2 receptors and mediated by potassium conductances. The switch to electrical silence involved the downregulation of voltage-gated A-type currents carried by Shaker and Shab, and the upregulation of voltage-independent leak currents through a two-pore-domain potassium channel that we term Sandman. Sandman is encoded by the CG8713 gene and translocates to the plasma membrane in response to dopamine. dFB-restricted interference with the expression of Shaker or Sandman decreased or increased sleep, respectively, by slowing the repetitive discharge of dFB neurons in the ON state or blocking their entry into the OFF state. Biophysical changes in a small population of neurons are thus linked to the control of sleep–wake state.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Saper, C. B., Fuller, P. M., Pedersen, N. P., Lu, J. & Scammell, T. E. Sleep state switching. Neuron 68, 1023–1042 (2010)
Donlea, J. M., Thimgan, M. S., Suzuki, Y., Gottschalk, L. & Shaw, P. J. Inducing sleep by remote control facilitates memory consolidation in Drosophila . Science 332, 1571–1576 (2011)
Donlea, J. M., Pimentel, D. & Miesenböck, G. Neuronal machinery of sleep homeostasis in Drosophila . Neuron 81, 860–872 (2014)
Liu, Q., Liu, S., Kodama, L., Driscoll, M. R. & Wu, M. N. Two dopaminergic neurons signal to the dorsal fan-shaped body to promote wakefulness in Drosophila . Curr. Biol. 22, 2114–2123 (2012)
Lima, S. Q. & Miesenböck, G. Remote control of behavior through genetically targeted photostimulation of neurons. Cell 121, 141–152 (2005)
Andretic, R., van Swinderen, B. & Greenspan, R. J. Dopaminergic modulation of arousal in Drosophila . Curr. Biol. 15, 1165–1175 (2005)
Kume, K., Kume, S., Park, S. K., Hirsh, J. & Jackson, F. R. Dopamine is a regulator of arousal in the fruit fly. J. Neurosci. 25, 7377–7384 (2005)
Ueno, T. et al. Identification of a dopamine pathway that regulates sleep and arousal in Drosophila . Nature Neurosci. 15, 1516–1523 (2012)
Shaw, P. J., Cirelli, C., Greenspan, R. J. & Tononi, G. Correlates of sleep and waking in Drosophila melanogaster . Science 287, 1834–1837 (2000)
Hendricks, J. C. et al. Rest in Drosophila is a sleep-like state. Neuron 25, 129–138 (2000)
Zemelman, B. V., Lee, G. A., Ng, M. & Miesenböck, G. Selective photostimulation of genetically chARGed neurons. Neuron 33, 15–22 (2002)
Claridge-Chang, A. et al. Writing memories with light-addressable reinforcement circuitry. Cell 139, 405–415 (2009)
Bokoch, G. M., Katada, T., Northup, J. K., Ui, M. & Gilman, A. G. Purification and properties of the inhibitory guanine nucleotide-binding regulatory component of adenylate cyclase. J. Biol. Chem. 259, 3560–3567 (1984)
Andrade, R. & Nicoll, R. A. Pharmacologically distinct actions of serotonin on single pyramidal neurones of the rat hippocampus recorded in vitro . J. Physiol. (Lond.) 394, 99–124 (1987)
Reale, V., Hannan, F., Hall, L. M. & Evans, P. D. Agonist-specific coupling of a cloned Drosophila melanogaster D1-like dopamine receptor to multiple second messenger pathways by synthetic agonists. J. Neurosci. 17, 6545–6553 (1997)
Connor, J. A. & Stevens, C. F. Prediction of repetitive firing behaviour from voltage clamp data on an isolated neurone soma. J. Physiol. (Lond.) 213, 31–53 (1971)
Timpe, L. C. et al. Expression of functional potassium channels from Shaker cDNA in Xenopus oocytes. Nature 331, 143–145 (1988)
Iverson, L. E., Tanouye, M. A., Lester, H. A., Davidson, N. & Rudy, B. A-type potassium channels expressed from Shaker locus cDNA. Proc. Natl Acad. Sci. USA 85, 5723–5727 (1988)
Littleton, J. T. & Ganetzky, B. Ion channels and synaptic organization: analysis of the Drosophila genome. Neuron 26, 35–43 (2000)
Harris-Warrick, R. M., Coniglio, L. M., Barazangi, N., Guckenheimer, J. & Gueron, S. Dopamine modulation of transient potassium current evokes phase shifts in a central pattern generator network. J. Neurosci. 15, 342–358 (1995)
Cirelli, C. et al. Reduced sleep in Drosophila Shaker mutants. Nature 434, 1087–1092 (2005)
Bushey, D., Huber, R., Tononi, G. & Cirelli, C. Drosophila hyperkinetic mutants have reduced sleep and impaired memory. J. Neurosci. 27, 5384–5393 (2007)
Koh, K. et al. Identification of SLEEPLESS, a sleep-promoting factor. Science 321, 372–376 (2008)
Enyedi, P. & Czirják, G. Molecular background of leak K+ currents: two-pore domain potassium channels. Physiol. Rev. 90, 559–605 (2010)
Schiavo, G., Matteoli, M. & Montecucco, C. Neurotoxins affecting neuroexocytosis. Physiol. Rev. 80, 717–766 (2000)
Hounsgaard, J., Hultborn, H., Jespersen, B. & Kiehn, O. Bistability of α-motoneurones in the decerebrate cat and in the acute spinal cat after intravenous 5-hydroxytryptophan. J. Physiol. (Lond.) 405, 345–367 (1988)
Marder, E., Abbott, L. F., Turrigiano, G. G., Liu, Z. & Golowasch, J. Memory from the dynamics of intrinsic membrane currents. Proc. Natl Acad. Sci. USA 93, 13481–13486 (1996)
Marder, E. & Thirumalai, V. Cellular, synaptic and network effects of neuromodulation. Neural Netw. 15, 479–493 (2002)
Baxter, D. A. & Byrne, J. H. Serotonergic modulation of two potassium currents in the pleural sensory neurons of Aplysia. J. Neurophysiol. 62, 665–679 (1989)
Nicola, S. M., Surmeier, J. & Malenka, R. C. Dopaminergic modulation of neuronal excitability in the striatum and nucleus accumbens. Annu. Rev. Neurosci. 23, 185–215 (2000)
Jenett, A. et al. A GAL4-driver line resource for Drosophila neurobiology. Cell Reports 2, 991–1001 (2012)
Friggi-Grelin, F. et al. Targeted gene expression in Drosophila dopaminergic cells using regulatory sequences from tyrosine hydroxylase. J. Neurobiol. 54, 618–627 (2003)
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999)
Pfeiffer, B. D. et al. Refinement of tools for targeted gene expression in Drosophila . Genetics 186, 735–755 (2010)
McGuire, S. E., Le, P. T., Osborn, A. J., Matsumoto, K. & Davis, R. L. Spatiotemporal rescue of memory dysfunction in Drosophila . Science 302, 1765–1768 (2003)
Ferris, J., Ge, H., Liu, L. & Roman, G. G (o) signaling is required for Drosophila associative learning. Nature Neurosci. 9, 1036–1040 (2006)
Klapoetke, N. C. et al. Independent optical excitation of distinct neural populations. Nature Methods 11, 338–346 (2014)
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila . Nature 448, 151–156 (2007)
Shaw, P. J., Tononi, G., Greenspan, R. J. & Robinson, D. F. Stress response genes protect against lethal effects of sleep deprivation in Drosophila . Nature 417, 287–291 (2002)
Buchner, E. Elementary movement detectors in an insect visual-system. Biol. Cybern. 24, 85–101 (1976)
Seelig, J. D. et al. Two-photon calcium imaging from head-fixed Drosophila during optomotor walking behavior. Nature Methods 7, 535–540 (2010)
Connor, J. A. & Stevens, C. F. Voltage clamp studies of a transient outward membrane current in gastropod neural somata. J. Physiol. (Lond.) 213, 21–30 (1971)
Acknowledgements
We thank S. Birman, R. Davis, B. Dickson, V. Jayaraman, L. Luo, G. Roman, G. Rubin, the Bloomington Stock Center, and the Vienna Drosophila Resource Center for flies. This work was supported by grants (to G.M.) from the Wellcome Trust, the Gatsby Charitable Foundation, the Oxford Martin School, and the National Institutes of Health. J.M.D. was the recipient of a postdoctoral fellowship from the Human Frontier Science Program; S.M.S. is a Commonwealth Scholar.
Author information
Authors and Affiliations
Contributions
D.P., J.M.D. and G.M. designed the study and analysed the results. All electrophysiological recordings were done by D.P.; J.M.D. performed molecular manipulations and behavioural analyses with the help of S.M.S. and A.J.F.T. C.B.T. developed instrumentation. G.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Optogenetic stimulation of dopaminergic neurons.
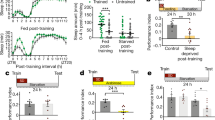
Dopaminergic neurons expressing CsChrimson under TH-GAL4 control were driven with 3 ms pulses of 630 nm light at the indicated frequencies. Optical power at the sample was ~28 mW cm−2. a, Examples of voltage responses to optical pulse trains. b, The ratio of light-evoked action potentials to optical pulses was close to 1 at driving frequencies between 5 and 20 Hz (n = 36 trials on 6 cells). Data are means ± s.e.m.
Extended Data Figure 2 Changes in sleep after interference with Dop1R2 signalling are consistent with diminished sensitivity to arousing dopamine.
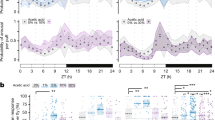
a, Sleep during a 24 h day in homozygous carriers of the Dop1R2MI08664 allele (red, n = 32 flies) and heterozygous controls (black, n = 31 flies). Data are means ± s.e.m. Two-way repeated-measures ANOVA detected a significant interaction between time of day and genotype (P < 0.0001). b, Sleep during a 24 h day in homozygous carriers of the Dop1R2MB05108 allele (red, n = 28 flies) and heterozygous controls (black, n = 32 flies). Data are means ± s.e.m. Two-way repeated-measures ANOVA failed to detect a significant interaction between time of day and genotype (P = 0.4736). c, Sleep in homozygous and heterozygous carriers of the Dop1R2MI08664 or Dop1R2MB05108 alleles (circles, individual flies; horizontal lines, group means). Mann–Whitney tests detected a significant effect of the Dop1R2MI08664 allele (P = 0.0219, red), but not of the Dop1R2MB05108 allele (P = 0.6750). The Dop1R2MB05108 allele contains a transposon insertion in a non-coding region of the Dop1R2 gene, which reduces mRNA levels in homozygous carriers by only 14% (ref. 4), thus explaining the lack of a phenotype. The inability of Dop1R2MB05108 to suppress the short-sleeping phenotype of flies with enhanced dopaminergic transmission4 therefore does not argue against a role of Dop1R2 in the dFB. d, Sleep during a 24 h day in flies expressing R23E10-GAL4-driven RNAi targeting Dop1R2 (red, n = 48 flies) and parental controls (open symbols: R23E10-GAL4, n = 48 flies; filled symbols: undriven UAS-Dop1R2RNAi, n = 32 flies). Data are means ± s.e.m. Two-way repeated-measures ANOVA detected a significant interaction between time of day and genotype (P < 0.0001). e, Average length of daytime sleep bouts in flies expressing R23E10-GAL4-driven RNAi targeting Dop1R2 and parental controls. Data are means ± s.e.m. One-way ANOVA detected a significant genotype effect (P = 0.0015); red indicates a significant difference from both parental controls in pairwise post-hoc comparisons. f, Sleep in flies with temperature-inducible R23E10-GAL4-driven expression of PTX and parental controls (circles, individual flies; horizontal lines, group means). Two-way ANOVA detected a significant interaction between genotype and temperature (P = 0.0143); blue indicates a significant difference between inducing and non-inducing temperatures in pairwise post-hoc comparisons. g, Average length of daytime sleep bouts in flies with temperature-inducible R23E10-GAL4-driven expression of PTX and parental controls. Data are means ± s.e.m. Two-way ANOVA detected a significant interaction between genotype and temperature (P = 0.0002); blue indicates a significant increase upon switching from non-inducing to inducing temperatures in pairwise post-hoc comparisons.
Extended Data Figure 3 Dopamine hyperpolarizes dFB neurons and inhibits their spiking.
a, Membrane potential of a dFB neuron during a 250 ms pulse of dopamine. b, Average amplitude of hyperpolarization evoked by dopamine in the indicated numbers of cells. Data are means ± s.e.m. Kruskal–Wallis test detected a significant difference between groups (P < 0.0001); asterisks indicate significant differences from controls in pairwise post-hoc comparisons.
Extended Data Figure 4 Optogenetic stimulation of dopaminergic neurons switches dFB neurons to quiescence.
Flies expressing CsChrimson under TH-GAL4 control in dopaminergic neurons were photostimulated with 3 ms pulses of 630 nm light at 20 Hz. a, Voltage responses to current steps were recorded in the same cell, before and after optogenetic stimulation of dopaminergic neurons (black and red traces). Red and grey traces in the OFF state (right) indicate current injections matching or exceeding those in the ON state, respectively (left). b, c, Time courses of changes in input resistance (Rm) and membrane time constant (τm) of dFB neurons during optogenetic stimulation of dopaminergic neurons (n = 7 cells). Data are means ± s.e.m. One-way repeated-measures ANOVA detected significant effects of time (P = 0.0135 for Rm; P = 0.0222 for τm).
Extended Data Figure 5 Membrane properties of dFB neurons in the ON state.
a, Input resistances (Rm) of the indicated numbers of cells. Data are means ± s.e.m. Kruskal–Wallis test failed to detect a significant difference between groups (P = 0.8997). b, Membrane time constants (τm) of the indicated numbers of cells. Data are means ± s.e.m. Kruskal–Wallis test failed to detect a significant difference between groups (P = 0.1682).
Extended Data Figure 6 Measurements of potassium currents in voltage clamp.
a, Voltage steps from a holding potential of −110 mV (top) elicited the full complement of potassium currents expressed by a dFB neuron (Itotal, bottom). b, Stepping the same neuron from a holding potential of −30 mV (top) elicited potassium currents lacking the A-type component (Inon-A, bottom). c, Digital subtraction of Inon-A (b, bottom) from Itotal (a, bottom) yielded an estimate of IA. Note the expanded timescale. d, Individual (grey) and average (black) A-type currents of seven dFB neurons, evoked by step depolarization to 40 mV. The magenta line represents a single-exponential fit to the average.
Extended Data Figure 7 Loss of Shaker and its interacting partners, Hyperkinetic and Sleepless, from dFB neurons has similar effects on sleep.
Sleep in flies expressing R23E10-GAL4-driven RNAi targeting Shaker, Hyperkinetic or sleepless and parental controls (circles, individual flies; horizontal lines, group means). One-way ANOVA detected a significant genotype effect (P < 0.0001); green indicates significant differences from both parental controls in pairwise post-hoc comparisons.
Rights and permissions
About this article
Cite this article
Pimentel, D., Donlea, J., Talbot, C. et al. Operation of a homeostatic sleep switch. Nature 536, 333–337 (2016). https://doi.org/10.1038/nature19055
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19055
This article is cited by
-
Phylogenetically distant animals sleep: why do sleep researchers care?
Biology & Philosophy (2024)
-
A rise-to-threshold process for a relative-value decision
Nature (2023)
-
A neural circuit linking learning and sleep in Drosophila long-term memory
Nature Communications (2022)
-
Neural Control of Action Selection Among Innate Behaviors
Neuroscience Bulletin (2022)
-
Compartment specific regulation of sleep by mushroom body requires GABA and dopaminergic signaling
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.