Abstract
During sleep, most animal species enter a state of reduced consciousness characterized by a marked sensory disconnect. Yet some processing of the external world must remain intact, given that a sleeping animal can be awoken by intense stimuli (for example, a loud noise or a bright light) or by soft but qualitatively salient stimuli (for example, the sound of a baby cooing or hearing one’s own name1,2,3). How does a sleeping brain retain the ability to process the quality of sensory information? Here we present a paradigm to study the functional underpinnings of sensory discrimination during sleep in Drosophila melanogaster. We show that sleeping vinegar flies, like humans, discern the quality of sensory stimuli and are more likely to wake up in response to salient stimuli. We also show that the salience of a stimulus during sleep can be modulated by internal states. We offer a prototypical blueprint detailing a circuit involved in this process and its modulation as evidence that the system can be used to explore the cellular underpinnings of how a sleeping brain experiences the world.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw data used for analysis are made available through the Supplementary Information and through a Zenodo repository (https://doi.org/10.5281/zenodo.5109970). Source data are provided with this paper.
Code availability
All scripts used for analysis are made available through the Supplementary Information and through a Zenodo repository (https://doi.org/10.5281/zenodo.5109970).
References
Langford, G. W., Meddis, R. & Pearson, A. J. D. Awakening latency from sleep for meaningful and non-meaningful stimuli. Psychophysiology 11, 1–5 (1974).
Buendia, N., Goode, M., Sierra, G. & Segundo, J. P. Responsiveness and discrimination during sleep. Experientia 19, 208–209 (1963).
Oswald, I., Taylor, A. M. & Treisman, M. Discriminative responses to stimulation during human sleep. Brain 83, 440–453 (1960).
Twyver, H. V. & Garrett, W. Arousal threshold in the rat determined by “meaningful” stimuli. Behav. Biol. 7, 205–215 (1972).
Siegel, J. & Langley, T. D. Arousal threshold in the cat as a function of sleep phase and stimulus significance. Experientia 21, 740–741 (1965).
Tannenbaum, P. L. et al. Orexin receptor antagonist-induced sleep does not impair the ability to wake in response to emotionally salient acoustic stimuli in dogs. Front. Behav. Neurosci. 8, 182 (2014).
Beckwith, E. J. & French, A. S. Sleep in Drosophila and its context. Front. Physiol. 10, 1167 (2019).
Geissmann, Q. et al. Ethoscopes: an open platform for high-throughput ethomics. PLoS Biol. 15, e2003026 (2017).
Geissmann, Q., Beckwith, E. J. & Gilestro, G. F. Most sleep does not serve a vital function: evidence from Drosophila melanogaster. Sci. Adv. 5, eaau9253 (2019).
Knaden, M., Strutz, A., Ahsan, J., Sachse, S. & Hansson, B. S. Spatial representation of odorant valence in an insect brain. Cell Rep. 1, 392–399 (2012).
Joseph, R. M., Devineni, A. V., King, I. F. G. & Heberlein, U. Oviposition preference for and positional avoidance of acetic acid provide a model for competing behavioral drives in Drosophila. Proc. Natl Acad. Sci. USA 106, 11352–11357 (2009).
Gao, X. J., Clandinin, T. R. & Luo, L. Extremely sparse olfactory inputs are sufficient to mediate innate aversion in Drosophila. PLoS ONE 10, e0125986 (2015).
Chakir, M., Peridy, O., Capy, P., Pla, E. & David, J. R. Adaptation to alcoholic fermentation in Drosophila: a parallel selection imposed by environmental ethanol and acetic acid. Proc. Natl Acad. Sci. USA 90, 3621–3625 (1993).
Faville, R., Kottler, B., Goodhill, G. J., Shaw, P. J. & van Swinderen, B. How deeply does your mutant sleep? Probing arousal to better understand sleep defects in Drosophila. Sci. Rep. 5, 8454 (2015).
Tamaki, M., Bang, J. W., Watanabe, T. & Sasaki, Y. Night watch in one brain hemisphere during sleep associated with the first-night effect in humans. Curr. Biol. 26, 1190–1194 (2016).
Newman, J. D. Neural circuits underlying crying and cry responding in mammals. Behav. Brain Res. 182, 155–165 (2007).
Ebrahim, I. O., Shapiro, C. M., Williams, A. J. & Fenwick, P. B. Alcohol and sleep I: effects on normal sleep. Alcohol. Clin. Exp. Res. 37, 539–549 (2013).
Masse, N. Y., Turner, G. C. & Jefferis, G. S. X. E. Olfactory information processing in Drosophila. Curr. Biol. 19, R700–R713 (2009).
Sayin, S. et al. A neural circuit arbitrates between persistence and withdrawal in hungry Drosophila. Neuron 104, 544–558.e6 (2019).
Hattori, D. et al. Representations of novelty and familiarity in a mushroom body compartment. Cell 169, 956–969.e17 (2017).
Seidner, G. et al. Identification of neurons with a privileged role in sleep homeostasis in Drosophila melanogaster. Curr. Biol. 25, 2928–2938 (2015).
Wolff, T., Iyer, N. A. & Rubin, G. M. Neuroarchitecture and neuroanatomy of the Drosophila central complex: a GAL4-based dissection of protocerebral bridge neurons and circuits. J. Comp. Neurol. 523, 997–1037 (2015).
Ni, J. D. et al. Differential regulation of the Drosophila sleep homeostat by circadian and arousal inputs. eLife 8, e40487 (2019).
Modi, M. N., Shuai, Y. & Turner, G. C. The Drosophila mushroom body: from architecture to algorithm in a learning circuit. Annu. Rev. Neurosci. 43, 465–484 (2020).
Devineni, A. V., Sun, B., Zhukovskaya, A. & Axel, R. Acetic acid activates distinct taste pathways in Drosophila to elicit opposing, state-dependent feeding responses. eLife 8, e47677 (2019).
Ko, K. I. et al. Starvation promotes concerted modulation of appetitive olfactory behavior via parallel neuromodulatory circuits. eLife 4, e08298 (2015).
Keene, A. C. et al. Clock and cycle limit starvation-induced sleep loss in Drosophila. Curr. Biol. 20, 1209–1215 (2010).
Lin, C.-C., Riabinina, O. & Potter, C. J. Olfactory behaviors assayed by computer tracking of Drosophila in a four-quadrant olfactometer. J. Vis. Exp. 114, e54346 (2016).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Talay, M. et al. Transsynaptic mapping of second-order taste neurons in flies by trans-Tango. Neuron 96, 783-795.e4 (2017).
Feinberg, E. H. et al. GFP reconstitution across synaptic partners (GRASP) defines cell contacts and synapses in living nervous systems. Neuron 57, 353–363 (2008).
Geissmann, Q., Rodriguez, L. G., Beckwith, E. J. & Gilestro, G. F. Rethomics: an R framework to analyse high-throughput behavioural data. PLoS ONE 14, e0209331 (2019).
French, A., Geissmann, Q., Beckwith, E. & Gilestro, G. Sensory processing during sleep in Drosophila melanogaster—ethoscope dataset. Zenodo https://doi.org/10.5281/zenodo.5109970 (2021).
Acknowledgements
We thank all members of the Gilestro laboratory for discussions and intellectual contributions and the fly community for sharing reagents and protocols; special thanks to G. Rubin for sharing unpublished information and to A. Bates for sharing some beautiful drawings on scidraw.io. A.S.F. was supported by the Biotechnology and Biological Sciences Research Council (BBSRC) through grants BB/M003930/1 and BB/R018839/1. E.J.B. was supported by EMBO ALTF 57–2014 and by the People Programme (Marie Curie Actions) of the European Union’s Eighth Framework Programme H2020 under REA grant agreement 705930. Q.G. was supported by a BBSRC DTP scholarship BB/J014575/1. We thank the Imperial College London Advanced Hackspace (ICAH) and the Facility for Imaging by Light Microscopy (FILM) at Imperial College London, part-supported by funding from the Wellcome Trust (grant 104931/Z/14/Z) and BBSRC (grant BB/L015129/1).
Author information
Authors and Affiliations
Contributions
A.F. and E.J.B. performed all of the experiments. Q.G. wrote the software for the collection and analysis of the data. A.F. and G.F.G. analysed the data. A.F. and G.F.G. devised and planned all of the experiments. All authors contributed to the preparation of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Occurrence and length of sleep bouts vary during the day.
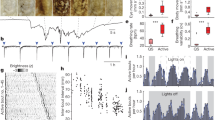
a, Schematics of the experimental setup. b, Average numbers of puffs per fly per hour in three conditions (0%; 5%; 10% acetic acid. Ns 106, 105, 103 respectively). The top grey bars show how experiments were conducted at eight independent, overlapping intervals. c, Mean response per fly at different time point during the day, grouped by concentration of acetic acid. Same dataset as in Fig. 1d. d, Total distribution of sleep bouts by length, binned at 1-minute intervals during three time points representing different types of sleep (ZT 4-6, ZT 13-15 and ZT18-20). The three grey vertical lines indicate the bins for 5-, 8-, and 12-minutes sleep respectively as chosen for the analysis in Fig. 1c–e. In all panels, errors are shown as bootstrapped 95% CI.
Extended Data Fig. 2 Changes in internal states affects arousal threshold during sleep.
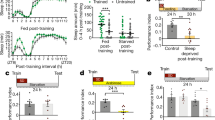
a, Schematic showing experimental procedures used in b, c and main Fig. 2a, b. b, Responses to 5% acetic acid in mock control flies (left) and flies previously intoxicated with vapours of 20% ethanol (right). Ethanol or mock exposure were limited to a 1-hour window between ZT15.5-16.5 and arousal was quantified and shown for ZT18-20. c, Quantification of the stimulus-evoked response in rested control flies (left) or flies that were previously deprived of sleep for 12h (right). Sleep deprivation was performed during the subjective night (ZT12-24) and waking to an olfactory stimulus (5% acetic acid) was measured during the subsequent day (ZT0-6). In all panels, errors are shown as bootstrapped 95% CI.
Extended Data Fig. 3 Inhibition of MB011B reduces arousal threshold.
a, Waking responses of control flies (MBON-GAL4/+) and those with temperature inhibited MBONs (MBON-GAL4/ShiTS) to 5% acetic acid between ZT15-24. b, 9 h response profile for MB011B ShiTS siliencing (light blue) compared to its appropriate parental controls (shades of grey). In all panels, errors are shown as bootstrapped 95% CI.
Extended Data Fig. 4 Activation or inhibition of some MBONs modulates sleep.
a, b, 24h sleep profiles (a) and mean sleep amount (b) exhibited by flies with inhibited MBON neurons (MBON/ShiTS: blue) or those carrying only the MBON-GAL4 (MBON-GAL4/+: grey solid) or only UAS-ShiTS (UAS-ShiTS /+: grey hashed) transgene over a 24hr baseline day. Ns underneath indicate the number of individual flies and refer to b and a. c, d, 24hr sleep profiles (c) and mean sleep amount (d) exhibited by flies with activated MBON neurons (MBON/dTRPA1:red) or those carrying only the MBON-GAL4 (MBON-GAL4/+: grey solid) or only UAS-dTRPA1 (UAS-dTRPA1: grey hashed) transgene over a 24hr baseline day. Ns underneath indicate the number of individual flies and refer to c, d. In all panels, errors are shown as bootstrapped 95% CI.
Extended Data Fig. 5 Activation or inhibition R23E10 neurons alters behavior.
a, Waking responses of control flies (UAS-ShiTS and UAS-TRPA1) and those with temperature manipulated dFSB neurons (23E10-GAL4/ShiTS and 23E10-GAL4/dTrpa1) to 5% acetic acid, between ZT18-20. Experiment performed at 29 °C. b, 24h sleep profile of dFSB inactivation through ShiTS (blue) compared to parental controls (grey). c, 24h sleep profile of dFSB forced activation through dTRPA1 (red) compared to parental controls (grey). d–f, Walking (d), micromovements (e) and X-position profiles (f) of control flies (UAS-ShiTS: grey hashed, R23E10-GAL4: grey) and those with inhibited dFSB neurons (R23E210-GAL4/UAS-ShiTS) over a 24 h baseline day. g–i, Walking (g), micromovements (h) and X-position profiles (i) of control flies (UAS-TRPA1: grey hashed, R23E10-GAL4: grey) and those with activated dFSB neurons (R23E210-GAL4/UAS-TRPA1) over a 24 h baseline day. Experiments preformed at 29 °C on 2-3 day old male flies in 12h:12h L:D cycle. In all panels, errors are shown as bootstrapped 95% CI.
Supplementary information
Supplementary Table 1
Technical summary of odourants used in Fig. 2
Supplementary Data 1
All raw data and scripts used for analysis
Rights and permissions
About this article
Cite this article
French, A.S., Geissmann, Q., Beckwith, E.J. et al. Sensory processing during sleep in Drosophila melanogaster. Nature 598, 479–482 (2021). https://doi.org/10.1038/s41586-021-03954-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03954-w
This article is cited by
-
Shared science’s time to shine
Lab Animal (2023)
-
Microengineered devices enable long-term imaging of the ventral nerve cord in behaving adult Drosophila
Nature Communications (2022)
-
Reduced neural feedback signaling despite robust neuron and gamma auditory responses during human sleep
Nature Neuroscience (2022)
-
Flies sense the world while sleeping
Nature (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.